This is the fourth in our series of articles about searching for unknown close family members, specifically; parents, grandparents, or siblings. However, these same techniques can be applied by genealogists to ancestors further back in time as well.
- I introduced the “In Search of” series in the article, DNA: In Search of…New Series Launches.
- In the second article, DNA: In Search of…What Do You Mean I’m Not Related to My Family? – and What Comes Next? we discussed the discovery that something was amiss when you don’t match a family member that you expect to match, then how to make sure a vial or upload mix-up didn’t happen. Next, I covered the basics of the four kinds of DNA tests you’ll be able to use to solve your mystery.
- In the third article, In Search of…Vendor Features, Strengths, and Testing Strategies, we discussed testing goals and strategies, including testing with and uploading to multiple autosomal DNA vendors, Y DNA, and mitochondrial DNA testing. We reviewed the vendor’s strengths and the benefits of combining vendor information and resources.
In this article, we discuss endogamy – how to determine if you have it, from what population, and how to follow the road signs.
After introductions, we will be covering the following topics:
- Pedigree collapse and endogamy
- Endogamous groups
- The challenge(s) of endogamy
- Endogamy and unknown close relatives (parents, grandparents)
- Ethnicity and Populations
- Matches
- AutoClusters
- Endogamous Relationships
- Endogamous DNA Segments
- “Are Your Parents Related?” Tool
- Surnames
- Projects
- Locations
- Y DNA, Mitochondrial DNA, and Endogamy
- Endogamy Tools Summary Tables
- Summary of Endogamy Tools by Vendor
- Summary of Endogamous Populations Identified by Each Tool
- Summary of Tools to Assist People Seeking Unknown Parents and Grandparents
What Is Endogamy and Why Does It Matter?
Endogamy occurs when a group or population of people intermarry among themselves for an extended period of time, without the introduction of many or any people from outside of that population.
The effect of this continual intermarriage is that the founders’ DNA simply gets passed around and around, eventually in small segments.
That happens because there is no “other” DNA to draw from within the population. Knowing or determining that you have endogamy helps make sense of DNA matching patterns, and those patterns can lead you to unknown relatives, both close and distant.
This Article
This article serves two purposes.
- This article is educational and relevant for all researchers. We discuss endogamy using multiple tools and examples from known endogamous people and populations.
- In order to be able to discern endogamy when we don’t know who our parents or grandparents are, we need to know what signs and signals to look for, and why, which is based on what endogamy looks like in people who know their heritage.

There’s no crystal ball – no definitive “one-way” arrow, but there are a series of indications that suggest endogamy.
Depending on the endogamous population you’re dealing with, those signs aren’t always the same.
If you’re sighing now, I understand – but that’s exactly WHY I wrote this article.
We’re covering a lot of ground, but these road markers are invaluable diagnostic tools.
I’ve previously written about endogamy in the articles:
Let’s start with definitions.
Pedigree Collapse and Endogamy
Pedigree collapse isn’t the same as endogamy. Pedigree collapse is when you have ancestors that repeat in your tree.

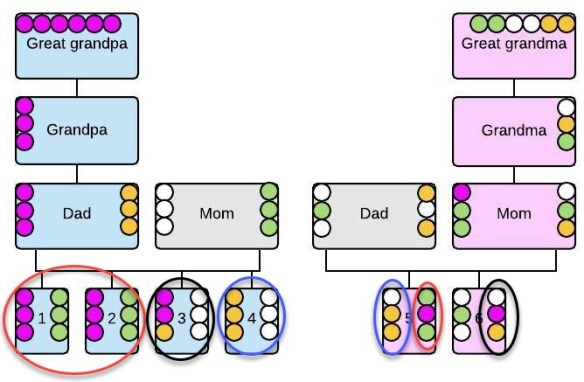
In this example, the parents of our DNA tester are first cousins, which means the tester shares great-grandparents on both sides and, of course, the same ancestors from there on back in their tree.
This also means they share more of those ancestors’ DNA than they would normally share.

John Smith and Mary Johnson are both in the tree twice, in the same position as great-grandparents. Normally, Tester Smith would carry approximately 12.5% of each of his great-grandparents’ DNA, assuming for illustration purposes that exactly 50% of each ancestor’s DNA is passed in each generation. In this case, due to pedigree collapse, 25% of Tester Smith’s DNA descends from John Smith, and another 25% descends from Mary Johnson, double what it would normally be. 25% is the amount of DNA contribution normally inherited from grandparents, not great-grandparents.
While we may find first cousin marriages a bit eyebrow-raising today, they were quite common in the past. Both laws and customs varied with the country, time, social norms, and religion.
Pedigree Collapse and Endogamy is NOT the Same
You might think that pedigree collapse and endogamy is one and the same, but there’s a difference. Pedigree collapse can lead to endogamy, but it takes more than one instance of pedigree collapse to morph into endogamy within a population. Population is the key word for endogamy.
The main difference is that pedigree collapse occurs with known ancestors in more recent generations for one person, while endogamy is longer-term and systemic in a group of people.
Picture a group of people, all descended from Tester Smith’s great-grandparents intermarrying. Now you have the beginnings of endogamy. A couple hundred or a few hundred years later, you have true endogamy.
In other words, endogamy is pedigree collapse on a larger scale – think of a village or a church.

My ancestors’ village of Schnait, in Germany, is shown above in 1685. One church and maybe 30 or 40 homes. According to church and other records, the same families had inhabited this village, and region, for generations. It’s a sure bet that both pedigree collapse and endogamy existed in this small community.
If pedigree collapse happens over and over again because there are no other people within the community to marry, then you have endogamy. In other words, with endogamy, you assuredly DO have historical pedigree collapse, generally back in time, often before you can identify those specific ancestors – because everyone descends from the same set of founders.
Endogamy Doesn’t Necessarily Indicate Recent Pedigree Collapse
With deep, historic endogamy, you don’t necessarily have recent pedigree collapse, and in fact, many people do not. Jewish people are a good example of this phenomenon. They shared ancestors for hundreds or thousands of years, depending on which group we are referring to, but in recent, known, generations, many Jewish people aren’t related. Still, their DNA often matches each other.
The good news is that there are telltale signs and signals of endogamy.
The bad news is that not all of these are obvious, meaning as an aid to people seeking clues about unknown close relatives, and other “signs” aren’t what they are believed to be.
Let’s step through each endogamy identifier, or “hint,” and then we will review how we can best utilize this information.
First, let’s take a look at groups that are considered to be endogamous.
Endogamous Groups
Jewish People – Specifically groups that were isolated from other groups of Jewish (and other) people; Ashkenazi (Germany, Northern France, and diaspora), Sephardic (Spanish, Iberia, and diaspora), Mizrahi (Israel, Middle Eastern, and diaspora,) Ethiopian Jews, and possibly Jews from other locations such as Mountain Jews from Kazakhstan and the Caucasus.
Acadians – Descendants of about 60 French families who settled in “Acadia” beginning about 1604, primarily on the island of Nova Scotia, and intermarried among themselves and with the Mi’kmaq people. Expelled by the English in 1755, they were scattered in groups to various diasporic regions where they continued to intermarry and where their descendants are found today. Some Acadians became the Cajuns of Louisiana.
Anabaptist Protestant Faiths – Amish, Mennonite, and Brethren (Dunkards) and their offshoots are Protestant religious sects founded in Europe in the 14th, 15th, and 16th centuries on the principle of baptizing only adults or people who are old enough to choose to follow the faith, or rebaptizing people who had been previously baptized as children. These Anabaptist faiths tend to marry within their own group or church and often expel those who marry outside of the faith. Many emigrated to the American colonies and elsewhere, seeking religious freedom. Occasionally those groups would locate in close proximity and intermarry, but not marry outside of other Anabaptist denominations.
Native American (Indigenous) People – all indigenous peoples found in North and South America before European colonization descended from a small number of original founders who probably arrived at multiple times.
Indigenous Pacific Islanders – Including indigenous peoples of Australia, New Zealand, and Hawaii prior to colonization. They are probably equally as endogamous as Native American people, but I don’t have specific examples to share.
Villages – European or other villages with little inflow or whose residents were restricted from leaving over hundreds of years.
Other groups may have significant multiple lines of pedigree collapse and therefore become endogamous over time. Some people from Newfoundland, French Canadians, and Mormons (Church of Jesus Christ of Latter-Day Saints) come to mind.
Endogamy is a process that occurs over time.
Endogamy and Unknown Relatives
If you know who your relatives are, you may already know you’re from an endogamous population, but if you’re searching for close relatives, it’s helpful to be able to determine if you have endogamous heritage, at least in recent generations.
If you know nothing about either parent, some of these tools won’t help you, at least not initially, but others will. However, as you add to your knowledge base, the other tools will become more useful.
If you know the identity of one parent, this process becomes at least somewhat easier.
In future articles, we will search specifically for parents and each of your four grandparents. In this article, I’ll review each of the diagnostic tools and techniques you can use to determine if you have endogamy, and perhaps pinpoint the source.
The Challenge
People with endogamous heritage are related in multiple, unknown ways, over many generations. They may also be related in known ways in recent generations.
If both of your parents share the SAME endogamous culture or group of relatives:
- You may have significantly more autosomal DNA matches than people without endogamy, unless that group of people is under-sampled. Jewish people have significantly more matches, but Native people have fewer due to under-sampling.
- You may experience a higher-than-normal cM (centiMorgan) total for estimated relationships, especially more distant relationships, 3C and beyond.
- You will have many matches related to you on both your maternal and paternal sides.
- Parts of your autosomal DNA will be the same on both your mother’s and father’s sides, meaning your DNA will be fully identical in some locations. (I’ll explain more in a minute.)
If either (or both) of your parents are from an endogamous population, you:
- Will, in some cases, carry identifying Y and mitochondrial DNA that points to a specific endogamous group. This is true for Native people, can be true for Jewish people and Pacific Islanders, but is not true for Anabaptist people.
One Size Does NOT Fit All
Please note that there is no “one size fits all.”
Each or any of these tools may provide relevant hints, depending on:
- Your heritage
- How many other people have tested from the relevant population group
- How many close or distant relatives have tested
- If your parents share the same heritage
- Your unique DNA inheritance pattern
- If your parents, individually, were fully endogamous or only partly endogamous, and how far back generationally that endogamy occurred
For example, in my own genealogy, my maternal grandmother’s father was Acadian on his father’s side. While I’m not fully endogamous, I have significantly more matches through that line proportionally than on my other lines.
I have Brethren endogamy on my mother’s side via her paternal grandmother.

Endogamous ancestors are shown with red stars on my mother’s pedigree chart, above. However, please note that her maternal and paternal endogamous ancestors are not from the same endogamous population.
However, I STILL have fewer matches on my mother’s side in total than on my father’s side because my mother has recent Dutch and recent German immigrants which reduces her total number of matches. Neither of those lines have had as much time to produce descendants in the US, and Europe is under-sampled when compared with the US where more people tend to take DNA tests because they are searching for where they came from.
My father’s ancestors have been in the US since it was a British Colony, and I have many more cousins who have tested on his side than mother’s.
If you looked at my pedigree chart and thought to yourself, “that’s messy,” you’d be right.
The “endogamy means more matches” axiom does not hold true for me, comparatively, between my parents – in part because my mother’s German and Dutch lines are such recent immigrants.
The number of matches alone isn’t going to tell this story.
We are going to need to look at several pieces and parts for more information. Let’s start with ethnicity.
Ethnicity and Populations
Ethnicity can be a double-edged sword. It can tell you exactly nothing you couldn’t discern by looking in the mirror, or, conversely, it can be a wealth of information.
Ethnicity reveals the parts of the world where your ancestors originated. When searching for recent ancestors, you’re most interested in majority ethnicity, meaning the 50% of your DNA that you received from each of your parents.
Ethnicity results at each vendor are easy to find and relatively easy to understand.

This individual at FamilyTreeDNA is 100% Ashkenazi Jewish.
If they were 50% Jewish, we could then estimate, and that’s an important word, that either one of their parents was fully Jewish, and not the other, or that two of their grandparents were Jewish, although not necessarily on the same side.
On the other hand, my mother’s ethnicity, shown below, has nothing remarkable that would point to any majority endogamous population, yet she has two.

The only hint of endogamy from ethnicity would be her ~1% Americas, and that isn’t relevant for finding close relatives. However, minority ancestry is very relevant for identifying Native ancestors, which I wrote about, here.
You can correlate or track your ethnicity segments to specific ancestors, which I discussed in the article, Native American & Minority Ancestors Identified Using DNAPainter Plus Ethnicity Segments, here.

Since I wrote that article, FamilyTreeDNA has added the feature of ethnicity or population Chromosome Painting, based on where each of your populations fall on your chromosomes.

In this example on chromosome 1, I have European ancestry (blue,) except for the pink Native segment, which occurs on the following segment in the same location on my mother’s chromosome 1 as well.

Both 23andMe, and FamilyTreeDNA provide chromosome painting AND the associated segment information so you can identify the relevant ancestors.
Ancestry is in the process of rolling out an ethnicity painting feature, BUT, it has no segment or associated matching information. While it’s interesting eye candy, it’s not terribly useful beyond the ethnicity information that Ancestry already provides. However, Jonny Perl at DNAPainter has devised a way to estimate Ancestry’s start and stop locations, here. Way to go Jonny!
Now all you need to do is convince your Ancestry matches to upload their DNA file to one of the three databases, FamilyTreeDNA, MyHeritage, and GEDMatch, that accept transfers, aka uploads. This allows matching with segment data so that you can identify who matches you on that segment, track your ancestors, and paint your ancestral segments at DNAPainter.
I provided step-by-step instructions, here, for downloading your raw DNA file from each vendor in order to upload the file to another vendor.
Ethnicity Sides
Three of the four DNA testing vendors, 23andMe, FamilyTreeDNA, and recently, Ancestry, attempt to phase your ethnicity DNA, meaning to assign it to one parental “side” or the other – both in total and on each chromosome.

Here’s Ancestry’s SideView, where your DNA is estimated to belong to parent 1 and parent 2. I detailed how to determine which side is which, here, and while that article was written specifically pertaining to Ancestry’s SideView, the technique is relevant for all the vendors who attempt to divide your DNA into parents, a technique known as phasing.
I say “attempt” because phasing may or may not be accurate, meaning the top chromosome may not always be parent 1, and the bottom chromosome may not always be chromosome 2.
Here’s an example at 23andMe.

See the two yellow segments. They are both assigned as Native. I happen to know one is from the mother and one is from the father, yet they are both displayed on the “top” chromosome, which one would interpret to be the same parent.
I am absolutely positive this is not the case because this is a close family member, and I have the DNA of the parent who contributed the Native segment on chromosome 1, on the top chromosome. That parent does not have a Native segment on chromosome 2 to contribute. So that Native segment had to be contributed by the other parent, but it’s also shown on the top chromosome.

The DNA segments circled in purple belong together on the same “side” and were contributed to the tester by the same parent. The Native segment on chromosome 2 abuts a purple African segment, suggesting perhaps that the ancestor who contributed that segment was mixed between those ethnicities. In the US, that suggests enslavement.
The other African segments, circled, are shown on the second chromosome in each pair.
To be clear, parent 1 is not assigned by the vendors to either mother or father and will differ by person. Your parent 1, or the parent on the top chromosome may be your mother and another person’s parent 1 may be their father.
As shown in this example, parents can vary by chromosome, a phenomenon known as “strand swap.” Occasionally, the DNA can even be swapped within a chromosome assignment.
You can, however, get an idea of the division of your DNA at any specific location. As shown above, you can only have a maximum of two populations of DNA on any one chromosome location.
In our example above, this person’s majority ancestry is European (blue.) On each chromosome where we find a minority segment, the opposite chromosome in the same location is European, meaning blue.
Let’s look at another example.
At FamilyTreeDNA, the person whose ethnicity painting is shown below has a Native American (pink) ancestor on their father’s side. FamilyTreeDNA has correctly phased or identified their Native segments as all belonging to the second chromosome in each pair.

Looking at chromosome 18, for example, most of their father’s chromosome is Native American (pink). The other parent’s chromosome is European (dark blue) at those same locations.
If one of the parents was of one ethnicity, and the other parent is a completely different ethnicity, then one bar of each chromosome would be all pink, for example, and one would be entirely blue, representing the other ethnicity.
Phasing ethnicity or populations to maternal and paternal sides is not foolproof, and each chromosome is phased individually.
Ethnicity can, in some cases, give you a really good idea of what you’re dealing with in terms of heritage and endogamy.
If someone had an Ashkenazi Jewish father and European mother, for example, one copy of each chromosome would be yellow (Ashkenazi Jewish), and one would be blue (European.)
However, if each of their parents were half European Jewish and half European (not Jewish), then their different colored segments would be scattered across their entire set of chromosomes.

In this case, both of the tester’s parents are mixed – European Jewish (green) and Western Europe (blue.) We know both parents are admixed from the same two populations because in some locations, both parents contributed blue (Western Europe), and in other locations, both contributed Jewish (green) segments.
Both MyHeritage and Ancestry provide a secondary tool that’s connected to ethnicity, but different and generally in more recent times.
Ancestry’s DNA Communities
While your ethnicity may not point to anything terribly exciting in terms of endogamy, Genetic Communities might. Ancestry says that a DNA Community is a group of people who share DNA because their relatives recently lived in the same place at the same time, and that communities are much smaller than ethnicity regions and reach back only about 50-300 years.

Based on the ancestors’ locations in the trees of me and my matches, Ancestry has determined that I’m connected to two communities. In my case, the blue group is clearly my father’s line. The orange group could be either parent, or even a combination of both.
My endogamous Brethren could be showing up in Maryland, Pennsylvania, and Ohio, but it’s uncertain, in part, because my father’s ancestral lines are found in Virginia, West Virginia, and Maryland too.
These aren’t useful for me, but they may be more useful for fully endogamous people, especially in conjunction with ethnicity.
My Acadian cousin’s European ethnicity isn’t informative.

However, viewing his DNA Communities puts his French heritage into perspective, especially combined with his match surnames.

I wrote about DNA Communities when it was introduced with the name Genetic Communities, here.
MyHeritage’s Genetic Groups
MyHeritage also provides a similar feature that shows where my matches’ ancestors lived in the same locations as mine.

One difference, though, is that testers can adjust their ethnicity results confidence level from high, above, to low, below where one of my Genetic Groups overlaps my ethnicity in the Netherlands.

You can also sort your matches by Genetic Groups.

The results show you not only who is in the group, but how many of your matches are in that group too, which provides perspective.

I wrote about Genetic Groups, here.
Next, let’s look at how endogamy affects your matches.
Matches
The number of matches that a person has who is from an entirely endogamous community and a person with no endogamy may be quite different.

FamilyTreeDNA provides a Family Matching feature that triangulates your matches and assigns them to your paternal or maternal side by using known matches that you have linked to their profile cards in your tree. You must link people for the Family Matching feature known as “bucketing” to be enabled.
The people you link are then processed for shared matches on the same chromosome segment(s). Triangulated individuals are then deposited in your maternal, paternal, and both buckets.

Obviously, your two parents are the best people to link, but if they haven’t tested (or uploaded their DNA file from another vendor) and you have other known relatives, link them using the Family Tree tab at the top of your personal page.

I uploaded my Ancestry V4 kit to use as an example for linking. Let’s pretend that’s my sister. If I had not already linked my Ancestry V4 kit to “my sister’s” profile card, I’d want to do that and link other known individuals the same way. Just drag and drop the match to the correct profile card.
Note that a full or half sibling will be listed as such at FamilyTreeDNA, but an identical twin will show as a potential parent/child match to you. You’re much more likely to find a parent than an identical twin, but just be aware.
I’ve created a table of FamilyTreeDNA bucketed match results, by category, comparing the number of matches in endogamous categories with non-endogamous.
|
Total Matches |
Maternal Matches |
Paternal Matches |
Both |
% Both |
% DNA Unassigned |
| 100% Jewish |
34,637 |
11,329 |
10,416 |
4,806 |
13.9 |
23.3 |
| 100% Jewish |
32,973 |
10,700 |
9,858 |
4,606 |
14 |
23.7 |
| 100% Jewish |
32,255 |
9,060 |
10,970 |
3,892 |
12 |
25.8 |
| 75% Jewish |
24,232 |
11,846 |
Only mother linked |
Only mother linked |
Only mother linked |
|
| 100% Acadian |
8093 |
3826 |
2299 |
1062 |
13 |
11 |
| 100% Acadian |
7828 |
3763 |
1825 |
923 |
11.8 |
17 |
| Not Endogamous |
6760 |
3845 |
1909 |
13 |
0.19 |
14.5 |
| Not Endogamous |
7723 |
1470 |
3317 |
6 |
0.08 |
38 |
| 100% Native American |
1,115 |
Unlinked |
Unlinked |
Unlinked |
|
|
| 100% Native American |
885 |
290 |
Unknown |
Can’t calculate without at least one link on both sides |
|
|
The 100% Jewish, Acadian, and Not Endogamous testers both have linked their parents, so their matches, if valid (meaning not identical by chance, which I discussed here,) will match them plus one or the other parent.
One person is 75% Jewish and has only linked their Jewish mother.
The Native people have not tested their parents, and the first Native person has not linked anyone in their tree. The second Native person has only linked a few maternal matches, but their mother has not tested. They are seeking their father.
It’s very difficult to find people who are fully Native as testers. Furthermore, Native people are under-sampled. If anyone knows of fully Native (or other endogamous) people who have tested and linked their parents or known relatives in their trees, and will allow me to use their total match numbers anonymously, please let me know.
As you can see, Jewish, Acadian, and Native people are 100% endogamous, but many more Jewish people than Native people have tested, so you CAN’T judge endogamy by the total number of matches alone.
In fact, in order:
- Fully Jewish testers have about 4-5 times as many matches as the Acadian and Non-endogamous testers
- Acadian and Non-endogamous testers have about 5-6 times as many matches as the Native American testers
- Fully Jewish people have about 30 times more matches than the Native American testers
If a person’s endogamy with a particular population is only on their maternal or paternal side, they won’t have a significant number of people related to both sides, meaning few people will fall into the “Both” bucket. People that will always be found in the ”Both” bucket are full siblings and their descendants, along with descendants of the tester, assuming their match is linked to their profiles in the tester’s tree.
In the case of our Jewish testers, you can easily see that the “Both” bucket is very high. The Acadians are also higher than one would reasonably expect without endogamy. A non-endogamous person might have a few matches on both sides, assuming the parents are not related to each other.
A high number of “Both” matches is a very good indicator of endogamy within the same population on both parents’ sides.
The percentage of people who are assigned to the “Both” bucket is between 11% and 14% in the endogamous groups, and less than 1% in the non-endogamous group, so statistically not relevant.
As demonstrated by the Native people compared to the Jewish testers, the total number of matches can be deceiving.
However, being related to both parents, as indicated by the “Both” bucket, unless you have pedigree collapse, is a good indicator of endogamy.
Of course, if you don’t know who your relatives are, you can’t link them in your tree, so this type of “hunt” won’t generally help people seeking their close family members.
However, you may notice that you’re matching people PLUS both of their parents. If that’s the case, start asking questions of those matches about their heritage.
A very high number of total matches, as compared to non-endogamous people, combined with some other hints might well point to Jewish heritage.
I included the % DNA Unassigned category because this category, when both parents are linked, is the percentage of matches by chance, meaning the match doesn’t match either of the tester’s parents. All of the people with people listed in “Both” categories have linked both of their parents, not just maternal and paternal relatives.
Matching Location at MyHeritage
MyHeritage provides a matching function by location. Please note that it’s the location of the tester, but that may still be quite useful.

The locations are shown in the most-matches to least-matches order. Clicking on the location shows the people who match you who are from that location. This would be the most useful in situations where recent immigration has occurred. In my case, my great-grandfather from the Netherlands arrived in the 1860s, and my German ancestors arrived in the 1850s. Neither of those groups are endogamous, though, unless it would be on a village level.
AutoClusters
Let’s shift to Genetic Affairs, a third-party tool available to everyone.
Using their AutoCluster function, Genetic Affairs clusters your matches together who match both each other and you.

This is an example of the first few clusters in my AutoCluster. You can see that I have several colored clusters of various sizes, but none are huge.
Compare that to the following endogamous cluster, sample courtesy of EJ Blom at Genetic Affairs.

If your AutoCluster at Genetic Affairs looks something like this, a huge orange blob in the upper left hand corner, you’re dealing with endogamy.
Please also note that the size of your cluster is also a function of both the number of testers and the match threshold you select. I always begin by using the defaults. I wrote about using Genetic Affairs, here.
If you tested at or transferred to MyHeritage, they too license AutoClusters, but have optimized the algorithm to tease out endogamous matches so that their Jewish customers, in particular, don’t wind up with a huge orange block of interrelated people.
You won’t see the “endogamy signature” huge cluster in the corner, so you’re less likely to be able to discern endogamy from a MyHeritage cluster alone.

The commonality between these Jewish clusters at MyHeritage is that they all tend to be rather uniform in size and small, with lots of grey connecting almost all the blocks.
Grey cells indicate people who match people in two colored groups. In other words, there is often no clear division in clusters between the mother’s side and the father’s side in Jewish clusters.
In non-endogamous situations, even if you can’t identify the parents, the clusters should still fall into two sides, meaning a group of clusters for each parent’s side that are not related to each other.
You can read more about Genetic Affairs clusters and their tools, here. DNAGedcom.com also provides a clustering tool.
Endogamous Relationships
Endogamous estimated relationships are sometimes high. Please note the word, “sometimes.”
Using the Shared cM Project tool relationship chart, here, at DNAPainter, people with heavy endogamy will discover that estimated relationships MAY be on the high side, or the relationships may, perhaps, be estimated too “close” in time. That’s especially true for more distant relationships, but surprisingly, it’s not always true. The randomness of inheritance still comes into play, and so do potential unknown relatives. Hence, the words “may” are bolded and underscored.
Unfortunately, it’s often stated as “conventional wisdom” that Jewish matches are “always” high, and first cousins appear as siblings. Let’s see what the actual data says.

At DNAPainter, you can either enter the amount of shared DNA (cM), or the percent of shared DNA, or just use the chart provided.

I’ve assembled a compilation of close relationships in kits that I have access to or from people who were generous enough to share their results for this article.
I’ve used Jewish results, which is a highly endogamous population, compared with non-endogamous testers.
The “Jewish Actual” column reports the total amount of shared DNA with that person. In other words, someone to their grandparent. The Average Range is the average plus the range from DNAPainter. The Percent Difference is the % difference between the actual number and the DNAPainter average.
You’ll see fully Jewish testers, at left, matching with their family members, and a Non-endogamous person, at right, matching with their same relative.
| Relationship |
Jewish Actual |
Percent Difference than Average |
Average -Range |
Non-endogamous Actual |
Percent Difference than Average |
| Grandparent |
2141 |
22 |
1754 (984-2482) |
1742 |
<1 lower |
| Grandparent |
1902 |
8.5 |
1754 (984-2482) |
1973 |
12 |
| Sibling |
3039 |
16 |
2613 (1613-3488) |
2515 |
3.5 lower |
| Sibling |
2724 |
4 |
2613 (1613-3488) |
2761 |
5.5 |
| Half-Sibling |
2184 |
24 |
1759 (1160-2436) |
2127 |
21 |
| Half-Sibling |
2128 |
21 |
1759 (1160-2436) |
2352 |
34 |
| Aunt/Uncle |
2066 |
18.5 |
1741 (1201-2282) |
1849 |
6 |
| Aunt/Uncle |
2031 |
16.5 |
1741 (1201-2282) |
2097 |
20 |
| 1C |
1119 |
29 |
866 (396-1397) |
959 |
11 |
| 1C |
909 |
5 |
866 (396-1397) |
789 |
9 lower |
| 1C1R |
514 |
19 |
433 (102-980) |
467 |
8 |
| 1C1R |
459 |
6 |
433 (102-980) |
395 |
9 lower |
These totals are from FamilyTreeDNA except one from GEDMatch (one Jewish Half-sibling).
Totals may vary by vendor, even when matching with the same person. 23andMe includes the X segments in the total cMs and also counts fully identical segments twice. MyHeritage imputation seems to err on the generous side.
However, in these dozen examples:
- You can see that the Jewish actual amount of DNA shared is always more than the average in the estimate.
- The red means the overage is more than 100 cM larger.
- The percentage difference is probably more meaningful because 100 cM is a smaller percentage of a 1754 grandparent connection than compared to a 433 cM 1C1R.
However, you can’t tell anything about endogamy by just looking at any one sample, because:
- Some of the Non-Endogamous matches are high too. That’s just the way of random inheritance.
- All of the actual Jewish match numbers are within the published ranges, but on the high side.
Furthermore, it can get more complex.
Half Endogamous
I requested assistance from Jewish genealogy researchers, and a lovely lady, Sharon, reached out, compiled her segment information, and shared it with me, granting permission to share with you. A HUGE thank you to Sharon!
Sharon is half-Jewish via one parent, and her half-sibling is fully Jewish. Their half-sibling match to each other at Ancestry is 1756 cM with a longest segment of 164 cM.
How does Jewish matching vary if you’re half-Jewish versus fully Jewish? Let’s look at 21 people who match both Sharon and her fully Jewish half-sibling.

Sharon shared the differences in 21 known Jewish matches with her and her half-sibling. I’ve added the Relationship Estimate Range from DNAPainter and colorized the highest of the two matches in yellow. Bolding in the total cM column shows a value above the average range for that relationship.
Total Matching cMs is on the left, with Longest Segment on the right.
While this is clearly not a scientific study, it is a representative sample.
The fully Jewish sibling carries more Jewish DNA, which is available for other Jewish matches to match as a function of endogamy (identical by chance/population), so I would have expected the fully Jewish sibling to match most if not all Jewish testers at a higher level than the half-Jewish sibling.
However, that’s not universally what we see.
The fully Jewish sibling is not always the sibling with the highest number of matches to the other Jewish testers, although the half-Jewish tester has the larger “Longest Segment” more often than not.
Approximately two-thirds of the time (13/21), the fully Jewish person does have a higher total matching cM, but about one-third of the time (8/21), the half-Jewish sibling has a higher matching cM.
About one-fourth of the time (5/21), the fully Jewish sibling has the longest matching segment, and about two-thirds of the time (13/21), the half-Jewish sibling does. In three cases, or about 14% of the time, the longest segment is equal which may indicate that it’s the same segment.
Because of endogamy, Jewish matches are more likely to have:
- Larger than average total cM for the specific relationship
- More and smaller matching segments
However, as we have seen, neither of those are definitive, nor always true. Jewish matches and relationships are not always overestimated.
Ancestry and Timber
Please note that Ancestry downweights some matches by removing some segments using their Timber algorithm. Based on my matches and other accounts that I manage, Ancestry does not downweight in the 2-3rd cousin category, which is 90 cM and above, but they do begin downweighting in the 3-4th cousin category, below 90 cM, where my “Extended Family” category begins.
If you’ve tested at Ancestry, you can check for yourself.

By clicking on the amount of DNA you share with your match on your match list at Ancestry, shown above, you will be taken to another page where you will be able to view the unweighted shared DNA with that match, meaning the amount of DNA shared before the downweighting and removal of some segments, shown below.

Given the downweighting, and the information in the spreadsheet provided by Sharon, it doesn’t appear that any of those matches would have been in a category to be downweighted.
Therefore, for these and other close matches, Timber wouldn’t be a factor, but would potentially be in more distant matches.
Endogamous Segments
Endogamous matches tend to have smaller and more segments. Small amounts of matching DNA tend to skew the total DNA cM upwards.
How and why does this happen?
Ancestral DNA from further back in time tends to be broken into smaller segments.
Sometimes, especially in endogamous situations, two smaller segments, at one time separated from each other, manage to join back together again and form a match, but the match is only due to ancestral segments – not because of a recent ancestor.
Please note that different vendors have different minimum matching cM thresholds, so smaller matches may not be available at all vendors. Remember that factors like Timber and imputation can affect matching as well.
Let’s take a look at an example. I’ve created a chart where two ancestors have their blue and pink DNA broken into 4 cM segments.

They have children, a blue child and a pink child, and the two children, shown above, each inherited the same blue 4 cM segment and the same pink 4 cM segment from their respective parents. The other unlabeled pink and blue segments are not inherited by these two children, so those unlabeled segments are irrelevant in this example.
The parents may have had other children who inherited those same 4 cM labeled pink and blue segments as well, and if not, the parents’ siblings were probably passing at least some of the same DNA down to their descendants too.
The blue and pink children had children, and their children had children – for several generations.
Time passed, and their descendants became an endogamous community. Those pink and blue 4 cM segments may at some time be lost during recombination in the descendants of each of their children, shown by “Lost pink” and “Lost blue.”
However, because there is only a very limited amount of DNA within the endogamous community, their descendants may regain those same segments again from their “other parent” during recombination, downstream.
In each generation, the DNA of the descendant carrying the original blue or pink DNA segment is recombined with their partner. Given that the partners are both members of the same endogamous community, the two people may have the same pink and/or blue DNA segments. If one parent doesn’t carry the pink 4 cM segment, for example, their offspring may receive that ancestral pink segment from the other parent.
They could potentially, and sometimes do, receive that ancestral segment from both parents.
In our example, the descendants of the blue child, at left, lost the pink 4 cM segment in generation 3, but a few generations later, in generation 11, that descendant child inherited that same pink 4 cM segment from their other parent. Therefore, both the 4 cM blue and 4 cM pink segments are now available to be inherited by the descendants in that line. I’ve shown the opposite scenario in the generational inheritance at right where the blue segment is lost and regained.
Once rejoined, that pink and blue segment can be passed along together for generations.
The important part, though, is that once those two segments butt up against each other again during recombination, they aren’t just two separate 4 cM segments, but one segment that is 8 cM long – that is now equal to or above the vendors’ matching threshold.
This is why people descended from endogamous populations often have the following matching characteristics:
- More matches
- Many smaller segment matches
- Their total cM is often broken into more, smaller segments
What does more, smaller segments, look like, exactly?
More, Smaller Segments
All of our vendors except Ancestry have a chromosome browser for their customers to compare their DNA to that of their matches visually.
Let’s take a look at some examples of what endogamous and non-endogamous matches look like.

For example, here’s a screen shot of a random Jewish second cousin match – 298 cM total, divided into 12 segments, with a longest segment of 58 cM,

A second Jewish 2C with 323 cM total, across 19 segments, with a 69 cM longest block.

A fully Acadian 2C match with 600 cM total, across 27 segments, with a longest segment of 69 cM.

A second Acadian 2C with 332 cM total, across 20 segments, with a longest segment of 42 cM.

Next, a non-endogamous 2C match with 217 cM, across 7 segments, with a longest segment of 72 cM.

Here’s another non-endogamous 2C example, with 169 shared cM, across 6 segments, with a longest segment of 70 cM.
Here’s the second cousin data in a summary table. The take-away from this is the proportion of total segments
| Tester Population |
Total cM |
Longest Block |
Total Segments |
| Jewish 2C |
298 |
58 |
12 |
| Jewish 2C |
323 |
69 |
19 |
| Acadian 2C |
600 |
69 |
27 |
| Acadian 2C |
332 |
42 |
20 |
| Non-endogamous 2C |
217 |
72 |
7 |
| Non-endogamous 2C |
169 |
70 |
6 |
You can see more examples and comparisons between Native American, Jewish and non-endogamous DNA individuals in the article, Concepts – Endogamy and DNA Segments.
I suspect that a savvy mathematician could predict endogamy based on longest block and total segment information.
Lara Diamond, a mathematician, who writes at Lara’s Jewnealogy might be up for this challenge. She just published compiled matching and segment information in her Ashkenazic Shared DNA Survey Results for those who are interested. You can also contribute to Laura’s data, here.
Endogamy, Segments, and Distant Relationships
While not relevant to searching for close relatives, heavily endogamous matches 3C and more distant, to quote one of my Jewish friends, “dissolve into a quagmire of endogamy and are exceedingly difficult to unravel.”
In my own Acadian endogamous line, I often simply have to label them “Acadian” because the DNA tracks back to so many ancestors in different lines. In other words, I can’t tell which ancestor the match is actually pointing to because the same DNA segments or segments is/are carried by several ancestors and their descendants due to founder effect.
The difference with the Acadians is that we can actually identify many or most of them, at least at some point in time. As my cousin, Paul LeBlanc, once said, if you’re related to one Acadian, you’re related to all Acadians. Then he proceeded to tell me that he and I are related 137 different ways. My head hurts!
It’s no wonder that endogamy is incredibly difficult beyond the first few generations when it turns into something like multi-colored jello soup.
“Are Your Parents Related?” Tool
There’s another tool that you can utilize to determine if your parents are related to each other.
To determine if your parents are related to each other, you need to know about ROH, or Runs of Homozygosity (ROH).
ROH means that the DNA on both strands or copies of the same chromosome is identical.
For a few locations in a row, ROH can easily happen just by chance, but the longer the segment, the less likely that commonality occurs simply by chance.
The good news is that you don’t need to know the identity of either of your parents. You don’t need either of your parent’s DNA tests – just your own. You’ll need to upload your DNA file to GEDmatch, which is free.
Click on “Are your parents related?”

GEDMatch analyzes your DNA to see if any of your DNA, above a reasonable matching threshold, is identical on both strands, indicating that you inherited the exact same DNA from both of your parents.
A legitimate match, meaning one that’s not by chance, will include many contiguous matching locations, generally a minimum of 500 SNPs or locations in a row. GEDmatch’s minimum threshold for identifying identical ancestral DNA (ROH) is 200 cM.

Here’s my result, including the graphic for the first two chromosomes. Notice the tiny green bars that show identical by chance tiny sliver segments.

I have no significant identical DNA, meaning my parents are not related to each other.
Next, let’s look at an endogamous example where there are small, completely identical segments across a person’s chromosome

This person’s Acadian parents are related to each other, but distantly.
Next, let’s look at a Jewish person’s results.

You’ll notice larger green matching ROH, but not over 200 contiguous SNPs and 7 cM.
GEDMatch reports that this Jewish person’s parents are probably not related within recent generations, but it’s clear that they do share DNA in common.
People whose parents are distantly related have relatively small, scattered matching segments. However, if you’re seeing larger ROH segments that would be large enough to match in a genealogical setting, meaning multiple greater than 7 cM and 500 SNPs,, you may be dealing with a different type of situation where cousins have married in recent generations. The larger the matching segments, generally, the closer in time.
Blogger Kitty Cooper wrote an article, here, about discovering that your parents are related at the first cousin level, and what their GEDMatch “Are Your Parents Related” results look like.
Let’s look for more clues.
Surnames
There MAY be an endogamy clue in the surnames of the people you match.
Viewing surnames is easier if you download your match list, which you can do at every vendor except Ancestry. I’m not referring to the segment data, but the information about your matches themselves.
I provided instructions in the recent article, How to Download Your DNA Match Lists and Segment Files, here.
If you suspect endogamy for any reason, look at your closest matches and see if there is a discernable trend in the surnames, or locations, or any commonality between your matches to each other.
For example, Jewish, Acadian, and Native surnames may be recognizable, as may locations.
You can evaluate in either or both of two ways:
- The surnames of your closest matches. Closest matches listed first will be your default match order.
- Your most frequently occurring surnames, minus extremely common names like Smith, Jones, etc., unless they are also in your closest matches. To utilize this type of matching, sort the spreadsheet in surname order and then scan or count the number of people with each surname.
Here are some examples from our testers.
Jewish – Closest surname matches.
- Roth
- Weiss
- Goldman
- Schonwald
- Levi
- Cohen
- Slavin
- Goodman
- Sender
- Trebatch
Acadian – Closest surname matches.
- Bergeron
- Hebert
- Bergeron
- Marcum
- Muise
- Legere
- Gaudet
- Perry
- Verlander
- Trombley
Native American – Closest surname matches.
- Ortega
- Begay
- Valentine
- Hayes
- Montoya
- Sun Bear
- Martin
- Tsosie
- Chiquito
- Yazzie
You may recognize these categories of surnames immediately.
If not, Google is your friend. Eliminate common surnames, then Google for a few together at a time and see what emerges.
The most unusual surnames are likely your best bets.
Projects
Another way to get some idea of what groups people with these surnames might belong to is to enter the surname in the FamilyTreeDNA surname search.
Go to the main FamilyTreeDNA page, but DO NOT sign on.
Scroll down until you see this image.

Type the surname into the search box. You’ll see how many people have tested with that surname, along with projects where project administrators have included that surname indicating that the project may be of interest to at least some people with that surname.

Here’s a portion of the project list for Cohen, a traditional Jewish surname.

These results are for Muise, an Acadian surname.
Clicking through to relevant surname projects, and potentially contacting the volunteer project administrator can go a very long way in helping you gather and sift information. Clearly, they have an interest in this topic.

For example, here’s the Muise surname in the Acadian AmerIndian project. Two great hints here – Acadian heritage and Halifax, Nova Scotia.
Repeat for the balance of surnames on your list to look for commonalities, including locations on the public project pages.
Locations
Some of the vendor match files include location information. Each person on your match list will have the opportunity at the vendor where they tested to include location information in a variety of ways, either for their ancestors or themselves.
Where possible, it’s easiest to sort or scan the download file for this type of information.
Ancestry does not provide or facilitate a match list, but you can still create your own for your closest 20 or 30 matches in a spreadsheet.
MyHeritage provides common surname and ancestral location information for every match. How cool is that!
Y DNA, Mitochondrial DNA, and Endogamy
Haplogroups for both Y and mitochondrial DNA can indicate and sometimes confirm endogamy. In other cases, the haplogroup won’t help, but the matches and their location information just might.
FamilyTreeDNA is the only vendor that provides Y DNA and mitochondrial DNA tests that include highly granular haplogroups along with matches and additional tools.
23andMe provides high-level haplogroups which may or may not be adequate to pinpoint a haplogroup that indicates endogamy.

Of course, only males carry Y DNA that tracks to the direct paternal (surname) line, but everyone carries their mother’s mitochondrial DNA that represents their mother’s mother’s mother’s, or direct matrilineal line.
Some haplogroups are known to be closely associated with particular ethnicities or populations, like Native Americans, Pacific Islanders, and some Jewish people.
Haplogroups reach back in time before genealogy and can give us a sense of community that’s not available by either looking in the mirror or through traditional records.

This Native American man is a member of high-level haplogroup Q-M242. However, some men who carry this haplogroup are not Native, but are of European or Middle Eastern origin.
I entered the haplogroup in the FamilyTreeDNA Discover tool, which I wrote about, here.

Checking the information about this haplogroup reveals that their common ancestor descended from an Asian man about 30,000 years ago.

The migration path in the Americans explains why this person would have an endogamous heritage.

Our tester would receive a much more refined haplogroup if he upgraded to the Big Y test at FamilyTreeDNA, which would remove all doubt.
However, even without additional testing, information about his matches at FamilyTreeDNA may be very illuminating.

The Q-M242 Native man’s Y DNA matches men with more granular haplogroups, shown above, at left. On the Haplogroup Origins report, you can see that these people have all selected the “US (Native American)” country option.
Another useful tool would be to check the public Y haplotree, here, and the public mitochondrial tree here, for self-reported ancestor location information for a specific haplogroup.

Here’s an example of mitochondrial haplogroup A2 and a few subclades on the public mitochondrial tree. You can see that the haplogroup is found in Mexico, the US (Native,) Canada, and many additional Caribbean, South, and Central American countries.
Of course, Y DNA and mitochondrial DNA (mtDNA) tell a laser-focused story of one specific line, each. The great news, if you’re seeking information about your mother or father, the Y is your father’s direct paternal (surname) line, and mitochondrial is your mother’s direct matrilineal line.
Y and mitochondrial DNA results combined with ethnicity, autosomal matching, and the wide range of other tools that open doors, you will be able to reveal a great deal of information about whether you have endogamous heritage or not – and if so, from where.
I’ve provided a resource for stepping through and interpreting your Y DNA results, here, and mitochondrial DNA, here.
Discover for Y DNA Only
If you’re a female, you may feel left out of Y DNA testing and what it can tell you about your heritage. However, there’s a back door.
You can utilize the Y DNA haplogroups of your closest autosomal matches at both FamilyTreeDNA and 23andMe to reveal information
Haplogroup information is available in the download files for both vendors, in addition to the Family Finder table view, below, at FamilyTreeDNA, or on your individual matches profile cards at both 23andMe and FamilyTreeDNA.

You can enter any Y DNA haplogroup in the FamilyTreeDNA Discover tool, here.

You’ll be treated to:
- Your Haplogroup Story – how many testers have this haplogroup (so far), where the haplogroup is from, and the haplogroup’s age. In this case, the haplogroup was born in the Netherlands about 250 years ago, give or take 200 years. I know that it was 1806 or earlier based on the common ancestor of the men who tested.
- Country Frequency – heat map of where the haplogroup is found in the world.
- Notable Connections – famous and infamous (this haplogroup’s closest notable person is Leo Tolstoy).
- Migration Map – migration path out of Africa and through the rest of the world.
- Ancient Connections – ancient burials. His closest ancient match is from about 1000 years ago in Ukraine. Their shared ancestor lived about 2000 years ago.
- Suggested Projects – based on the surname, projects that other matches have joined, and haplogroups.
- Scientific Details – age estimates, confidence intervals, graphs, and the mutations that define this haplogroup.
I wrote about the Discover tool in the article, FamilyTreeDNA DISCOVER Launches – Including Y DNA Haplogroup Ages.
Endogamy Tools Summary Tables
Endogamy is a tough nut sometimes, especially if you’re starting from scratch. In order to make this topic a bit easier and to create a reference tool for you, I’ve created three summary tables.
- Various endogamy-related tools available at each vendor which will or may assist with evaluating endogamy
- Tools and their ability to detect endogamy in different groups
- Tools best suited to assist people seeking information about unknown parents or grandparents
Summary of Endogamy Tools by Vendor
Please note that GEDMatch is not a DNA testing vendor, but they accept uploads and do have some tools that the testing vendors do not.
| Tool |
23andMe |
Ancestry |
FamilyTreeDNA |
MyHeritage |
GEDMatch |
| Ethnicity |
Yes |
Yes |
Yes |
Yes |
Use the vendors |
| Ethnicity Painting |
Yes + segments |
Yes, limited |
Yes + segments |
Yes |
|
| Ethnicity Phasing |
Yes |
Partial |
Yes |
No |
|
| DNA Communities |
No |
Yes |
No |
No |
|
| Genetic Groups |
No |
No |
No |
Yes |
|
| Family Matching aka Bucketing |
No |
No |
Yes |
No |
|
| Chromosome Browser |
Yes |
No |
Yes |
Yes |
Yes |
| AutoClusters |
Through Genetic Affairs |
No |
Through Genetic Affairs |
Yes, included |
Yes, with subscription |
| Match List Download |
Yes, restricted # of matches |
No |
Yes |
Yes |
Yes |
| Projects |
No |
No |
Yes |
No |
|
| Y DNA |
High-level haplogroup only |
No |
Yes, full haplogroup with Big Y, matching, tools, Discover |
No |
|
| Mitochondrial DNA |
High-level haplogroup only |
No |
Yes, full haplogroup with mtFull, matching, tools |
No |
|
| Public Y Tree |
No |
No |
Yes |
No |
|
| Public Mito Tree |
No |
No |
Yes |
No |
|
| Discover Y DNA – public |
No |
No |
Yes |
No |
|
| ROH |
No |
No |
No |
No |
Yes |
Summary of Endogamous Populations Identified by Each Tool
The following chart provides a guideline for which tools are useful for the following types of endogamous groups. Bolded tools require that both parents be descended from the same endogamous group, but several other tools give more definitive results with higher amounts of endogamy.
Y and mitochondrial DNA testing are not affected by admixture, autosomal DNA or anything from the “other” parent.
| Tool |
Jewish |
Acadian |
Anabaptist |
Native |
Other/General |
| Ethnicity |
Yes |
No |
No |
Yes |
Pacific Islander |
| Ethnicity Painting |
Yes |
No |
No |
Yes |
Pacific Islander |
| Ethnicity Phasing |
Yes, if different |
No |
No |
Yes, if different |
Pacific Islander, if different |
| DNA Communities |
Yes |
Possibly |
Possibly |
Yes |
Pacific Islander |
| Genetic Groups |
Yes |
Possibly |
Possibly |
Yes |
Pacific Islander |
| Family Matching aka Bucketing |
Yes |
Yes |
Possibly |
Yes |
Pacific Islander |
| Chromosome Browser |
Possibly |
Possibly |
Yes, once segments or ancestors identified |
Possibly |
Pacific Islander, possibly |
| Total Matches |
Yes, compared to non-endogamous |
No |
No |
No |
No, unknown |
| AutoClusters |
Yes |
Yes |
Uncertain, probably |
Yes |
Pacific Islander |
| Estimated Relationships High |
Not always |
Sometimes |
No |
Sometimes |
Uncertain, probably |
| Relationship Range High |
Possibly, sometimes |
Possibly |
Possibly |
Possibly |
Pacific Islander, possibly |
| More, Smaller Segments |
Yes |
Yes |
Probably |
Yes |
Pacific Islander, probably |
| Parents Related |
Some but minimal |
Possibly |
Uncertain |
Probably similar to Jewish |
Uncertain, Possibly |
| Surnames |
Probably |
Probably |
Probably Not |
Possibly |
Possibly |
| Locations |
Possibly |
Probably |
Probably Not |
Probably |
Probably Pacific Islander |
| Projects |
Probably |
Probably |
Possibly |
Possibly |
Probably Pacific Islander |
| Y DNA |
Yes, often |
Yes, often |
No |
Yes |
Pacific Islander |
| Mitochondrial DNA |
Yes, often |
Sometimes |
No |
Yes |
Pacific Islander |
| Y public tree |
Probably not alone |
No |
No |
Yes |
Pacific Islander |
| MtDNA public tree |
Probably not |
No |
No |
Yes |
Pacific Islander |
| Y DNA Discover |
Yes |
Possibly |
Probably not, maybe projects |
Yes |
Pacific Islander |
Summary of Endogamy Tools to Assist People Seeking Unknown Parents and Grandparents
This table provides a summary of when each of the various tools can be useful to:
- People seeking unknown close relatives
- People who already know who their close relatives are, but are seeking additional information or clues about their genealogy
I considered rating these on a 1 to 10 scale, but the relative usefulness of these tools is dependent on many factors, so different tools will be more or less useful to different people.
For example, ethnicity is very useful if someone is admixed from different populations, or even 100% of a specific endogamous population. It’s less useful if the tester is 100% European, regardless of whether they are seeking close relatives or not. Conversely, even “vanilla” ethnicity can be used to rule out majority or recent admixture with many populations.
| Tools |
Unknown Close Relative Seekers |
Known Close Relatives – Enhance Genealogy |
| Ethnicity |
Yes, to identify or rule out populations |
Yes |
| Ethnicity Painting |
Yes, possibly, depending on population |
Yes, possibly, depending on population |
| Ethnicity Phasing |
Yes, possibly, depending on population |
Yes, possibly, depending on population |
| DNA Communities |
Yes, possibly, depending on population |
Yes, possibly, depending on population |
| Genetic Groups |
Possibly, depending on population |
Possibly, depending on population |
| Family Matching aka Bucketing |
Not if parents are entirely unknown, but yes if one parent is known |
Yes |
| Chromosome Browser |
Unlikely |
Yes |
| AutoClusters |
Yes |
Yes, especially at MyHeritage if Jewish |
| Estimated Relationships High |
Not |
No |
| Relationship Range High |
Not reliably |
No |
| More, Smaller Segments |
Unlikely |
Unlikely other than confirmation |
| Match List Download |
Yes |
Yes |
| Surnames |
Yes |
Yes |
| Locations |
Yes |
Yes |
| Projects |
Yes |
Yes |
| Y DNA |
Yes, males only, direct paternal line, identifies surname lineage |
Yes, males only, direct paternal line, identifies and correctly places surname lineage |
| Mitochondrial DNA |
Yes, both sexes, direct matrilineal line only |
Yes, both sexes, direct matrilineal line only |
| Public Y Tree |
Yes for locations |
Yes for locations |
| Public Mito Tree |
Yes for locations |
Yes for locations |
| Discover Y DNA |
Yes, for heritage information |
Yes, for heritage information |
| Parents Related – ROH |
Possibly |
Less useful |
Acknowledgments
A HUGE thank you to several people who contributed images and information in order to provide accurate and expanded information on the topic of endogamy. Many did not want to be mentioned by name, but you know who you are!!!
If you have information to add, please post in the comments.
_____________________________________________________________
Follow DNAexplain on Facebook, here or follow me on Twitter, here.
Share the Love!
You’re always welcome to forward articles or links to friends and share on social media.
If you haven’t already subscribed (it’s free,) you can receive an email whenever I publish by clicking the “follow” button on the main blog page, here.
You Can Help Keep This Blog Free
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Uploads
Genealogy Products and Services
My Book
Genealogy Books
Genealogy Research