When I first began as a surname administrator for the Estes project, more than a decade ago, I wrote an “intro” basics document for anyone who might be interested in testing. This saved me from having to repeat myself again and again. I believe this is the 8th version of that document. Genetic genealogy keeps changing, for the better, with more tests and tools available, so more to explain.
DNA testing for genealogy didn’t exist a few years ago. In 1999, the first tests were performed for genetic genealogy and this wonderful tool which would revolutionize genealogy forever was born into the consumer marketplace from the halls of academia, thanks to one very persistent genealogist, Bennett Greenspan, now President of Family Tree DNA.
Initially we had more questions than answers. If it’s true that we have some amount of DNA from all of our ancestors, how can we tell which pieces are from which ancestor? How much can we learn from our DNA? Where did we come from both individually and as population subgroups? How can DNA help me knock down those genealogy brick walls?
In just a few short years, we have answers for most of these questions. However, in this still infant science we continue to learn every day. But before we discuss the answers, let’s talk for just a minute about how DNA works.
DNA – The Basics
Every human has 23 pairs of chromosomes (think of them as recipe books), which contain most of your DNA, functional units of which are known as genes (think of them as chapters). One chromosome of each pair comes from a person’s mother and the other from their father. Due to the mixing, called recombination, of DNA that occurs during meiosis prior to sperm and egg development, each chromosome in 22 of the 23 pairs, which are known as autosomes, has DNA (think of it as ingredients) from both the corresponding parent’s parents (and their ancestors before them).

Two portions of our DNA are not combined with that of the other parent. The 23rd chromosome, in the box above, determines the sex of the individual. Two X chromosomes produce a female and an X and a Y chromosome produce a male. Women do not have a Y chromosome (otherwise they would be males) so they cannot contribute a Y chromosome to male offspring. Given this scenario, males inherit their father’s Y chromosome unmixed with the mother’s DNA, and an X chromosome from their mother, unmixed with their father’s DNA.
This inheritance pattern is what makes it possible for us to use the Y chromosome to compare against other men of the same surname to see if they share a common ancestor, because if they do, their Y chromosome DNA will match, either exactly or nearly so, because it has been passed intact directly from those paternal ancestors.
Autosomal DNA, X chromosomal DNA and, in males, Y chromosomal DNA are all found in the nucleus of a cell. A fourth type of DNA call mitochondrial DNA, or mtDNA for short, resides within cells but outside the cell’s nucleus. Mitochondrial DNA packets are the cell’s powerhouse as they provide the entire body with energy.
For both genders, mitochondria DNA is inherited only from the mother. Men inherit their mother’s mtDNA, but do not pass it on to their offspring. Women have their mother’s mtDNA and pass it to both their female and male offspring. Given this scenario, women inherit their mother’s mtDNA unmixed with the father’s and pass it on generation to generation from female to female. This inheritance pattern is what makes it possible for us to compare our mitochondrial DNA with that of others to determine whether we share a common maternal ancestor.

Autosomal DNA, the rest of your DNA, those other 22 chromosomes that are not the X/Y chromosome and not the mitochondrial DNA, tends to be transferred in groupings, which ultimately give us traits like Mother’s blue eyes, Grandpa’s chin or Dad’s stocky build. Sometimes these inherited traits can be less positive, like deformities, diseases or tendencies like alcoholism. How this occurs and what genes or combinations of genes are responsible for transferring particular traits is still being deciphered.
Sometimes we inherit conflicting genes from our parents and the resolution of which trait is exhibited is called gene expression. For example, if you inherit a gene for blue eyes and brown eyes, you can’t have both, so the complex process of gene expression determines which color of eyes you will have. However, this type of genetics along with medical genetics does not concern us when we are using genetics for genealogy. Let’s focus initially on the unrecombined Y chromosomal DNA, called Y DNA for short, and mtDNA as genealogical tools.
How Can Unrecombined DNA Help Us With Genealogy?
I’m so glad you asked.
During normal cell combination, called meiosis, each ancestor’s autosomal DNA gets watered down or divided by roughly half with each generation, meaning each child gets half of the DNA carried by each parent.
However, that isn’t true of the Y DNA or mtDNA. In the following example of just 4 generations, we see that the Y DNA, the blue box on the left, is passed down the paternal line intact and the son has the exact same Y DNA as his paternal great-grandfather.
Similarly, the round red doughnut shaped O represents the mitochondrial DNA (mtDNA) and it is passed down the maternal side, so both the daughter and the son will have the exact same mtDNA as the maternal great-grandmother (but only the female child will pass it on).

The good news is that you may well have noticed that the surname is passed down the same blue paternal path, so if this is a Jones family, the Y DNA travels right along with the surname. How it can help us with genealogy now becomes obvious, because if we can test different male descendents who also bear the Jones surname, if they share a common ancestor somewhere in recent time (the last several hundred years), their DNA will match, or nearly so. Surname projects have been created by volunteer administrators at Family Tree DNA to facilitate coordination and comparison of individuals carrying the same or similar surnames.
Mitochondrial DNA (mtDNA) is useful as well, but not as easily for genealogical purposes since the maternal surname traditionally changes with each generation.
There have been several remarkable success stories using mitochondrial DNA, but they are typically more difficult to coordinate because of the challenges presented by the last name changes. Sometimes joining regional projects is more useful for finding mtDNA matches than joining surname projects. A case in point is the Cumberland Gap projects, both Y DNA and mtDNA, which have helped many people whose families lived in close proximity of the Cumberland Gap (at the intersection of Va., Tn. and Ky.) connect with their genetic cousins. What mtDNA as well as Y DNA testing can easily do for us is to confirm, or put to bed forever, rumors of Native American, European, African or Asian ancestry in that direct line.
What About Mutations?
Another really good question.
Y DNA testing actually tests either 12, 25, 37, 67 or 111 locations on the Y chromosome, depending on which test you select. What is actually reported at these locations is the number of exact repeats of that segment of DNA. Occasionally, either a segment is dropped or one is added. This is a normal process and typically affects nothing. However, for genealogy, these changes or mutations are wonderful, as the number of segments in a particular location will typically be the same from generation to generation. These mutations differentiate us and our families over time. Without mutations, all of our DNA would look exactly alike and there would be no genetic genealogy.
For mitochondrial DNA, you can test at the entry level, the intermediate “plus” level and at the full sequence level. If you think of the full sequence level, which tests the entire mitochondria, as a clock face, the entry level test tests from 5 till the hour to “noon” so from 11AM to 12 on the clock face. The second intermediate level tests from “noon” to 5 after, or 1PM. The full sequence level tests the entire clock face. Ultimately, if it’s matches you’re looking for, you’ll want the full sequence test to provide you with the best matches and the ones closest to you in time, plus it provides you with your full haplogroup, or clan, designation.
When a change, called a mutation, does occur at a particular location, it is then passed from father to son (or mother to daughter) and on down that line. That mutation, called a “line marker mutation” is then forever associated with that line of the family. If you test different males with the same surname, and they match except for only a couple of minor differences, you can be assured that they do in fact share a common ancestor in a genealogically relevant timeframe.
A father can potentially sire several sons, some with no mutations, and others with different mutations, as shown by the red mutation bar in the following illustration.

In the above example, John Patrick Kenney had two sons, one with no mutation and Paul Edward Kenney who had one mutation. All of the male descendents of Paul Edward Kenney have his mutation and a second mutation is added to this line at a new location in the generation above Stan Kenny.
John Patrick Kenney’s son who had no mutations sired a son Joseph Kenney, who had a mutation in yet a different location than either of the mutations in the Paul Edward Kenney line.
In the span of time between 1478 and 2004, this grouping of Kenney/Kenny families has accumulated 4 distinct lines as you can see across the bottom of the diagram, line 3 with no mutations, line 1 with 2 mutations, and two other lines with only one mutation each, but those mutations are not in the same location so they are easily differentiated in descendants testing today. These are called “line marker” mutations and allow testers to quickly and easily see which line of the Kenny family they descend from.
What Do the Results Look Like?
Y DNA results are reported in the following format at Family Tree DNA where locus means the location number, the DYS# means the name of that marker location, and the number of alleles means the number of repeats of DNA found in that location. This is a partial screen shot from the Family Tree DNA results page for a participant.

This is interesting, but the power of DNA testing isn’t in what your numbers alone look like, but in how they compare with others of similar surnames. So, you’re provided with a list of people that you match, along with access to their Gedcom file if they have uploaded one, most distant ancestor information, and most importantly, their e-mail address by clicking on the little envelope right after their name.

As a DNA Surname Project Administrator of several projects, I combine the groupings of participants into logical groupings based on their DNA patterns and their genealogy. Haplogroup projects are grouped by subgroup and mutations, and surname projects are grouped by matching family group.
The following table is an example from my Estes surname project which has very successfully identified the various sons of the immigrant ancestor, Abraham Estes born in 1647. Based on his descendent lines’ DNA, we have even successfully reconstructed what Abraham’s DNA looked like, shown in green, through a process called triangulation, so we have a firm basis for comparison, and everyone is compared to Abraham. Mutations are highlighted in yellow.
I have shown only an example of the full chart below. Moses through John R’s line does have line marker mutations on markers that are not shown here. Elisha’s line matches Abraham’s exactly. We have had 4 descendents test from various sons of Elisha and so far we have found no mutations.
 To form a baseline within a family, we generally test two individuals from two separate lines of the common ancestor, just in case an undocumented adoption has occurred. If these two individuals match, except for minor mutations, then we know basically what the DNA of your ancestor looks like and others can then test and compare results against that established line.
To form a baseline within a family, we generally test two individuals from two separate lines of the common ancestor, just in case an undocumented adoption has occurred. If these two individuals match, except for minor mutations, then we know basically what the DNA of your ancestor looks like and others can then test and compare results against that established line.
If you’re a female and can’t test for Y DNA markers, you’re not left out. You’ll need to use traditional genealogy to find male lineal descendants of your ancestor that carry the family name. Consider offering a scholarship for a descendent of that line to be tested and then advertise on Rootsweb lists and boards, on Yahoo groups, on Facebook and anyplace else that you think would be effective.
Mitochondrial results look slightly different from Y DNA, but the match information is in essence the same.
What Else Can We Tell?
The results of your tests not only tell you about your genealogy, they can also tell you about your deep ancestry and identify your deep ancestral clan.
Have you ever wondered where your ancestors came from before contemporary times? We know that for the most part surnames did not exist before 1066, and in some places did not exist until much later. The likelihood of us ever knowing where our ancestors were prior to 1066, unless we are extremely lucky, is very remote using conventional genealogical research methods.
However, now with the results of our DNA, we can peer through that keyhole and unlock that door. Based on the results of our tests, and the relative rarity of the combined numbers, humans are grouped together in clans called haplogroups. We know who was a member of which clan by both the tests shown above and a different kind of test, called a SNP (pronounced snip) test.
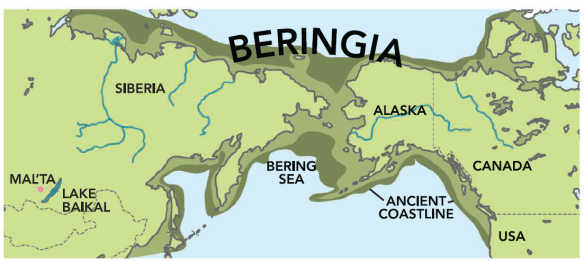
Population geneticists use this type of information to determine how groups of people migrated, and when. We may well be able to tell if our clan is Celtic, or Viking, African, Native American or related to Genghis Khan, for example. Based on our clan type, we may be able to tell where our group resided during the last ice age, and then trace their path from there to England or America over hundreds or thousands of years. While this sounds farfetched, it certainly isn’t and many people are discovering their deep ancestry. For example, we know that the Estes clan wintered the last ice age in Anatolia, and we know this because that is where other people who have this very rare combination of marker values are found in greater numbers than anyplace else on earth.
How Can I Test My Family?
It’s easy to get started. For Y DNA testing, you only need one male volunteer that carries your surname who is descended from your oldest progenitor by the same surname. To order a test kit, be sure to join a surname project for the best pricing. You can check on various surname projects by going to www.familytreedna.com and entering the surname in the search box on the right hand side of the page where it says “Search Your Last Name.”

I searched for Estes and the information returned tells me how many people, both male and female, have tested with that surname, if an Estes project exists, and the link, and any other projects where the administrator has specifically entered the Estes surname. So join the surname project and be sure to check out any others shown.

Anyone, males or females can test their mitochondrial DNA. To test your own mitochondrial DNA, just order a test kit, and then follow the branch on your pedigree chart directly up your maternal line of the tree (your mother, her mother, her mother, etc.) to see whose mitochondrial DNA you carry.
Autosomal, the Third Kind of DNA Testing
In the past two or three years, autosomal DNA testing has really come into its own. This type of testing does not focus on one line, like the Y-line DNA focuses only on the direct paternal surname line and the mitochondrial focuses only on the direct maternal line. The Y DNA and mtDNA are wonderful tests and provide you with huge amounts of information, but they can’t tell you anything about your other lines…not unless you can find a cousin from that other surname line and beg to have his or her DNA tested. This process (the testing, not the begging) is called building your DNA pedigree chart.
You can see an example of my DNA pedigree chart below. Being a female, I obviously can’t test for any Y DNA lines, so I had to find cousins to test for those lines. I can test for the direct mitochondrial line, but that still leaves most of the 14 great-great-grandparents with no information at all. By mining surname projects and begging cousins to test, I have filled in a number of these slots, but certainly not all.

But the time comes that you can’t complete the chart, or you have other genealogy questions to answer, and you’ll need to move to the third type of DNA testing, autosomal.
Autosomal testing provides you with two primary features.
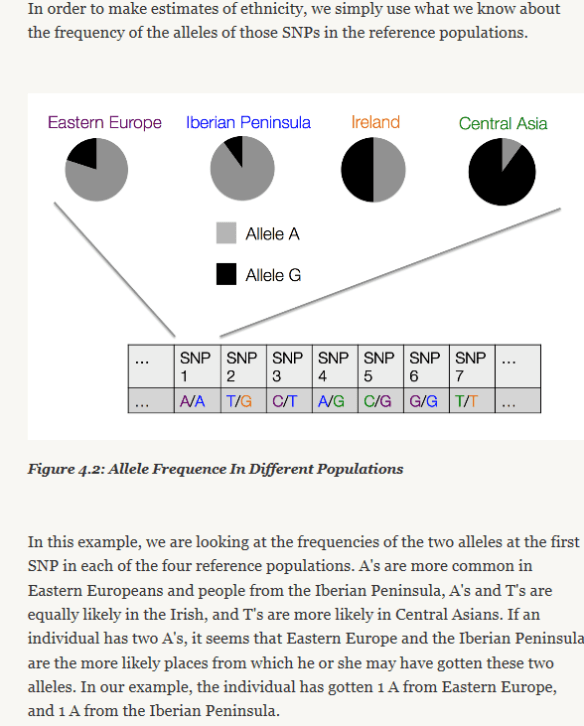
First, autosomal testing provides you with percentages of ethnicity. This may or may not excite you. Understand that when you’re looking for that elusive Native American great-great-great-grandmother, that you may or may not carry enough or a large enough piece of her DNA to be identified. But you’ll never know if you don’t test.

Second, you receive a list of cousin matches. These are people who match you on your autosomal results. This means that they are related to you on one line or another. It’s up to you to figure out which line, but there are tools and techniques to utilize. You probably won’t recognize the names of most of your matches, and you may or may not recognize a common ancestor. In some cases, the genealogy isn’t far enough back or there are other challenges in identifying a common ancestor. However, some huge brick walls have fallen for people and continue to fall daily by using autosomal tools to identify common ancestral families.

I wrote a series on “The Autosomal Me” which describes in detail how to utilize your Autosomal results.
Ok, now you’re convinced. You want to see who you match and meet those new cousins just waiting.
Summary – Who Can Test For What???
Just to be sure we all understand, here’s a handy chart that summarizes who can test for what at Family Tree DNA and what you discover!

What About The Test…
You may wonder why I recommend Family Tree DNA for testing. It’s simple. They are the only DNA testing company that offers the full range of tests and tools needed by genetic genealogists. They are the oldest company and have the largest data base, in addition to tools that facilitate using multiple types of test results together. Family Tree DNA has been wonderful to work with, sponsors free surname, haplogroup, geographic and special interest projects and are infinitely patient and extremely helpful. They are also a partner to the National Geographic Society and participants from the Genographic project can transfer results into the Family Tree DNA database for free.
Testing is done at Family Tree DNA using a cheek swab that looks like a Q-tip.

A test kit is shown above. Just swab the inside of your cheek, put the swab back in the vial and mail back to the lab. It’s that easy.
To see someone collecting a sample from receiving the envelope in the mail to mailing it off again, click here http://www.davedorsey.com/dna.html.
Receiving your Results
After you receive your Y DNA or mitochondrial results at Family Tree DNA on your personal page, please consider our Y-Line or Mitochondrial DNA Personal DNA Reports. Family Tree DNA customers who have minimally tested at 37 markers for the Y DNA or the mtDNA full sequence for mitochondrial can also order their reports directly through Family Tree DNA on their personal page.
What you discover from your own DNA will be priceless – and there is no other way to make these discoveries other than DNA testing. Your DNA results are notes in bottles that have sailed over time from your ancestor to you. Begin your adventure today, open that bottle and see what secrets your ancestors sent!
Be sure to sign up for the this blog to keep current with genetic genealogy. There is great introductory and educational material there as well, and it’s free. You can sign up by clicking on the little grey “follow” button in the upper right hand corner of the main blog page.
Happy ancestor hunting!!!