Looking in the rear view mirror, what a year! Some days it’s been hard to catch your breath things have been moving so fast.
What were the major happenings, how did they affect genetic genealogy and what’s coming in 2019?
The SNiPPY Award
First of all, I’m giving an award this year. The SNiPPY.
Yea, I know it’s kinda hokey, but it’s my way of saying a huge thank you to someone in this field who has made a remarkable contribution and that deserves special recognition.
Who will it be this year?
Drum roll…….
The 2018 SNiPPY goes to…
DNAPainter – The 2018 SNiPPY award goes to DNAPainter, without question. Applause, everyone, applause! And congratulations to Jonny Perl, pictured below at Rootstech!
Jonny Perl created this wonderful, visual tool that allows you to paint your matches with people on your chromosomes, assigning the match to specific ancestors.
I’ve written about how to use the tool with different vendors results and have discovered many different ways to utilize the painted segments. The DNA Painter User Group is here on Facebook. I use DNAPainter EVERY SINGLE DAY to solve a wide variety of challenges.
What else has happened this year? A lot!
Ancient DNA – Academic research seldom reports on Y and mitochondrial DNA today and is firmly focused on sequencing ancient DNA. Ancient genome sequencing has only recently been developed to a state where at least some remains can be successfully sequenced, but it’s going great guns now. Take a look at Jennifer Raff’s article in Forbes that discusses ancient DNA findings in the Americas, Europe, Southeast Asia and perhaps most surprising, a first generation descendant of a Neanderthal and a Denisovan.
Inroads were made into deeper understanding of human migration in the Americas as well in the paper Early human dispersals within the Americas by Moreno-Mayer et al.
I look for 2019 and on into the future to hold many more revelations thanks to ancient DNA sequencing as well as using those sequences to assist in understanding the migration patterns of ancient people that eventually became us.
Barbara Rae-Venter and the Golden State Killer Case
Using techniques that adoptees use to identify their close relatives and eventually, their parents, Barbara Rae-Venter assisted law enforcement with identifying the man, Joseph DeAngelo, accused (not yet convicted) of being the Golden State Killer (GSK).
A very large congratulations to Barbara, a retired patent attorney who is also a genealogist. Nature recognized Ms. Rae-Venter as one of 2018’s 10 People Who Mattered in Science.
DNA in the News
DNA is also represented on the 2018 Nature list by Viviane Slon, a palaeogeneticist who discovered an ancient half Neanderthal, half Denisovan individual and sequenced their DNA and He JianKui, a Chinese scientist who claims to have created a gene-edited baby which has sparked widespread controversy. As of the end of the year, He Jiankui’s research activities have been suspended and he is reportedly sequestered in his apartment, under guard, although the details are far from clear.
In 2013, 23andMe patented the technology for designer babies and I removed my kit from their research program. I was concerned at the time that this technology knife could cut two ways, both for good, eliminating fatal disease-causing mutations and also for ethically questionable practices, such as eugenics. I was told at the time that my fears were unfounded, because that “couldn’t be done.” Well, 5 years later, here we are. I expect the debate about the ethics and eventual regulation of gene-editing will rage globally for years to come.
Elizabeth Warren’s DNA was also in the news when she took a DNA test in response to political challenges. I wrote about what those results meant scientifically, here. This topic became highly volatile and politicized, with everyone seeming to have a very strongly held opinion. Regardless of where you fall on that opinion spectrum (and no, please do not post political comments as they will not be approved), the topic is likely to surface again in 2019 due to the fact that Elizabeth Warren has just today announced her intention to run for President. The good news is that DNA testing will likely be discussed, sparking curiosity in some people, perhaps encouraging them to test. The bad news is that some of the discussion may be unpleasant at best, and incorrect click-bait at worst. We’ve already had a rather unpleasant sampling of this.
Law Enforcement and Genetic Genealogy
The Golden State Killer case sparked widespread controversy about using GedMatch and potentially other genetic genealogy data bases to assist in catching people who have committed violent crimes, such as rape and murder.
GedMatch, the database used for the GSK case has made it very clear in their terms and conditions that DNA matches may be used for both adoptees seeking their families and for other uses, such as law enforcement seeking matches to DNA sequenced during a criminal investigation. Since April 2018, more than 15 cold case investigations have been solved using the same technique and results at GedMatch. Initially some people removed their DNA from GedMatch, but it appears that the overwhelming sentiment, based on uploads, is that people either aren’t concerned or welcome the opportunity for their DNA matches to assist apprehending criminals.
Parabon Nanolabs in May established a genetic genealogy division headed by CeCe Moore who has worked in the adoptee community for the past several years. The division specializes in DNA testing forensic samples and then assisting law enforcement with the associated genetic genealogy.
Currently, GedMatch is the only vendor supporting the use of forensic sample matching. Neither 23anMe nor Ancestry allow uploaded data, and MyHeritage and Family Tree DNA’s terms of service currently preclude this type of use.
MyHeritage
Wow talk about coming onto the DNA world stage with a boom.
 MyHeritage went from a somewhat wobbly DNA start about 2 years ago to rolling out a chromosome browser at the end of January and adding important features such as SmartMatching which matches your DNA and your family trees. Add triangulation to this mixture, along with record matching, and you’re got a #1 winning combination.
MyHeritage went from a somewhat wobbly DNA start about 2 years ago to rolling out a chromosome browser at the end of January and adding important features such as SmartMatching which matches your DNA and your family trees. Add triangulation to this mixture, along with record matching, and you’re got a #1 winning combination.
It was Gilad Japhet, the MyHeritage CEO who at Rootstech who christened 2018 “The Year of the Segment,” and I do believe he was right. Additionally, he announced that MyHeritage partnered with the adoption community by offering 15,000 free kits to adoptees.
In November, MyHeritage hosted MyHeritage LIVE, their first user conference in Oslo, Norway which focused on both their genealogical records offerings as well as DNA. This was a resounding success and I hope MyHeritage will continue to sponsor conferences and invest in DNA. You can test your DNA at MyHeritage or upload your results from other vendors (instructions here). You can follow my journey and the conference in Olso here, here, here, here and here.
GDPR
GDPR caused a lot of misery, and I’m glad the implementation is behind us, but the the ripples will be affecting everyone for years to come.
GDPR, the European Data Protection Regulation which went into effect on May 25, 2018 has been a mixed and confusing bag for genetic genealogy. I think the concept of users being in charge and understanding what is happened with their data, and in this case, their data plus their DNA, is absolutely sound. The requirements however, were created without any consideration to this industry – which is small by comparison to the Googles and Facebooks of the world. However, the Googles and Facebooks of the world along with many larger vendors seem to have skated, at least somewhat.
Other companies shut their doors or restricted their offerings in other ways, such as World Families Network and Oxford Ancestors. Vendors such as Ancestry and Family Tree DNA had to make unpopular changes in how their users interface with their software – in essence making genetic genealogy more difficult without any corresponding positive return. The potential fines, 20 million plus Euro for any company holding data for EU residents made it unwise to ignore the mandates.
In the genetic genealogy space, the shuttering of both YSearch and MitoSearch was heartbreaking, because that was the only location where you could actually compare Y STR and mitochondrial HVR1/2 results. Not everyone uploaded their results, and the sites had not been updated in a number of years, but the closure due to GDPR was still a community loss.
Today, mitoydna.org, a nonprofit comprised of genetic genealogists, is making strides in replacing that lost functionality, plus, hopefully more.
On to more positive events.
Family Tree DNA
In April, Family Tree DNA announced a new version of the Big Y test, the Big Y-500 in which at least 389 additional STR markers are included with the Big Y test, for free. If you’re lucky, you’ll receive between 389 and 439 new markers, depending on how many STR markers above 111 have quality reads. All customers are guaranteed a minimum of 500 STR markers in total. Matching was implemented in December.
These additional STR markers allow genealogists to assemble additional line marker mutations to more granularly identify specific male lineages. In other words, maybe I can finally figure out a line marker mutation that will differentiate my ancestor’s line from other sons of my founding ancestor😊
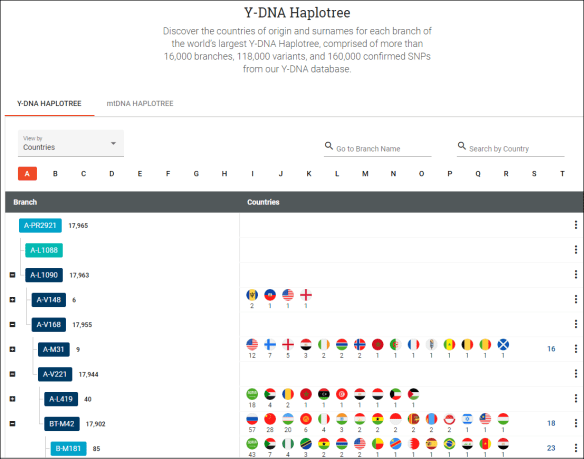
In June, Family Tree DNA announced that they had named more than 100,000 SNPs which means many haplogroup additions to the Y tree. Then, in September, Family Tree DNA published their Y haplotree, with locations, publicly for all to reference.
I was very pleased to see this development, because Family Tree DNA clearly has the largest Y database in the industry, by far, and now everyone can reap the benefits.
In October, Family Tree DNA published their mitochondrial tree publicly as well, with corresponding haplogroup locations. It’s nice that Family Tree DNA continues to be the science company.
You can test your Y DNA, mitochondrial or autosomal (Family Finder) at Family Tree DNA. They are the only vendor offering full Y and mitochondrial services complete with matching.
2018 Conferences
Of course, there are always the national conferences we’re familiar with, but more and more, online conferences are becoming available, as well as some sessions from the more traditional conferences.
I attended Rootstech in Salt Lake City in February (brrrr), which was lots of fun because I got to meet and visit with so many people including Mags Gaulden, above, who is a WikiTree volunteer and writes at Grandma’s Genes, but as a relatively expensive conference to attend, Rootstech was pretty miserable. Rootstech has reportedly made changes and I hope it’s much better for attendees in 2019. My attendance is very doubtful, although I vacillate back and forth.
On the other hand, the MyHeritage LIVE conference was amazing with both livestreamed and recorded sessions which are now available free here along with many others at Legacy Family Tree Webinars.
Family Tree University held a Virtual DNA Conference in June and those sessions, along with others, are available for subscribers to view.
The Virtual Genealogical Association was formed for those who find it difficult or impossible to participate in local associations. They too are focused on education via webinars.
Genetic Genealogy Ireland continues to provide their yearly conference sessions both livestreamed and recorded for free. These aren’t just for people with Irish genealogy. Everyone can benefit and I enjoy them immensely.
Bottom line, you can sit at home and educate yourself now. Technology is wonderful!
2019 Conferences
In 2019, I’ll be speaking at the National Genealogical Society Family History Conference, Journey of Discovery, in St. Charles, providing the Special Thursday Session titled “DNA: King Arthur’s Mighty Genetic Lightsaber” about how to use DNA to break through brick walls. I’ll also see attendees at Saturday lunch when I’ll be providing a fun session titled “Twists and Turns in the Genetic Road.” This is going to be a great conference with a wonderful lineup of speakers. Hope to see you there.
There may be more speaking engagements at conferences on my 2019 schedule, so stay tuned!
The Leeds Method
In September, Dana Leeds publicized The Leeds Method, another way of grouping your matches that clusters matches in a way that indicates your four grandparents.
I combine the Leeds method with DNAPainter. Great job Dana!
Genetic Affairs
In December, Genetic Affairs introduced an inexpensive subscription reporting and visual clustering methodology, but you can try it for free.
I love this grouping tool. I have already found connections I didn’t know existed previously. I suggest joining the Genetic Affairs User Group on Facebook.
DNAGedcom.com
 I wrote an article in January about how to use the DNAGedcom.com client to download the trees of all of your matches and sort to find specific surnames or locations of their ancestors.
I wrote an article in January about how to use the DNAGedcom.com client to download the trees of all of your matches and sort to find specific surnames or locations of their ancestors.
However, in December, DNAGedcom.com added another feature with their new DNAGedcom client just released that downloads your match information from all vendors, compiles it and then forms clusters. They have worked with Dana Leeds on this, so it’s a combination of the various methodologies discussed above. I have not worked with the new tool yet, as it has just been released, but Kitty Cooper has and writes about it here. If you are interested in this approach, I would suggest joining the Facebook DNAGedcom User Group.
Rootsfinder
I have not had a chance to work with Rootsfinder beyond the very basics, but Rootsfinder provides genetic network displays for people that you match, as well as triangulated views. Genetic networks visualizations are great ways to discern patterns. The tool creates match or triangulation groups automatically for you.
Training videos are available at the website and you can join the Rootsfinder DNA Tools group at Facebook.
Chips and Imputation
Illumina, the chip maker that provides the DNA chips that most vendors use to test changed from the OmniExpress to the GSA chip during the past year. Older chips have been available, but won’t be forever.
The newer GSA chip is only partially compatible with the OmniExpress chip, providing limited overlap between the older and the new results. This has forced the vendors to use imputation to equalize the playing field between the chips, so to speak.
This has also caused a significant hardship for GedMatch who is now in the position of trying to match reasonably between many different chips that sometimes overlap minimally. GedMatch introduced Genesis as a sandbox beta version previously, but are now in the process of combining regular GedMatch and Genesis into one. Yes, there are problems and matching challenges. Patience is the key word as the various vendors and GedMatch adapt and improve their required migration to imputation.
DNA Central
In June Blaine Bettinger announced DNACentral, an online monthly or yearly subscription site as well as a monthly newsletter that covers news in the genetic genealogy industry.
Many educators in the industry have created seminars for DNACentral. I just finished recording “Getting the Most out of Y DNA” for Blaine.
Even though I work in this industry, I still subscribed – initially to show support for Blaine, thinking I might not get much out of the newsletter. I’m pleased to say that I was wrong. I enjoy the newsletter and will be watching sessions in the Course Library and the Monthly Webinars soon.
If you or someone you know is looking for “how to” videos for each vendor, DNACentral offers “Now What” courses for Ancestry, MyHeritage, 23andMe, Family Tree DNA and Living DNA in addition to topic specific sessions like the X chromosome, for example.
Social Media
2018 has seen a huge jump in social media usage which is both bad and good. The good news is that many new people are engaged. The bad news is that people often given faulty advice and for new people, it’s very difficult (nigh on impossible) to tell who is credible and who isn’t. I created a Help page for just this reason.
You can help with this issue by recommending subscribing to these three blogs, not just reading an article, to newbies or people seeking answers.
- https://dna-explained.com/ (this blog)
- https://thegeneticgenealogist.com/ (Blaine Bettinger’s)
- https://www.legalgenealogist.com/ (Judy Russell’s)
Always feel free to post links to my articles on any social media platform. Share, retweet, whatever it takes to get the words out!
The general genetic genealogy social media group I would recommend if I were to select only one would be Genetic Genealogy Tips and Techniques. It’s quite large but well-managed and remains positive.
I’m a member of many additional groups, several of which are vendor or interest specific.
Genetic Snakeoil
Now the bad news. Everyone had noticed the popularity of DNA testing – including shady characters.
Be careful, very VERY careful who you purchase products from and where you upload your DNA data.
If something is free, and you’re not within a well-known community, then YOU ARE THE PRODUCT. If it sounds too good to be true, it probably is. If it sounds shady or questionable, it’s probably that and more, or less.
If reputable people and vendors tell you that no, they really can’t determine your Native American tribe, for example, no other vendor can either. Just yesterday, a cousin sent me a link to a “tribe” in Canada that will, “for $50, we find one of your aboriginal ancestors and the nation stamps it.” On their list of aboriginal people we find one of my ancestors who, based on mitochondrial DNA tests, is clearly NOT aboriginal. Snake oil comes in lots of flavors with snake oil salesmen looking to prey on other people’s desires.
When considering DNA testing or transfers, make sure you fully understand the terms and conditions, where your DNA is going, who is doing what with it, and your recourse. Yes, read every single word of those terms and conditions. For more about legalities, check out Judy Russell’s blog.
Recommended Vendors
All those DNA tests look yummy-good, but in terms of vendors, I heartily recommend staying within the known credible vendors, as follows (in alphabetical order).
For genetic genealogy for ethnicity AND matching:
- 23andMe
- Ancestry
- Family Tree DNA
- GedMatch (not a vendor because they don’t test DNA, but a reputable third party)
- MyHeritage
You can read about Which DNA Test is Best here although I need to update this article to reflect the 2018 additions by MyHeritage.
Understand that both 23andMe and Ancestry will sell your DNA if you consent and if you consent, you will not know who is using your DNA, where, or for what purposes. Neither Family Tree DNA, GedMatch, MyHeritage, Genographic Project, Insitome, Promethease nor LivingDNA sell your DNA.
The next group of vendors offers ethnicity without matching:
- Genographic Project by National Geographic Society
- Insitome
- LivingDNA (currently working on matching, but not released yet)
Health (as a consumer, meaning you receive the results)
- 23andMe (limited health)
- Promethease
Medical (as a contributor, meaning you are contributing your DNA for research)
- 23andMe
- Ancestry
- DNA.Land (not a testing vendor, doesn’t test DNA)
There are a few other niche vendors known for specific things within the genetic genealogy community, many of whom are mentioned in this article, but other than known vendors, buyer beware. If you don’t see them listed or discussed on my blog, there’s probably a reason.
What’s Coming in 2019
Just like we couldn’t have foreseen much of what happened in 2018, we don’t have access to a 2019 crystal ball, but it looks like 2019 is taking off like a rocket. We do know about a few things to look for:
- MyHeritage is waiting to see if envelope and stamp DNA extractions are successful so that they can be added to their database.
- www.totheletterDNA.com is extracting (attempting to) and processing DNA from stamps and envelopes for several people in the community. Hopefully they will be successful.
- LivingDNA has been working on matching since before I met with their representative in October of 2017 in Dublin. They are now in Beta testing for a few individuals, but they have also just changed their DNA processing chip – so how that will affect things and how soon they will have matching ready to roll out the door is unknown.
- Ancestry did a 2018 ethnicity update, integrating ethnicity more tightly with Genetic Communities, offered genetic traits and made some minor improvements this year, along with adding one questionable feature – showing your matches the location where you live as recorded in your profile. (23andMe subsequently added the same feature.) Ancestry recently said that they are promising exciting new tools for 2019, but somehow I doubt that the chromosome browser that’s been on my Christmas list for years will be forthcoming. Fingers crossed for something new and really useful. In the mean time, we can download our DNA results and upload to MyHeritage, Family Tree DNA and GedMatch for segment matching, as well as utilize Ancestry’s internal matching tools. DNA+tree matching, those green leaf shared ancestor hints, is still their strongest feature.
- The Family Tree DNA Conference for Project Administrators will be held March 22-24 in Houston this year, and I’m hopeful that they will have new tools and announcements at that event. I’m looking forward to seeing many old friends in Houston in March.
Here’s what I know for sure about 2019 – it’s going to be an amazing year. We as a community and also as individual genealogists will be making incredible discoveries and moving the ball forward. I can hardly wait to see what quandaries I’ve solved a year from now.
What mysteries do you want to unravel?
I’d like to offer a big thank you to everyone who made 2018 wonderful and a big toast to finding lots of new ancestors and breaking down those brick walls in 2019.
Happy New Year!!!
______________________________________________________________
Disclosure
I receive a small contribution when you click on some (but not all) of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research




























































 ay that the
ay that the 









