Mother’s Day is approaching, so I’m writing articles about mitochondrial DNA inspired by the most common questions in the Mitochondrial DNA for Genealogy Facebook group. I’ll be adding these articles to the Mitochondrial DNA Resource page, here.
FamilyTreeDNA has already started their Mother’s Day sale where both the mitochondrial DNA test and Family Finder are both on sale. Take a look.
I can’t believe how much the prices have dropped over the years – as the technology has improved. I took the full sequence mitochondrial DNA test when it was first offered and I think it was something like $800, as was the first autosomal test I ordered lo those many years ago.
Today, these tests are $139 and $59, respectively, and are critical tools for everyone’s genealogy.
Where Did My Mitochondrial DNA Haplogroup Come From?
This is one of the most common questions about mitochondrial DNA. Everyone wants to know something about their haplogroup.
The answer is multi-faceted and depends on the question you’re actually trying to answer.
There are really two flavors of this question:
- Where did my ancestors come from in a genealogical timeframe?
- Where did my ancestors come from before I can find them in genealogical records?
Clearly, the timeframes involved vary to some extent, because when records end varies for each ancestral line. Generally speaking, genealogy records don’t extend back beyond 500 years or so. Whenever your genealogy records end, that’s where your haplogroup and match information becomes critically important to your research.
Fortunately, we have tools to answer both types of questions which actually form a continuum.
Some answers rely on having taken a mitochondrial DNA test at FamilyTreeDNA and some don’t.
- We’ll discuss finding haplogroup information for people who have taken a (preferably full sequence) mitochondrial DNA test at FamilyTreeDNA.
- We’ll discuss how people who have obtained their haplogroups through autosomal testing at other vendors can find information.
- We’ll talk about finding haplogroup information when other family members have tested who carry the mitochondrial DNA of ancestors that you do not.
Tools exist for each of these situations.
Genealogical Timeframe
If you’re trying to answer the question of where other people who carry your haplogroup are found in the world, that question can be further subdivided:
- Where are the earliest known matrilineal ancestors of my mitochondrial DNA matches located?
- Where are other mitochondrial DNA testers who carry my haplogroup, even if I don’t match them, found in the world?
Let’s start at FamilyTreeDNA and then move to public resources.
FamilyTreeDNA
Mitochondrial DNA Tests
FamilyTreeDNA provides a great deal of information for people who have taken a mitochondrial DNA test. We’ll step through each tab on a tester’s personal page that’s relevant to haplogroups.
To find the location of your matches’ most distant ancestors, you need to have taken the mitochondrial DNA test at FamilyTreeDNA in order to obtain results and matches. I know this might seem like an obvious statement, but you’d be surprised how many people don’t realize that there are separate tests for Y and mitochondrial DNA.
Your most detailed, and therefore most accurate and specific results will result from taking the Full Sequence test, called the mtFull test and sometimes abbreviated as FMS (full mitochondrial sequence.)
Taking a full sequence test means you’ve tested all three different regions of the mitochondria, HVR1, HVR2, and the Coding Region. Don’t worry about those details. Today, the Full Sequence test is the only test you can order, but people who tested earlier could order a partial test. Those people can easily upgrade today.

click on images to enlarge
You can see, in the upper right-hand corner of the mitochondrial section of my personal page, above, that I’ve taken both tests. The “Plus” test is the HVR1 and HVR2 portion of the test.
If you haven’t taken any mitochondrial DNA test, then the mitochondrial section doesn’t show on your personal page.

If your Plus and Full buttons are both greyed out, that means you took the HVR1 level test only, and you can click on either button to upgrade.

If your “Full” button is greyed out, that means you haven’t tested at that level and you can click on the Full button to upgrade.
Entering Ancestor Information is Important
Genealogy is a collaborative sport and entering information about our ancestors is important – both for our own genealogy and for other testers too.
Your matches may or may not enter their ancestor’s information in all three locations where it can be useful:
- Earliest Known Ancestor (found under the dropdown beneath your name in the upper right-hand corner of your personal page, then “Account Settings,” then “Genealogy,” then “Earliest Known Ancestors”)
- Matches Map (found on your Y or mtDNA personal page tab or “Update Location” on Earliest Known Ancestors tab)
- Uploading or creating a tree (found under myTree at the very top of your personal page)
Please enter your information by following the notes above, or you can follow the step-by-step instructions, here. You’ll be glad you did.
Your Haplogroup

You’ll find your haplogroup name under the Badges section of your personal page as well as at the top of the mtDNA section.

click all images to enlarge
The mtDNA section at FamilyTreeDNA has five tabs that each provides different pieces of the puzzle of where your ancestors, and therefore your haplogroups, came from.
Checking all of these tabs in the mtDNA section of your results is critical to gather every piece of evidence provided by your matches and the scientists as well. Let’s take a look at each one and what they reveal about your haplogroup.
Let’s start with your matches.
Matches
On the matches page, you’ll only be matched with people who carry the same haplogroup – or at least the same base haplogroup.
The haplogroup level of your matches depends on the level of test they have taken. In other words, if your match has only taken the HVR1 level test, and they only have a base haplogroup of J, then you’ll only see them, and their haplogroup J, on your HVR1 match page. If they have tested at a higher level and you match them at the HVR1 level, you’ll see the most specific haplogroup possible as determined by the level they tested.
The (default) match page shows your matches at the highest-level test you have tested. In my case, that’s the “HVR1, HVR2, Coding Region” because I’ve taken the full sequence test which tests the entire mitochondria.
At the full sequence level match page, I’ll only see people who match me on the same extended haplogroup. In my case, that’s J1c2f.

Viewing your matches’ Earliest Known Ancestor shows where their ancestors were located, which provides clues as to where your common haplogroup was found in the world at that time. Based on those results, the geographic distribution, what you know about your own ancestors, and how far back in time, your matches’ information may be an important clue about your own ancestry.
Generally, the closer your matches, meaning the fewer mutations difference, the closer in time you share a common ancestor. I say “generally,” because mutations don’t happen on a time schedule and can happen in any generation.
The number of mutations is shown in the column “Genetic Distance.” Genetic Distance is the number of mutations difference between you and your match. So a 3 in the GD column means 3 mutations difference. A GD of 0 is an exact match. At the HVR1 and HVR2 levels, no genetic distance is provided because only exact matches are shown at those levels.
The little blue pedigree icons on the Matches page indicate people who have created or uploaded trees. You’ll definitely want to take a look at those. Sometimes you’ll discover that your matches have added more generations in their tree than is shown in the Earliest Known Ancestor field.
Is Taking the Full Sequence Test Important?
Why is taking the full sequence test important? Looking at my HVR1 matches, below, provides the perfect example.

This shows my first four HVR1-only matches. In other words, these people match me on a small subset of my mitochondrial DNA. About 1000 locations of the total 16,569 are tested in the HVR1 region. You can see that utilizing the HVR1 region, only, the people I match exactly in that region have different extended, or full haplogroups, assigned when taking the full sequence test.
Crystal and Katherine have both taken the full sequence test as indicated by FMS (full mitochondrial sequence,) and they are both haplogroup J1c2f, but Peter is haplogroup J1c2g – a different haplogroup.
Peter is shown as an exact match to me at the HVR1 level, but he has a different full haplogroup, so he won’t be shown as a match at the HVR1/HVR2/Coding Region (full sequence) level.
Crystal and Katherine will match me at the full sequence level if we have three or fewer mutations difference in total.
Susan has only tested to the HVR1 level, so she can only be assigned to haplogroup J from those 1000 locations. That tells us that (at least) one of mutations that defines haplogroup J resides in the HVR1 region.
At the HVR1 matching level, I’ll be matched with everyone I match exactly so long as they are in haplogroup J, the common denominator haplogroup of everyone at that level.
If Susan were to test at the full sequence level, she would obtain a full haplogroup and I might continue to match her at the full sequence level if she is haplogroup J1c2f and matches me with three or fewer mutations difference. At the full sequence level, I’ll only match people who match my haplogroup exactly and match at a genetic distance of 0, 1, 2 or 3.
Now, let’s look at the Ancestral Origins tab.
Ancestral Origins

The Ancestral Origins tab is organized by Country within match level. In the example above, I’ve shown exact matches or GD=0.
The match total on the Ancestral Origins tab shows the number of people whose ancestors were from various locations – as entered by the testers.
The most common places for my full sequence exact matches are in Norway and Sweden. That’s interesting because my ancestor was found in Germany in the 1600s.
There is also a comments column, to the right, not shown here, which may hold additional information of interest such as “Ashkenazi” or “Sicily” or “Canary Islands.”
The Country Total column is interesting too because it tells you how many people are in the database who have indicated that location as ancestral. The Match Percentage column is pretty much irrelevant unless your haplogroup is extremely rare.
Matches Map
The matches map falls into the “picture is worth 1000 words category.”

This is the map of the earliest known matrilineal ancestor locations of my full sequence matches.
My ancestor is the white pin in Germany. Red=exact match, orange=1 mutation difference, yellow=2 mutations difference. I have no GD=3 matches showing.

By clicking on any pin, you can see additional information about the ancestor of the tester.
You can also select an option on the map to view lower testing levels, such as my HVR1 matches shown below.

While some people are tempted to ignore the HVR1 or HVR2 Matches Maps, I don’t.
If the question you’re trying to answer is where your haplogroup came from, viewing the map of where people are located who may match you more distantly in time is useful. While we know for sure that some of these people have different full haplogroups, we also know that they are all members of haplogroup J plus some subclade. Therefore, these matches shared a common haplogroup J ancestor.
J subgroups are clearly European but some are found in Anatolia, the path out of Africa to Europe, although that could be a function of back-migration.
When looking at match maps, keep two things in mind:
- The information is provided by testers. It’s possible for them to misunderstand what is meant by providing the information for their earliest known “direct maternal ancestor.” I can’t tell you how many male names I’ve seen here. Clearly, the tester misunderstood the purpose and what was being asked – because men don’t pass mitochondrial DNA to their offspring. Check the pins for surnames that seem to fit the pin location, and that pins have been accurately placed.
- Testing bias. In other words, lots of people have tested in the US as compared to Europe, and probably more people in the UK than say, Turkey. Testing is still illegal in France.
Haplogroup Origins

While the Ancestral Origins tab is organized by the locations of your matches ancestors, the Haplogroup Origins tab is focused on your haplogroup by match level only.
In many cases, the numbers will match your Ancestral Origins exactly, but for other test levels, the numbers will be different.

For example, at the HVR1/HVR2 level, I can easily see at a glance the locations where my haplogroup is found, and the number of my matches in those various locations.
This page is reflective of where the haplogroup itself is found, according to your matches.
There may be other people with the same haplogroup that you don’t match and won’t be reflected on this page. We’ll see them either in projects or on the Public Mitochondrial Tree in following sections.
Migration Map

The migration map tab shows the path between Mitochondrial Eve who lived in African about 145,000 years ago and your haplogroup today. For haplogroups J, Eve’s descendant left African and traveled through the Middle East and on into Southwest Asia before turning left and migrating throughout Europe.
Clearly, the vast majority of this migration occurred before genealogy, but not all, or you wouldn’t be here today.
Thousands of my ancestors brought my mitochondrial DNA from Africa through Anatolia, through Europe, to Scandinavia, and back to Germany – then on to the US where it continued being passed on for five more generations before reaching me.
Additional Features – Other Tools
On your personal page, scroll down below your Mitochondrial DNA results area and you’ll see Public Haplotrees under the Other Tools tab.

This tree is available to FamilyTreeDNA customers as well as the public.
Public Mitochondrial DNA Haplotree
The public mitochondrial haplotree provided by FamilyTreeDNA includes location information and is available to everyone, customer or not, for free. Please note that only full sequence results were used to construct this tree, so partial results, meaning haplogroups of people who tested at the HVR1/2 levels only, are not included because the haplogroup cannot be refined at that level.
If you’ve received a haplogroup from a different test at another vendor, you can use this public tool to obtain location information. FamilyTreeDNA has the single largest repository of mitochondrial tests in the world, having tested customers for 21 years, and they have made this tree with location information available for everyone.
If you are a customer, you can sign in and access this tree from your account, above.

If you access the haplotree in this manner, be sure to select the mtDNA tree, not the Y DNA tree which is the default.
Or you can simply access the mtDNA the same way as the public, below.
Go to the main FamilyTreeDNA page by clicking here.
On the main page, scroll to the very bottom – yes, just keep scrolling.

At the very bottom, in the footer, you’ll see “Community.” (Hint, if you don’t see Community at the very bottom of this page, you’re probably signed in to your account.)
Click on “mtDNA Haplotree.”

Next, you’ll see the beginning, or root, of the mitochondrial DNA tree, with the RSRS at the top of the page. The tree structure and haplogroups are defined at Phylotree Build 17, here. All of the main daughter haplogroups, such as “J,” are displayed beneath or you can select them across the top.
Enter the haplogroup name in the “Branch Name” field in the upper right. For me, that’s J1c2f.
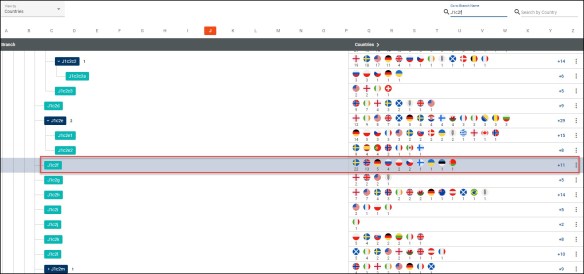
I don’t match all of the J1c2f people in the database, because there more total country designations shown here (82) than I have full sequence matches with locations provided (50 from my Ancestral Origins page.)

If you click on the three dots at right, you’ll see a Country Report which provides details for this haplogroup and downstream haplogroups, if there are any. I wrote about that, in detail, here.
There are no “J1c2f plus a daughter” haplogroups defined today, so there is nothing listed downstream.
However, that’s not always the case. There may be a downstream clade that you’re not a member of, meaning you don’t carry that haplogroup-defining mutation.
Or, you may have tested someplace that provides you with a partial haplogroup, so you don’t know if you have a subclade or not. You can still glean useful information from partial haplogroups.
Partial Haplogroups From Autosomal Tests
There’s nothing “wrong” with partial haplogroups. It’s nice to know at least some history about your matrilineal ancestry. What you don’t receive, of course, aside from matching, is more recent, genealogical, information.
Both 23andMe and LivingDNA provide autosomal customers with partial mitochondrial haplogroups. Both of these vendors tend to be accurate as far as they go, as opposed to other vendors, who shall remain unnamed, that are often inaccurate.
Autosomal tests don’t specifically test the mitochondrial DNA directly like a full sequence mitochondrial DNA test does, but they do use “probes” that scan specific haplogroup defining locations. Of course, each of the autosomal chips has a finite number of locations and every location that is used for either mitochondrial or Y DNA haplogroups is a space the vendors can’t use for autosomal locations.
Therefore, customers receive partial haplogroups.
In my case, I’ve received J1c at LivingDNA and J1c2 at 23andMe.

Both vendors provide basic information about your haplogroup, along with migration maps. Wikipedia also provides basic haplogroup information. Google is your friend – “mitochondrial haplogroup J Wikipedia.”
DNA Projects
Most haplogroups have a DNA project at FamilyTreeDNA. Note that these projects are administered by volunteers, so your mileage will vary in terms of participant grouping, along with whether or not results or maps are displayed. You can just google for “mitochondrial haplogroup J DNA project at FamilyTreeDNA” and you’ll find the project or perhaps multiple projects to select from. Some haplogroups have a main “J” project and perhaps a subproject, like “J1c,” for example.

You can join the project, either from this page if you’ve tested at FamilyTreeDNA, or from your personal page via the “myProjects” tab at the top of your personal page.
If you’re looking for public haplogroup information, click on “DNA Results.”

If the Haplogroup J DNA testers have joined this project, authorized displaying their results in projects, and provided ancestor information, you will be able to see that on the “Results” page. Projects are often grouped by haplogroup subgroup. Please note that the default page display size is 25, so scroll to the bottom to see how many pages are in the project. Multiply that number times 25 (182 pages total X 25 = 4550) and change the page display size to that number (4550, in this case.)
One of the most useful tools for haplogroup discovery is the project map which offers the same subgroups as the project groupings.

You can select “All” on the dropdown to display the locations of the earliest known ancestors of everyone in this haplogroup project, or you can select a subclade. This map is displaying haplogroup J1c2 as an example of my partial haplogroup.
The Public Mitochondrial Tree and Partial Haplogroups
To find more comprehensive information for partial haplogroups, I can use the free mitochondrial tree at FamilyTreeDNA. While projects only reflect information for people who have joined those particular projects, the tree provides more comprehensive information.
Anyone with a partial haplogroup can still learn a great deal. Like with any haplogroup, you can view where tester’s ancestors lived in the world.
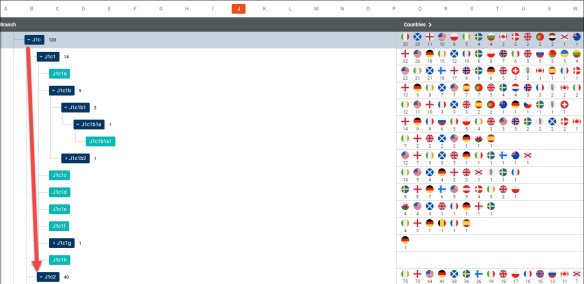
In this case, it doesn’t matter whether I’m looking at partial haplogroups J1c or J1c2, there are many subgroups that I could potentially belong to.
In fact, haplogroup J1c has subclades through J1c17, so there are pages and pages of haplogroup subclade candidates.
Does a Full Haplogroup Really Matter?
How much difference can there be? Is J1c or J1c2 good enough? Good questions.
It depends – on what you want to know.
- For general interest, perhaps.
- For genealogy, no.
Genealogists need the most granular results possible to obtain the most information possible. You don’t know what you don’t know. But how much might that be, aside from full sequence matches?
There’s a significant difference in the country details of haplogroup J1c, J1c2 and J1c2f. I created a chart of the top 10 locations, and how many people’s ancestors are found there for J1c, J1c2, and J1c2f.

Wow, that’s a big difference.
How accurately do J1c and J1c2 results reflect the locations in my full J1c2f haplogroup? I color-coded the results and removed the locations from J1c and J1c2 that are not reflected in J1c2f.

As it turns out, the 5 most frequent locations in J1c and the top 3 locations in J1c2 aren’t even in the top 10 of J1c2f. Obtaining a full haplogroup is important.
Current and Past Populations
It’s worth noting that where a current population is found is not always indicative of where an ancestral population was found.
Of course, with genealogy, we can look back a few generations by seeing where the ancestors of our close and distant matches were found.
My earliest known ancestor is found in a marriage record in 1647 in Wirbenz, Germany when she was 26 years old. However, the majority of my exact mitochondrial DNA matches are not found in Germany, or even in Europe, but in Scandinavia. I’m sure there’s a story there to be told, possibly related to the Thirty Years’ War which began in 1618 and devastated Germany. The early German records where she lived were destroyed.
Even in the abbreviated genealogical timeframe where records and surnames exist, as compared to the history of mankind and womankind, we can see examples of population migration and shift with weather, warfare, and opportunity.
We can’t peer further back in time, at least not without ancient DNA, except by a combination of general history, haplogroup inference, and noting where branching from our mother clade occurred.
We know that people move. Sometimes populations were small and the entire population moved to a new location.
Sometimes, the entire population didn’t move, the but descendants of the migrating group survived to take DNA tests, while the population remaining in the original location has no present-day descendants.
Sometimes descendants of both groups survived.
Of course, throughout history, mutations continued to occur in all lines, forming new genetic branches – haplogroups.
Thank goodness they did, because mutations, or lack thereof, are incredibly important clues to genealogy as well as being our breadcrumbs back into the mists of distant time. Those haplogroup-defining mutations are the umbilical cord that allows us to connect with those distant ancestors.
These tools, especially used together, are the best way to answer the question, “Where did my Mitochondrial DNA Haplogroup Come From?”
Where did your haplogroup come from?
_____________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Products and Services
Books
Genealogy Research