Not only have we put 2020 in the rear-view mirror, thankfully, we’re at the 20-year, two-decade milestone. The point at which genetics was first added to the toolbox of genealogists.
It seems both like yesterday and forever ago. And yes, I’ve been here the whole time, as a spectator, researcher, and active participant.
Let’s put this in perspective. On New Year’s Eve, right at midnight, in 2005, I was able to score kit number 50,000 at Family Tree DNA. I remember this because it seemed like such a bizarre thing to be doing at midnight on New Year’s Eve. But hey, we genealogists are what we are.
I knew that momentous kit number which seemed just HUGE at the time was on the threshold of being sold, because I had inadvertently purchased kit 49,997 a few minutes earlier.
Somehow kit 50,000 seemed like such a huge milestone, a landmark – so I quickly bought kits, 49,998, 49,999, and then…would I get it…YES…kit 50,000. Score!

That meant that in the 5 years FamilyTreeDNA had been in business, they had sold on an average of 10,000 kits per year, or 27 kits a day. Today, that’s a rounding error. Then it was momentous!
In reality, the sales were ramping up quickly, because very few kits were sold in 2000, and roughly 20,000 kits had been sold in 2005 alone. I know this because I purchased kit 28,429 during the holiday sale a year earlier.
Of course, I had no idea who I’d test with that momentous New Year’s Eve Y DNA kit, but I assuredly would find someone. A few months later, I embarked on a road trip to visit an elderly family member with that kit in tow. Thank goodness I did, and they agreed and swabbed on the spot, because they are gone today and with them, the story of the Y line and autosomal DNA of their branch.
In the past two decades, almost an entire generation has slipped away, and with them, an entire genealogical library held in their DNA.
Today, more than 40 million people have tested with the four major DNA testing companies, although we don’t know exactly how many.
Lots of people have had more time to focus on genealogy in 2020, so let’s take a look at what’s important? What’s going on and what matters beyond this month or year?
How has this industry changed in the last two decades, and where it is going?
Reflection
This seems like a good point to reflect a bit.

Professor Dan Bradley reflecting on early genetic research techniques in his lab at the Smurfit Institute of Genetics at Trinity College in Dublin. Photo by Roberta Estes
In the beginning – twenty years ago, there were two companies who stuck their toes in the consumer DNA testing water – Oxford Ancestors and Family Tree DNA. About the same time, Sorenson Genomics and GeneTree were also entering that space, although Sorenson was a nonprofit. Today, of those, only FamilyTreeDNA remains, having adapted with the changing times – adding more products, testing, and sophistication.
Bryan Sykes who founded Oxford Ancestors announced in 2018 that he was retiring to live abroad and subsequently passed away in 2020. The website still exists, but the company has announced that they have ceased sales and the database will remain open until Sept 30, 2021.
James Sorenson died in 2008 and the assets of Sorenson Molecular Genealogy Foundation, including the Sorenson database, were sold to Ancestry in 2012. Eventually, Ancestry removed the public database in 2015.
Ancestry dabbled in Y and mtDNA for a while, too, destroying that database in 2014.
Other companies, too many to remember or mention, have come and gone as well. Some of the various company names have been recycled or purchased, but aren’t the same companies today.
In the DNA space, it was keep up, change, die or be sold. Of course, there was the small matter of being able to sell enough DNA kits to make enough money to stay in business at all. DNA processing equipment and a lab are expensive. Not just the equipment, but also the expertise.
The Next Wave
As time moved forward, new players entered the landscape, comprising the “Big 4” testing companies that constitute the ponds where genealogists fish today.
23andMe was the first to introduce autosomal DNA testing and matching. Their goal and focus was always medical genetics, but they recognized the potential in genealogists before anyone else, and we flocked to purchase tests.
Ancestry settled on autosomal only and relies on the size of their database, a large body of genealogy subscribers, and a widespread “feel-good” marketing campaign to sell DNA kits as the gateway to “discover who you are.”
FamilyTreeDNA did and still does offer all 3 kinds of tests. Over the years, they have enhanced both the Y DNA and mitochondrial product offerings significantly and are still known as “the science company.” They are the only company to offer the full range of Y DNA tests, including their flagship Big Y-700, full sequence mitochondrial testing along with matching for both products. Their autosomal product is called Family Finder.
MyHeritage entered the DNA testing space a few years after the others as the dark horse that few expected to be successful – but they fooled everyone. They have acquired companies and partnered along the way which allowed them to add customers (Promethease) and tools (such as AutoCluster by Genetic Affairs), boosting their number of users. Of course, MyHeritage also offers users a records research subscription service that you can try for free.
In summary:
One of the wonderful things that happened was that some vendors began to accept compatible raw DNA autosomal data transfer files from other vendors. Today, FamilyTreeDNA, MyHeritage, and GEDmatch DO accept transfer files, while Ancestry and 23andMe do not.
The transfers and matching are free, but there are either minimal unlock or subscription plans for advanced features.
There are other testing companies, some with niche markets and others not so reputable. For this article, I’m focusing on the primary DNA testing companies that are useful for genealogy and mainstream companion third-party tools that complement and enhance those services.
The Single Biggest Change
As I look back, the single biggest change is that genetic genealogy evolved from the pariah of genealogy where DNA discussion was banned from the (now defunct) Rootsweb lists and summarily deleted for the first few years after introduction. I know, that’s hard to believe today.
Why, you ask?
Reasons varied from “just because” to “DNA is cheating” and then morphed into “because DNA might do terrible things like, maybe, suggest that a person really wasn’t related to an ancestor in a lineage society.”
Bottom line – fear and misunderstanding. Change is exceedingly difficult for humans, and DNA definitely moved the genealogy cheese.
From that awkward beginning, genetic genealogy organically became a “thing,” a specific application of genealogy. There was paper-trail traditional genealogy and then the genetic aspect. Today, for almost everyone, genealogy is “just another tool” in the genealogist’s toolbox, although it does require focused learning, just like any other tool.
DNA isn’t separate anymore, but is now an integral part of the genealogical whole. Having said that, DNA can’t solve all problems or answer all questions, but neither can traditional paper-trail genealogy. Together, each makes the other stronger and solves mysteries that neither can resolve alone.
Synergy.
I fully believe that we have still only scratched the surface of what’s possible.
Inheritance
As we talk about the various types of DNA testing and tools, here’s a quick graphic to remind you of how the different types of DNA are inherited.

- Y DNA is inherited paternally for males only and informs us of the direct patrilineal (surname) line.
- Mitochondrial DNA is inherited by everyone from their mothers and informs us of the mother’s matrilineal (mother’s mother’s mother’s) line.
- Autosomal DNA can be inherited from potentially any ancestor in random but somewhat predictable amounts through both parents. The further back in time, the less identifiable DNA you’ll inherit from any specific ancestor. I wrote about that, here.
What’s Hot and What’s Not
Where should we be focused today and where is this industry going? What tools and articles popped up in 2020 to help further our genealogy addiction? I already published the most popular articles of 2020, here.
This industry started two decades ago with testing a few Y DNA and mitochondrial DNA markers, and we were utterly thrilled at the time. Both tests have advanced significantly and the prices have dropped like a stone. My first mitochondrial DNA test that tested only 400 locations cost more than $800 – back then.
Y DNA and mitochondrial DNA are still critically important to genetic genealogy. Both play unique roles and provide information that cannot be obtained through autosomal DNA testing. Today, relative to Y DNA and mitochondrial DNA, the biggest challenge, ironically, is educating newer genealogists about their potential who have never heard about anything other than autosomal, often ethnicity, testing.
We have to educate in order to overcome the cacophony of “don’t bother because you don’t get as many matches.”
That’s like saying “don’t use the right size wrench because the last one didn’t fit and it’s a bother to reach into the toolbox.” Not to mention that if everyone tested, there would be a lot more matches, but I digress.
If you don’t use the right tool, and all of the tools at your disposal, you’re not going to get the best result possible.
The genealogical proof standard, the gold standard for genealogy research, calls for “a reasonably exhaustive search,” and if you haven’t at least considered if or how Y
DNA and mitochondrial DNA along with autosomal testing can or might help, then your search is not yet exhaustive.
I attempt to obtain the Y and mitochondrial DNA of every ancestral line. In the article, Search Techniques for Y and Mitochondrial DNA Test Candidates, I described several methodologies to find appropriate testing candidates.
Y DNA – 20 Years and Still Critically Important
Y DNA tracks the Y chromosome for males via the patrilineal (surname) line, providing matching and historical migration information.
We started 20 years ago testing 10 STR markers. Today, we begin at 37 markers, can upgrade to 67 or 111, but the preferred test is the Big Y which provides results for 700+ STR markers plus results from the entire gold standard region of the Y chromosome in order to provide the most refined results. This allows genealogists to use STR markers and SNP results together for various aspects of genealogy.

I created a Y DNA resource page, here, in order to provide a repository for Y DNA information and updates in one place. I would encourage anyone who can to order or upgrade to the Big Y-700 test which provides critical lineage information in addition to and beyond traditional STR testing. Additionally, the Big Y-700 test helps build the Y DNA haplotree which is growing by leaps and bounds.
More new SNPs are found and named EVERY SINGLE DAY today at FamilyTreeDNA than were named in the first several years combined. The 2006 SNP tree listed a grand total of 459 SNPs that defined the Y DNA tree at that time, according to the ISOGG Y DNA SNP tree. Goran Rundfeldt, head of R&D at FamilyTreeDNA posted this today:
2020 was an awful year in so many ways, but it was an unprecedented year for human paternal phylogenetic tree reconstruction. The FTDNA Haplotree or Great Tree of Mankind now includes:
– 37,534 branches with 12,696 added since 2019 – 51% growth!
defined by
– 349,097 SNPs with 131,820 added since 2019 – 61% growth!
In just one year, 207,536 SNPs were discovered and assigned FT SNP names. These SNPs will help define new branches and refine existing ones in the future.
The tree is constructed based on high coverage chromosome Y sequences from:
– More than 52,500 Big Y results
– Almost 4,000 NGS results from present-day anonymous men that participated in academic studies
Plus an additional 3,000 ancient DNA results from archaeological remains, of mixed quality and Y chromosome coverage at FamilyTreeDNA.
Wow, just wow.
These three new articles in 2020 will get you started on your Y DNA journey!
Mitochondrial DNA – Matrilineal Line of Humankind is Being Rewritten
The original Oxford Ancestor’s mitochondrial DNA test tested 400 locations. The original Family Tree DNA test tested around 1000 locations. Today, the full sequence mitochondrial DNA test is standard, testing the entire 16,569 locations of the mitochondria.

Mitochondrial DNA tracks your mother’s direct maternal, or matrilineal line. I’ve created a mitochondrial DNA resource page, here that includes easy step-by-step instructions for after you receive your results.
New articles in 2020 included the introduction of The Million Mito Project. 2021 should see the first results – including a paper currently in the works.
The Million Mito Project is rewriting the haplotree of womankind. The current haplotree has expanded substantially since the first handful of haplogroups thanks to thousands upon thousands of testers, but there is so much more information that can be extracted today.
Y and Mitochondrial Resources
If you don’t know of someone in your family to test for Y DNA or mitochondrial DNA for a specific ancestral line, you can always turn to the Y DNA projects at Family Tree DNA by searching here.

The search provides you with a list of projects available for a specific surname along with how many customers with that surname have tested. Looking at the individual Y DNA projects will show the earliest known ancestor of the surname line.
Another resource, WikiTree lists people who have tested for the Y DNA, mitochondrial DNA and autosomal DNA lines of specific ancestors.

Click on images to enlarge
On the left side, my maternal great-grandmother’s profile card, and on the right, my paternal great-great-grandfather. You can see that someone has tested for the mitochondrial DNA of Nora (OK, so it’s me) and the Y DNA of John Estes (definitely not me.)
MitoYDNA, a nonprofit volunteer organization created a comparison tool to replace Ysearch and Mitosearch when they bit the dust thanks to GDPR.
MitoYDNA accepts uploads from different sources and allows uploaders to not only match to each other, but to view the STR values for Y DNA and the mutation locations for the HVR1 and HVR2 regions of mitochondrial DNA. Mags Gaulden, one of the founders, explains in her article, What sets mitoYDNA apart from other DNA Databases?.
If you’ve tested at nonstandard companies, not realizing that they didn’t provide matching, or if you’ve tested at a company like Sorenson, Ancestry, and now Oxford Ancestors that is going out of business, uploading your results to mitoYDNA is a way to preserve your investment. PS – I still recommend testing at FamilyTreeDNA in order to receive detailed results and compare in their large database.
CentiMorgans – The Word of Two Decades
The world of autosomal DNA turns on the centimorgan (cM) measure. What is a centimorgan, exactly? I wrote about that unit of measure in the article Concepts – CentiMorgans, SNPs and Pickin’ Crab.
Fortunately, new tools and techniques make using cMs much easier. The Shared cM Project was updated this year, and the results incorporated into a wonderfully easy tool used to determine potential relationships at DNAPainter based on the number of shared centiMorgans.
Match quality and potential relationships are determined by the number of shared cMs, and the chromosome browser is the best tool to use for those comparisons.
Chromosome Browser – Genetics Tool to View Chromosome Matches
Chromosome browsers allow testers to view their matching cMs of DNA with other testers positioned on their own chromosomes.

My two cousins’ DNA where they match me on chromosomes 1-4, is shown above in blue and red at Family Tree DNA. It’s important to know where you match cousins, because if you match multiple cousins on the same segment, from the same side of your family (maternal or paternal), that’s suggestive of a common ancestor, with a few caveats.
Some people feel that a chromosome browser is an advanced tool, but I think it’s simply standard fare – kind of like driving a car. You need to learn how to drive initially, but after that, you don’t even think about it – you just get in and go. Here’s help learning how to drive that chromosome browser.
Triangulation – Science Plus Group DNA Matching Confirms Genealogy
The next logical step after learning to use a chromosome browser is triangulation. If fact, you’re seeing triangulation above, but don’t even realize it.
The purpose of genetic genealogy is to gather evidence to “prove” ancestral connections to either people or specific ancestors. In autosomal DNA, triangulation occurs when:
- You match at least two other people (not close relatives)
- On the same reasonably sized segment of DNA (generally 7 cM or greater)
- And you can assign that segment to a common ancestor

The same two cousins are shown above, with triangulated segments bracketed at MyHeritage. I’ve identified the common ancestor with those cousins that those matching DNA segments descend from.
MyHeritage’s triangulation tool confirms by bracketing that these cousins also match each other on the same segment, which is the definition of triangulation.
I’ve written a lot about triangulation recently.
If you’d prefer a video, I recorded a “Top Tips” Facebook LIVE with MyHeritage.
Why is Ancestry missing from this list of triangulation articles? Ancestry does not offer a chromosome browser or segment information. Therefore, you can’t triangulate at Ancestry. You can, however, transfer your Ancestry DNA raw data file to either FamilyTreeDNA, MyHeritage, or GEDmatch, all three of which offer triangulation.
Step by step download/upload transfer instructions are found in this article:
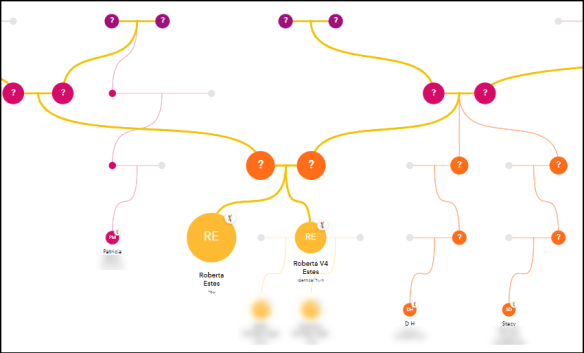
Clustering Matches and Correlating Trees
Based on what we’ve seen over the past few years, we can no longer depend on the major vendors to provide all of the tools that genealogists want and need.
Of course, I would encourage you to stay with mainstream products being used by a significant number of community power users. As with anything, there is always someone out there that’s less than honorable.
2020 saw a lot of innovation and new tools introduced. Maybe that’s one good thing resulting from people being cooped up at home.

Third-party tools are making a huge difference in the world of genetic genealogy. My favorites are Genetic Affairs, their AutoCluster tool shown above, DNAPainter and DNAGedcom.
These articles should get you started with clustering.
If you like video resources, here’s a MyHeritage Facebook LIVE that I recorded about how to use AutoClusters:
I created a compiled resource article for your convenience, here:
I have not tried a newer tool, YourDNAFamily, that focuses only on 23andMe results although the creator has been a member of the genetic genealogy community for a long time.
Painting DNA Makes Chromosome Browsers and Triangulation Easy
DNAPainter takes the next step, providing a repository for all of your painted segments. In other words, DNAPainter is both a solution and a methodology for mass triangulation across all of your chromosomes.

Here’s a small group of people who match me on the same maternal segment of chromosome 1, including those two cousins in the chromosome browser and triangulation sections, above. We know that this segment descends from Philip Jacob Miller and his wife because we’ve been able to identify that couple as the most distant ancestor intersection in all of our trees.
It’s very helpful that DNAPainter has added the functionality of painting all of the maternal and paternal bucketed matches from Family Tree DNA.
All you need to do is to link your known matches to your tree in the proper place at FamilyTreeDNA, then they do the rest by using those DNA matches to indicate which of the rest of your matches are maternal and paternal. Instructions, here. You can then export the file and use it at DNAPainter to paint all of those matches on the correct maternal or paternal chromosomes.
Here’s an article providing all of the DNAPainter Instructions and Resources.
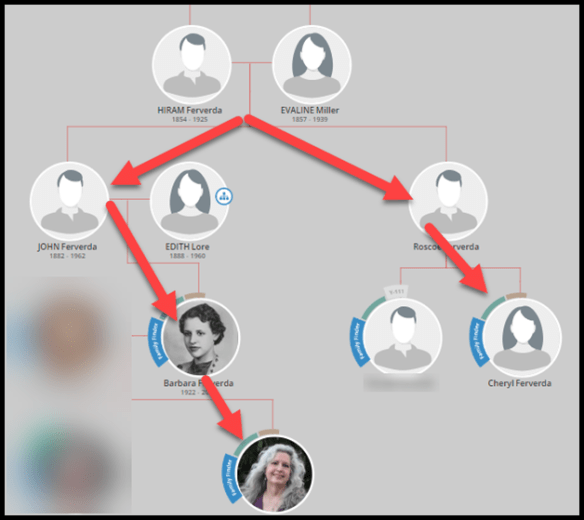
DNA Matches Plus Trees Enhance Genealogy
Of course, utilizing DNA matching plus finding common ancestors in trees is one of the primary purposes of genetic genealogy – right?
Vendors have linked the steps of matching DNA with matching ancestors in trees.
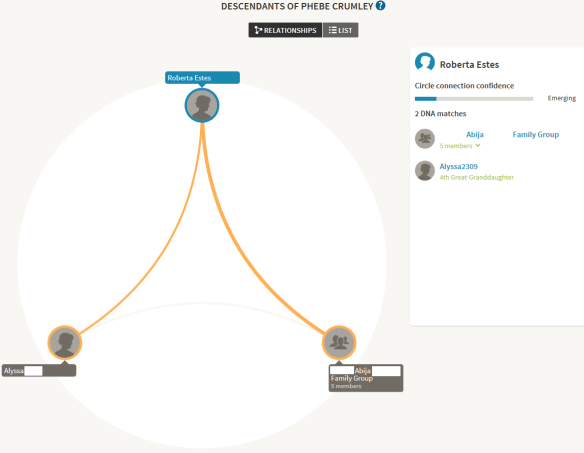
Genetic Affairs take this a step further. If you don’t have an ancestor in your tree, but your matches have common ancestors with each other, Genetic Affairs assembles those trees to provide you with those hints. Of course, that common ancestor might not be relevant to your genealogy, but it just might be too!

click to enlarge
This tree does not include me, but two of my matches descend from a common ancestor and that common ancestor between them might be a clue as to why I match both of them.
Ethnicity Continues to be Popular – But Is No Shortcut to Genealogy
Ethnicity is always popular. People want to “do their DNA” and find out where they come from. I understand. I really do. Who doesn’t just want an answer?
Of course, it’s not that simple, but that doesn’t mean it’s not disappointing to people who test for that purpose with high expectations. Hopefully, ethnicity will pique their curiosity and encourage engagement.
All four major vendors rolled out updated ethnicity results or related tools in 2020.
The future for ethnicity, I believe, will be held in integrated tools that allow us to use ethnicity results for genealogy, including being able to paint our ethnicity on our chromosomes as well as perform segment matching by ethnicity.
For example, if I carry an African segment on chromosome 1 from my father, and I match one person from my mother’s side and one from my father’s side on that same segment – one or the other of those people should also have that segment identified as African. That information would inform me as to which match is paternal and which is maternal
Not only that, this feature would help immensely tracking ancestors back in time and identifying their origins.
Will we ever get there? I don’t know. I’m not sure ethnicity is or can be accurate enough. We’ll see.
Transition to Digital and Online
Sometimes the future drags us kicking and screaming from the present.
With the imposed isolation of 2020, conferences quickly moved to an online presence. The genealogy community has all pulled together to make this work. The joke is that 2020’s most used phrase is “can you hear me?” I can vouch for that.
Of course while the year 2020 is over, the problem isn’t and is extending at least through the first half of 2021 and possibly longer. Conferences are planned months, up to a year, in advance and they can’t turn on a dime, so don’t even begin to expect in-person conferences until either late in 2021 or more likely, 2022 if all goes well this year.
I expect the future will eventually return to in-person conferences, but not entirely.
Finding ways to be more inclusive allows people who don’t want to or can’t travel or join in-person to participate.
I’ve recorded several sessions this year, mostly for 2021. Trust me, these could be a comedy, mostly of errors😊
I participated in four MyHeritage Facebook LIVE sessions in 2020 along with some other amazing speakers. This is what “live” events look like today!

Screenshot courtesy MyHeritage
A few days ago, I asked MyHeritage for a list of their LIVE sessions in 2020 and was shocked to learn that there were more than 90 in English, all free, and you can watch them anytime. Here’s the MyHeritage list.
By the way, every single one of the speakers is a volunteer, so say a big thank you to the speakers who make this possible, and to MyHeritage for the resources to make this free for everyone. If you’ve ever tried to coordinate anything like this, it’s anything but easy.
Additonally, I’ve created two Webinars this year for Legacy Family Tree Webinars.
Geoff Rasmussen put together the list of their top webinars for 2020, and I was pleased to see that I made the top 10! I’m sure there are MANY MORE you’d be interested in watching. Personally, I’m going to watch #6 yet today! Also, #9 and #22. You can always watch new webinars for free for a few days, and you can subscribe to watch all webinars, here.
The 2021 list of webinar speakers has been announced here, and while I’m not allowed to talk about something really fun that’s upcoming, let’s just say you definitely have something to look forward to in the springtime!
Also, don’t forget to register for RootsTech Connect which is entirely online and completely free, February 25-27, here.

Thank you to Penny Walters for creating this lovely graphic.
There are literally hundreds of speakers providing sessions in many languages for viewers around the world. I’ve heard the stats, but we can’t share them yet. Let me just say that you will be SHOCKED at the magnitude and reach of this conference. I’m talking dumbstruck!
During one of our zoom calls, one of the organizers says it feels like we’re constructing the plane as we’re flying, and I can confirm his observation – but we are getting it done – together! All hands on deck.
I’ll be presenting an advanced session about triangulation as well as a mini-session in the FamilySearch DNA Resource Center about finding your mother’s ancestors. I’ll share more information as it’s released and I can.
Companies and Owners Come & Go
You probably didn’t even notice some of these 2020 changes. Aside from the death of Bryan Sykes (RIP Bryan,) the big news and the even bigger unknown is the acquisition of Ancestry by Blackstone. Recently the CEO, Margo Georgiadis announced that she was stepping down. The Ancestry Board of Directors has announced an external search for a new CEO. All I can say is that very high on the priority list should be someone who IS a genealogist and who understands how DNA applies to genealogy.
Other changes included:
In the future, as genealogy and DNA testing becomes ever more popular and even more of a commodity, company sales and acquisitions will become more commonplace.
Some Companies Reduced Services and Cut Staff
I understand this too, but it’s painful. The layoffs occurred before Covid, so they didn’t result from Covid-related sales reductions. Let’s hope we see renewed investment after the Covid mess is over.
In a move that may or may not be related to an attempt to cut costs, Ancestry removed 6 and 7 cM matches from their users, freeing up processing resources, hardware, and storage requirements and thereby reducing costs.
I’m not going to beat this dead horse, because Ancestry is clearly not going to move on this issue, nor on that of the much-requested chromosome browser.
Later in the year, 23andMe also removed matches and other features, although, to their credit, they have restored at least part of this functionality and have provided ethnicity updates to V3 and V4 kits which wasn’t initially planned.
It’s also worth noting that early in 2020, 23andMe laid off 100 people as sales declined. Since that time, 23andMe has increasingly pushed consumers to pay to retest on their V5 chip.
About the same time, Ancestry also cut their workforce by about 6%, or about 100 people, also citing a slowdown in the consumer testing market. Ancestry also added a health product.
I’m not sure if we’ve reached market saturation or are simply seeing a leveling off. I wrote about that in DNA Testing Sales Decline: Reason and Reasons.
Of course, the pandemic economy where many people are either unemployed or insecure about their future isn’t helping.
The various companies need some product diversity to survive downturns. 23andMe is focused on medical research with partners who pay 23andMe for the DNA data of customers who opt-in, as does Ancestry.
Both Ancestry and MyHeritage provide subscription services for genealogy records.
FamilyTreeDNA is part of a larger company, GenebyGene whose genetics labs do processing for other companies and medical facilities.
A huge thank you to both MyHeritage and FamilyTreeDNA for NOT reducing services to customers in 2020.
Scientific Research Still Critical & Pushes Frontiers
Now that DNA testing has become a commodity, it’s easy to lose track of the fact that DNA testing is still a scientific endeavor that requires research to continue to move forward.
I’m still passionate about research after 20 years – maybe even more so now because there’s so much promise.
Research bleeds over into the consumer marketplace where products are improved and new features created allowing us to better track and understand our ancestors through their DNA that we and our family members inherit.
Here are a few of the research articles I published in 2020. You might notice a theme here – ancient DNA. What we can learn now due to new processing techniques is absolutely amazing. Labs can share files and information, providing the ability to “reprocess” the data, not the DNA itself, as more information and expertise becomes available.
Of course, in addition to this research, the Million Mito Project team is hard at work rewriting the tree of womankind.
If you’d like to participate, all you need to do is to either purchase a full sequence mitochondrial DNA kit at FamilyTreeDNA, or upgrade to the full sequence if you tested at a lower level previously.
Predictions

Predictions are risky business, but let me give it a shot.
Looking back a year, Covid wasn’t on the radar.
Looking back 5 years, neither Genetic Affairs nor DNAPainter were yet on the scene. DNAAdoption had just been formed in 2014 and DNAGedcom which was born out of DNAAdoption didn’t yet exist.
In other words, the most popular tools today didn’t exist yet.
GEDmatch, founded in 2010 by genealogists for genealogists was 5 years old, but was sold in December 2019 to Verogen.
We were begging Ancestry for a chromosome browser, and while we’ve pretty much given up beating them, because the horse is dead and they can sell DNA kits through ads focused elsewhere, that doesn’t mean genealogists still don’t need/want chromosome and segment based tools. Why, you’d think that Ancestry really doesn’t want us to break through those brick walls. That would be very bizarre, because every brick wall that falls reveals two more ancestors that need to be researched and spurs a frantic flurry of midnight searching. If you’re laughing right now, you know exactly what I mean!
Of course, if Ancestry provided a chromosome browser, it would cost development money for no additional revenue and their customer service reps would have to be able to support it. So from Ancestry’s perspective, there’s no good reason to provide us with that tool when they can sell kits without it. (Sigh.)
I’m not surprised by the management shift at Ancestry, and I wouldn’t be surprised to see several big players go public in the next decade, if not the next five years.
As companies increase in value, the number of private individuals who could afford to purchase the company decreases quickly, leaving private corporations as the only potential buyers, or becoming publicly held. Sometimes, that’s a good thing because investment dollars are infused into new product development.
What we desperately need, and I predict will happen one way or another is a marriage of individual tools and functions that exist separately today, with a dash of innovation. We need tools that will move beyond confirming existing ancestors – and will be able to identify ancestors through our DNA – out beyond each and every brick wall.

If a tester’s DNA matches to multiple people in a group descended from a particular previously unknown couple, and the timing and geography fits as well, that provides genealogical researchers with the hint they need to begin excavating the traditional records, looking for a connection.
In fact, this is exactly what happened with mitochondrial DNA – twice now. A match and a great deal of digging by one extremely persistent cousin resulting in identifying potential parents for a brick-wall ancestor. Autosomal DNA then confirmed that my DNA matched with 59 other individuals who descend from that couple through multiple children.
BUT, we couldn’t confirm those ancestors using autosomal DNA UNTIL WE HAD THE NAMES of the couple. DNA has the potential to reveal those names!
I wrote about that in Mitochondrial DNA Bulldozes Brick Wall and will be discussing it further in my RootsTech presentation.
The Challenge

We have most of the individual technology pieces today to get this done. Of course, the combined technological solution would require significant computing resources and processing power – just at the same time that vendors are desperately trying to pare costs to a minimum.
Some vendors simply aren’t interested, as I’ve already noted.
However, the winner, other than us genealogists, of course, will be the vendor who can either devise solutions or partner with others to create the right mix of tools that will combine matching, triangulation, and trees of your matches to each other, even if you don’t’ share a common ancestor.
We need to follow the DNA past the current end of the branch of our tree.

Each triangulated segment has an individual history that will lead not just to known ancestors, but to their unknown ancestors as well. We have reached critical mass in terms of how many people have tested – and more success would encourage more and more people to test.
There is a genetic path over every single brick wall in our genealogy.
Yes, I know that’s a bold statement. It’s not future Jetson’s flying-cars stuff. It’s doable – but it’s a matter of commitment, investment money, and finding a way to recoup that investment.
I don’t think it’s possible for the one-time purchase of a $39-$99 DNA test, especially when it’s not a loss-leader for something else like a records or data subscription (MyHeritage and Ancestry) or a medical research partnership (Ancestry and 23andMe.)
We’re performing these analysis processes manually and piecemeal today. It’s extremely inefficient and labor-intensive – which is why it often fails. People give up. And the process is painful, even when it does succeed.
This process has also been made increasingly difficult when some vendors block tools that help genealogists by downloading match and ancestral tree information. Before Ancestry closed access, I was creating theories based on common ancestors in my matches trees that weren’t in mine – then testing those theories both genetically (clusters, AutoTrees and ThruLines) and also by digging into traditional records to search for the genetic connection.
For example, I’m desperate to identify the parents of my James Lee Clarkson/Claxton, so I sorted my spreadsheet by surname and began evaluating everyone who had a Clarkson/Claxton in their tree in the 1700s in Virginia or North Carolina. But I can’t do that anymore now, either with a third-party tool or directly at Ancestry. Twenty million DNA kits sold for a minimum of $79 equals more than 1.5 billion dollars. Obviously, the issue here is not a lack of funds.
Including Y and mitochondrial DNA resources in our genetic toolbox not only confirms accuracy but also provides additional hints and clues.
Sometimes we start with Y DNA or mitochondrial DNA, and wind up using autosomal and sometimes the reverse. These are not competing products. It’s not either/or – it’s *and*.
Personally, I don’t expect the vendors to provide this game-changing complex functionality for free. I would be glad to pay for a subscription for top-of-the-line innovation and tools. In what other industry do consumers expect to pay for an item once and receive constant life-long innovations and upgrades? That doesn’t happen with software, phones nor with automobiles. I want vendors to be profitable so that they can invest in new tools that leverage the power of computing for genealogists to solve currently unsolvable problems.
Every single end-of-line ancestor in your tree represents a brick wall you need to overcome.
If you compare the cost of books, library visits, courthouse trips, and other research endeavors that often produce exactly nothing, these types of genetic tools would be both a godsend and an incredible value.
That’s it.
That’s the challenge, a gauntlet of sorts.
Who’s going to pick it up?
I can’t answer that question, but I can say that 23andMe can’t do this without supporting extensive trees, and Ancestry has shown absolutely no inclination to support segment data. You can’t achieve this goal without segment information or without trees.
Among the current players, that leaves two DNA testing companies and a few top-notch third parties as candidates – although – as the past has proven, the future is uncertain, fluid, and everchanging.
It will be interesting to see what I’m writing at the end of 2025, or maybe even at the end of 2021.
Stay tuned.
_____________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Products and Services
Genealogy Research
Books