I love all the various trees at FamilyTreeDNA – and I’m not referring just to traditional genealogy trees with people, names, and dates. I’m talking about phylogenetic or haplogroup trees – the ones you use to understand your Y-DNA and mitochondrial DNA haplogroups, origins – and more. These trees tell you ABOUT your ancestors, those people in the more traditional genealogy tree, and the combination of both is powerful.
This article introduces the various trees available at FamilyTreeDNA, when and where you’ll find them, and what they can do for you.
Haplogroup Trees
Phylogenetic, or haplogroup trees, provide a genetic path from you, or the tester, today, back in time to Y-Line Adam, or Mitochondrial Eve – the first two humans who lived AND have descendants today.
Let’s start by explaining about Y-DNA and mitochondrial DNA (mtDNA), their inheritance path, and what they mean to you.
Y-DNA
Only men have a Y-chromosome, so only biological males can test their Y-DNA.
Y-Line Adam, Y-DNA haplogroup A-PR2921, lived about 232,000 BCE, or 234,000 years ago.
Is it possible that one day someone will test whose results push that date back somewhat? Yes, of course, as we are always learning, and many testers split branches.
Today, all 711,000+ modern descendants who have tested carry the mutation named A-PR2921 as their oldest SNP (single nucleotide polymorphism), or haplogroup-defining mutation in their Y-DNA. That’s because we all descend from that one man.
If you’re a male, Y-DNA testing tells you about your direct paternal line by matching with other men who have also taken a Y-DNA test, and by revealing valuable information from before the adoption of surnames. There’s no other way to reach that far back in time.
If you’re a female, you can recruit males in your family to test.
The Big Y-700 test provides the deepest-reaching and most refined Y-DNA test available, which is essential for both genealogy and tree-building.
Mitochondrial DNA
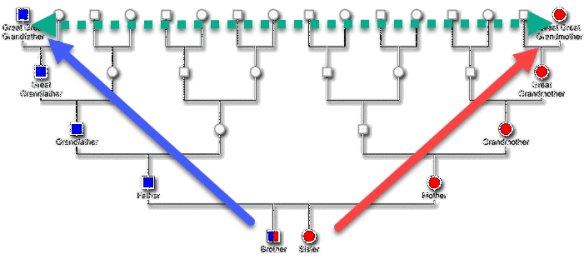
All people have mitochondrial DNA, inherited from their mother directly through her matrilineal line – meaning her mother, her mother, her mother, and so forth – directly up your tree through all mothers.
Everyone inherits their mitochondrial DNA (mtDNA) from their mother, but only females pass it on. Both males and females in the current generation, meaning you, can (and should) test their mitochondrial DNA.
Mitochondrial Eve, mitochondrial DNA haplogroup L, lived about 141,000 BCE, or about 143,000 years ago. All 315,000 testers descend from this one woman.
Like with Y-Line Adam, one day the results of future testers may push this date further back in time. A full sequence mitochondrial DNA test, mtFull, is necessary to test all 16,569 mitochondrial locations.
Test Types
FamilyTreeDNA has been in business for more than 25 years. Technology has advanced dramatically during that time. While they continue to offer new tests and products, they strive to maintain value for their original testers.
Even though some early testers may have joined their ancestors, matching with their test results is still beneficial to us.
Present-day DNA testers can still derive value by matching the earlier, lower-level, lower-resolution tests. Not as much value as if the original tester had taken a higher-level test, but those tests may not have been available at that time.
Matches, surnames, genealogy, locations, and haplogroups provide us with valuable information. The more people who test, the larger the pool becomes, and the better our chances of discovering something that refines our understanding of our ancestors – and identifies who they are.
Before we look at the trees available, let’s take a look at where haplogroups come from. Different level tests assign different levels of haplogroups, based on how much is tested.
Let’s answer two common questions:
- Where can you find your haplogroup, and what does it mean?
- How can haplogroups be different for people who descend from the same ancestor?
Where Do Haplogroups Come From?
Since the beginning, FamilyTreeDNA has always provided their customers with haplogroup information. Haplogroups are very genealogically useful today, but initially, 25 years ago, they were only able to provide essentially continental-level origin information for your particular line. That too was useful, and helped to identify and eliminate common lineages – just not as useful as today.
Science and testing have both come a long way. Present-day testers still match with people who only tested at a lower level. You never know what you might find at that level – a match to someone who has not taken the current tests, but is still very relevant because they share your ancestor. In fact, they may be the only tester who does.
For Y-DNA testers, you’ll notice several match categories that reflect different testing levels – along with the number of matches at each level. At one time, you could purchase each one of these tests individually, then later upgrade to higher-level tests. Today, only the 37 and 111 marker tests, and the Big Y-700, which scans the entire gold-standard region of the Y chromosome, are available. Higher level tests include the lower-level tests.
Different types of tests provide either a predicted or a confirmed haplogroup which shows on your match list.
Without getting all sciency on you – the 12-111 marker tests test targeted STRs, or short tandem repeats, which can’t be used for haplogroup assignment and confirmation. They can and are used to compare to other testers for matching because the number of repeats, or stutters, are inherited on the Y chromosome. The Big Y test scans the Y chromosome for SNPs, single nucleotide polymorphisms, which are stable mutations that define haplogroups. I wrote about this in the article, STRs vs SNPs, Multiple DNA Personalities.
Some haplogroups are much further down the tree, or more current, than others. Your most current haplogroup, only available with the Big Y-700 test, is the best because it brings you the closest to current in time, often placing you within family branches. The Big Y-700 scans about 23 million locations on the Y chromosome, revealing both known and unknown mutations, not just a few markers, making it the most refined and relevant test genealogically.
Each higher-level test includes the lower-level tests. You can see what tests your matches have taken by looking beneath their names on your match list. In this case, these Estes men who match my cousin have taken the Family Finder (or uploaded an autosomal transfer), and taken the mtFull test. One match initially took the Big Y-500 but has since upgraded to the Big Y-700, and the other originally tested at the 111 marker level, and has since upgraded as well.
The Big Y-700 includes all lower-level tests, such as the Big Y-500 (now obsolete), the 111, 67, 37, 25, and 12 marker STR tests. You still match with people who only tested at those levels, plus everyone else who ordered a more refined test.
The haplogroup you receive is more or less refined, based on the test level you take.
| Y-DNA Test Type | Haplogroup Provided | Relevance | Upgradable |
| Y-DNA STR 12-111 marker tests (only 37 and 111 are available today – the rest are obsolete) | Predicted based on STRs – very reliable at the level predicted | Predicted (not confirmed) haplogroup that was generally formed a couple thousand years ago, or earlier | Yes, if enough quality DNA remains. Only 37, 111, and the Big Y-700 tests are available today. Recommend the upgrade to Big Y-700. |
| Individual SNP test (now obsolete) | Confirms a predicted haplogroup or tests a single SNP to confirm a closer haplogroup | Relevant at the level tested – either positive or negative result was reported | Individual SNP tests have now been replaced by Big Y-700, which covers all individual SNPs that were available to test, plus much more. |
| Big Y-500 test (now obsolete) | Confirmed haplogroup within range of that test’s ability, replaced by much more granular Big Y-700 | Big Y-700 is more refined and moves the tester towards more current haplogroups, so more genealogically significant | Yes, upgrade to Big Y-700 if enough DNA remains, or tester can re-swab |
| Big Y-700 – scans the entire gold-standard region of the Y chromosome – approximately 23 million base pairs | Top-of-the-line SNP-confirmed test, most granular and refined. Scans for known and previously unknown mutations. Extremely accurate. | Generally advances the tester into a genealogical timeframe, and often divides testers into multiple lineages descended from a known common ancestor | No more advanced test is available. |
| Family Finder autosomal test or transfer | Confirmed to mid-range level if possible. Not all transfer files have Y-DNA or mtDNA SNPs so you get what you get. | Useful in autosomal matching for locating people you may be related to you with that surname. | Ask the match if they are willing to take a Y-DNA test, if relevant, or sponsor a testing scholarship for them. |
Family Finder haplogroups are relatively new at FamilyTreeDNA. Each chip level that FamilyTreeDNA has used for testing over the years, and the chips that other vendors have used, contain different SNPs (or none at all on the Ancestry test) that can be measured for some level of haplogroup. Other vendors generally don’t quality-control for either Y-DNA or mtDNA SNPs because they don’t use them. This is a “you get what you get” freebie.
That said, most Family Finder haplogroups are closer in time, or “better” than the predicted R-M269, the most common haplogroup in Europe, often reported with STR testing.
Not everyone with a transfer kit receives a haplogroup. Due to quality and reliability issues, you cannot see haplogroups on your autosomal match list for those who only have a haplogroup through an autosomal transfer.
Using our male Estes testers as an example, we find the following haplogroup results at the various testing levels:
| Haplogroup | Haplogroup Formation Date | Ancestor or Haplogroup Formation Location | Haplogroup Source |
| R-M269 | 4450 BCE (6450 years ago) | Between Ukraine and Kazakhstan, north of the Black and Caspian Seas | Predicted from 12-111 STR marker tests |
| R-BY487 | 700 CE (1300 years ago) | UK, Scotland/England | Family Finder DNA SNP Confirmed |
| R-BY482 | 1550 CE | Robert Eastye b 1555 Ringwould, Kent, England | Big Y-700 |
| R-BY490 | 1700 CE | Silvester Eastye b 1596 Kent, England | Big Y-700 |
| R-ZS3700 | 1750 CE | Moses Estes 1711 VA | Big Y-700 |
| R-BY154784 | 1850 CE | Joseph Estes b c 1790 VA or TN | Big Y-700 |
All of these are valid and accurate haplogroups – some are just closer in time and much more useful than others. All of these men have R-M269, because it is a parent haplogroup of all of those downstream haplogroups. The Big-Y tested men beginning with R-BY482 don’t share the haplogroups below them, because they don’t have those mutations that are downstream on the tree. However, the men at the bottom with R-BY154784 have all of the SNPs above them.
Note that all haplogroup formation dates are ranges. I’m showing the midpoint here.
When upgrading, if the original tester is deceased, select the highest-level test available, as there may not be enough DNA to run more than one test. When I offer scholarships now, I always just offer the Big Y-700 test to avoid future issues.
If the tester you need is no longer available, consider the possibility that other people, family members perhaps, might be available to test to represent this same line.
Next, let’s look at mitochondrial test levels and haplogroups.
| Mitochondrial DNA Test Type | Haplogroup Provided | Relevance | Upgradable |
| HVR1 & HVR2 tests (no longer available) | Predicted based on around 1000 markers – very reliable at the level predicted | Predicted haplogroup, not confirmed, generally formed a couple thousand years ago or earlier | Yes, if enough quality DNA remains. Only the mtFull test is available today. |
| mtFull, full sequence test | Tests all 16,569 SNP locations in the entire mitochondria. Most granular and refined. Extremely accurate. | Often brings tester into genealogical timeframe, especially with the new Mitotree. Divides testers into multiple haplotype lineages, sometimes descended from known common ancestor. | No upgrade needed to receive new Mitotree and mtDNA Discover benefits. |
| Family Finder autosomal test or transfer | Coming soon. Will be the same criteria and caveats as Y-DNA SNPs. | May be able to find a similar or upstream haplogroup that might point to a common ancestor. | Ask autosomal match if they are willing to take a mtFull test, if relevant, or sponsor a scholarship for them. |
Ok, now that we understand more about haplogroups, how they are determined, and where yours came from, let’s look at all of the trees at FamilyTreeDNA.
Trees Within Your Y-DNA and Mitochondrial DNA Account
Let’s start with trees found within your personal account, so sign in.
Each tree has a different purpose and unique benefits.
Tree #1 – Your Matches Genealogy Trees
Each of your matches may have provided links to genealogical trees. They may show trees in multiple places too; at MyHeritage, an archived tree at FamilyTreeDNA, and a WikiTree link. I makes notes about their trees in the comments field, and I also keep a spreadsheet to look for commonalities.
Tree #2 – Haplogroups and SNPs for Y-DNA Testers
Next, for Y-DNA testers, click on the Y-DNA Results and Tools.
You’ll see the Haplotree & SNPs tile on the dashboard.
The Haplotree and SNPs link takes you to a phylogenetic tree that defaults to your haplogroup, where you can view:
- Variants – SNP mutations that define your haplogroup
- Surnames with this haplogroup – so long as there are multiple public testers
- Countries – self-reported for earliest known ancestors (EKA)
- Recommended Projects – haplogroup projects only – others such as surname projects are found in Discover under Suggested Projects
Tree #3 – The Block Tree for Big Y Testers
People who have taken the Big Y-700 test have a separate section that includes tools for the Big-Y test that aren’t relevant for the 12-111 STR marker tests.
Big Y testers will see the Block Tree tile on their dashboard.
The block tree is an alternative way of displaying matches on a phylogenetic tree. While the Discover Time Tree is viewed left to right, this tree is displayed top to bottom, with each mutation being represented by one grey bar on the scale at left. Each mutation corresponds to approximately 100 years, which is a rough average for the frequency of Y-chromosomal mutations.
People with 30 mutations or fewer are shown as matches, with the goal of reaching back about 1500 years.
Each large block shows the mutation for which the haplogroup is named, such as R-BY482, at the top. The mutations, known as variants, shown below that haplogroup name, are found in the results of each person in that haplogroup, but in the future, people without those mutations, or with additional mutations, will form a new branching haplogroup.
The green “Private Variants” at the bottom of the branches display the average number of mutations of people within that group awaiting another tester to have the same mutations, so a new branch can be formed. I view Private Mutations as “haplogroups in waiting.”
Discover
In addition to the haplogroup trees shown in your account at FamilyTreeDNA, there are several additional trees in Discover for both Y-DNA and mitochondrial DNA. Discover, updated weekly, is a suite of tools for both Y-DNA and mitochondrial DNA that, cumulatively, provides a book about your haplogroup results.
Discover comes in two flavors:
- The publicly available free version with limited functionality
- Your private version with expanded functionality available from within your account
You can access Discover, here if you’d like to follow along.
Discover is a publicly available free tool introduced in the fall of 2023 that provides more than a dozen reports, enabling a deeper understanding of all haplogroups.
Just select Y-DNA or mtDNA and enter your haplogroup of choice.
Think of these menu choices, in the sidebar, as chapters in your personal book. Every chapter has something interesting to tell you. Please read them – don’t just scan.
In addition to the free version, if you have taken a Big-Y or mitochondrial DNA full sequence test at FamilyTreeDNA, you’ll have additional information available.
For mitochondrial DNA results, just click on the pink Discover tile.
For Y-DNA results, click on the blue Discover tile.
Within Discover, you’ll find three distinct trees.
Trees #4 and #5 – Y-DNA and Mitochondrial DNA Time Trees
The Time Tree shows your Y-DNA or mitochondrial DNA haplogroup displayed on a timeline, along with:
- A self-reported ancestral country indicator for every person’s DNA in that haplogroup
- Haplotype groupings indicating exact matches between everyone in that haplotype.
A haplotype is a grouping of people whose DNA matches exactly, including unstable or hypervariable locations too unreliable to use for haplogroup formation. However, those mutations may be relevant for genealogical matching.
I wrote about haplogroups and haplotypes here and here.
Tree #6 and #7 – Y-DNA and Mitochondrial DNA Class Tree View
The Classic Tree is available for both Y-DNA and mitochondrial DNA.
On the Classic Mitotree View, you can display and filter the tree, including haplotypes, in seven ways, as shown in the dropdown “Display Options.”
Tree #8 and #9 – Y-DNA and Mitochondrial DNA Tree Branch Comparison
Have you ever seen two haplogroups and wondered how closely they are related? Compare provides that answer.
Here, I’m comparing my haplogroup to that of a family member. Everyone is related, but how long ago are we related on our matrilineal lines?
Haplogroup J1c2f compared with haplogroup V216a shows that our common ancestor lived a VERY long time ago – about 55,000 years in the past, someplace in the fertile crescent.
For either Y-DNA or mitochondrial DNA, you can compare two haplogroups. This provides specific information about those two branches of the tree, and where they intersect. To view more about the common ancestor, just pop R+10398 into Discover and learn more about when and where that ancestor lived.
Trees #10 and #11 – Match Time Trees
Match Time Trees are one of the most useful Discover features.
In addition to the Time Trees and Classic Trees provided for everyone in Discover, test takers will also have a Match Time Tree that shows all of your matches, organized genetically.
For mtFull testers, your matches are organized by haplotype cluster. People in your haplotype cluster are your exact matches.
I have over 100 full sequence matches, so I’m only showing the first few in this screenshot. In addition to the match’s name, their EKA (earliest known ancestor) is shown, if provided.
On the Y-DNA Match Time Tree, links are provided to genealogical trees of the tester, which could be an archived FamilyTreeDNA tree, a MyHeritage tree, WikiTree, or some combination.
You can actually see your matches’ WikiTree tree on your Match Time Tree by enabling another feature.
Trees #12 and #13 – WikiTree Tree Integration
While you’re still on the Match Time Tree page for either Y-DNA or mitochondrial DNA, click on Display Options, above the Time Tree, and enable WikiTree Connections. Unfortunately, the default for this great feature is “off.”
I’ve enabled “Share Mode” at the top to obfuscate the names of the testers, and I’ve adjusted the vertical spacing so you can see more in my examples. You’ll notice the grey lines with dots inside circles. I think of these as beads or maybe knots on a rope, but they actually represent a line of ancestors.
Each tester with one of those grey dot bars has connected themselves to their ancestors at WikiTree, a public one-world tree. Living people are not shown, hence the dash marks to the immediate left of the tester’s name.
By mousing over any of the dots, aka ancestors, you can view information about this ancestor of this Estes tester at WikiTree. Ancestors appear in genealogical order in their relevant place on the Time Tree. How cool is that!!!
WikiTree, like any tree, public or private, can have errors. Always verify any tree using original source documents.
As far as I’m concerned, the Match Time Tree is one of the very best features of both Y-DNA and mitochondrial DNA testing and matching. There are so many options to select from, so take some time to look around.
Your Personal Version of Discover is Best
Y-DNA Discover and mtDNA Discover can both be useful for any level of haplogroup, but the best results are obtained when clicking through from the tester’s FamilyTreeDNA account. Big Y and full sequence mitochondrial DNA customers receive additional information, not available in the free, public version of Discover, including
- The Match Time Tree
- Including WikiTree integration
- Globetrekker (Y-DNA, mtDNA coming eventually)
- Up to 30 Ancient Connections, as compared to 3 in the free version
- Up to 30 Notable Connections, as compared to 3 in the free version
Tree #14 – Group Time Trees
I absolutely love Group Time Trees. They are similar to Match Time Trees, but unlike Match Time Trees, are publicly viewable for Group Projects if the volunteer project administrators have enabled this feature for the project.
There are two ways to access Group Time Trees – through publicly accessible Discover or directly through any project.
In Discover, select Group Project in the dropdown.
Then type the name of the surname project you’re seeking. You’ll be presented with a menu if the surname you’ve entered is found in multiple projects, or administrators have listed it as “of interest” in their project.
I clicked on the Estes project.
Viewing the Estes DNA Project, under DNA Results, you can see the various options.
Selecting Y-DNA Results Overview displays the project results by administrator-defined group. The teal groups all descend through Abraham Estes through various sons.
However, by clicking the Group Time Tree instead, you can view all these testers and their results in a Match Time Tree format, arranged genetically.
Clicking on the Group Time Tree link takes you to the Group Time Tree for this project. A menu is displayed at left, based on how the administrator has grouped the project.
I’ve selected several groups that I know descend from the original Estes ancestor from Kent, England. Testers who have joined the Estes project and granted permission for their results to be displayed publicly are automatically grouped genetically, at right, with their surname and EKA (earliest known ancestor), assuming they have entered that information.
Earliest Known Ancestors (EKA)
You’ve probably noticed that earliest known ancestors, along with their locations, are used in many places.
Please enter both your direct paternal (father, father, to father’s line) and direct matrilineal (mother, mother, to mother’s line) earliest known ancestors, along with their locations. I wrote about how to do that in “Earliest Known Ancestors” at FamilyTreeDNA in 3 Easy Steps, here.
Trees #15 and #16 – Public Trees
In addition to trees within testers’ accounts, Discover trees, Group Time Trees, and WikiTree tree integration, FamilyTreeDNA provides two additional public trees.
FamilyTreeDNA made the Y-DNA and mitochondrial DNA haplogroup trees freely available years ago, at the bottom of their main company public page – without signing in.
These trees are still actively maintained today and are free for everyone to use.
To find these trees, scroll all the way to the very bottom of the page, in the footer, to the Community section. Yes, I know, it’s a bit like a scavenger hunt!
You can select to view either the Y-DNA or mtDNA tree. I love this tree, because it shows how many SNP-confirmed people have been tested. That number does not include the thousands of academic and public samples that may be utilized to help define haplogroups, and that you’ll sometimes see in your Ancient and Notable Connections.
So, if you receive a new haplogroup, but you don’t see a new match on your list or on the Block Tree, it’s probably because you match a high-quality academic sample.
The trees display from the root, meaning the oldest haplogroup is shown at the top. In the Y-DNA tree, above, haplogroup A-PR2921 is “Y-Adam”.
You can select any haplogroup on the bar across the top, search by country, or select a specific branch name to view.
The tree itself is viewable by country, as shown above, or by variant, meaning the haplogroup-defining mutations, shown below.
Additionally, for the Y-DNA tree, you can choose to display by surname, so long as there are two or more testers with that identically spelled surname who share this haplogroup and who have given permission for public display.
Please note that these people are all SNP-tested and confirmed at the level reported, but they are NOT all Big-Y testers.
This feature alone can be genealogy-changing because they may be surnames associated with your ancestors in records, or they may just be neighbors. Or maybe you thought they were “just neighbors,” but they are actually related.
At one time, customers could order an individual SNP test for R-M269 to confirm their predicted haplogroup. That test is no longer available, but anyone who took that test to confirm R-M269 and never tested or received results (like Family Finder) at a more granular level will be reported at R-M269. Note that 687 is the number of distinct surnames shown, not the total number of testers.
The three “hamburger dots” on the right side provide options for a user-reported Country Report based on the location of their earliest known ancestor, and a Surname Report. The surname report for R-M269 shows a total of 2448 testers who share those 687 surnames.
It’s a Whole Forest
Who knew there were 16 unique trees available at FamilyTreeDNA!
Each tree has a unique purpose and provides information not available elsewhere.
Take a look and see what kind of information is waiting for you – and don’t forget to check back often.
_____________________________________________________________
Share the Love!
You’re always welcome to forward articles or links to friends and share on social media.
Subscribe!
If you haven’t already subscribed, it’s free. You’ll receive an e-mail whenever I publish by clicking the “follow” button at the top of the main blog page, here.
Help Keep This Blog Free
I receive a small commission when you click a vendor link in my articles and purchase that item. This does NOT increase your price but helps me keep the lights on and this informational blog free for everyone. Please click on the affiliate links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Uploads
- FamilyTreeDNA – Y-DNA, mitochondrial and autosomal DNA testing
- MyHeritage DNA – Autosomal DNA test
- AncestryDNA – Autosomal DNA test
- AncestryDNA Plus Traits
- 23andMe Ancestry – Autosomal DNA only, no Health
- 23andMe Ancestry Plus Health
Genealogy Products and Services
- MyHeritage Subscription with Free Trial
- Legacy Family Tree Webinars – Genealogy and DNA classes, subscription-based, some free
- Legacy Family Tree Software – Genealogy software for your computer
- OldNews – Old Newspapers with links to save to MyHeritage trees
- MyHeritage Omni comprehensive “everything included” subscription plan
- Newspapers.com – Search newspapers for your ancestors
- NewspaperArchive – Search different newspapers for your ancestors
My Books
- DNA for Native American Genealogy – by Roberta Estes, for those ordering the e-book from anyplace, or paperback within the United States
- DNA for Native American Genealogy – for those ordering the paperback outside the US
- The Complete Guide to FamilyTreeDNA – Y-DNA, Mitochondrial, Autosomal and X-DNA – for those ordering the e-book from anyplace, or paperback within the United States
- The Complete Guide to FamilyTreeDNA – Y-DNA, Mitochondrial, Autosomal and X-DNA for those ordering the paperback from outside the US
Genealogy Books
- Genealogical.com – Lots of wonderful genealogy research books
- American Ancestors – Wonderful selection of genealogy books
Genealogy Research
- Legacy Tree Genealogists – Professional genealogy research