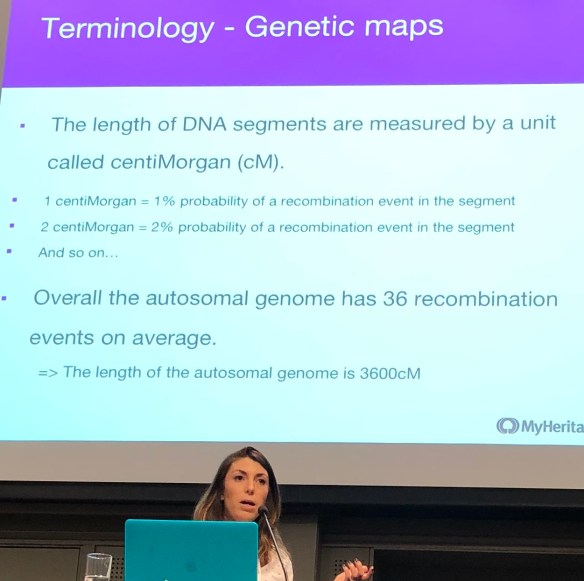
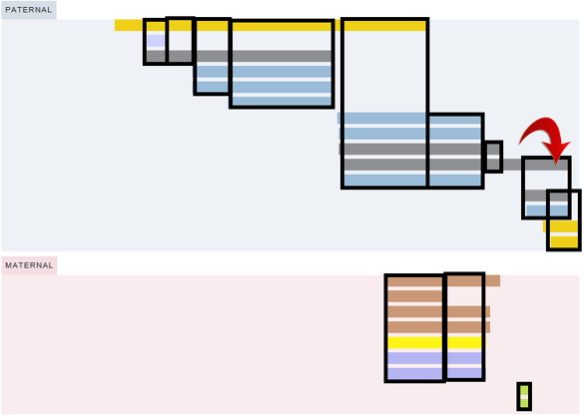
The official dates of RootsTech 2022 were March 3-5, but the sessions and content in the vendor booths are still available. I’ve compiled a list of the sessions focused on DNA, with web links on the RootsTech YouTube channel
YouTube reports the number of views, so I was able to compile that information as of March 8, 2022.
I do want to explain a couple of things to add context to the numbers.
Most speakers recorded their sessions, but a few offered live sessions which were recorded, then posted later for participants to view. However, there have been glitches in that process. While the sessions were anticipated to be available an hour or so later, that didn’t quite happen, and a couple still aren’t posted. I’m sure the presenters are distressed by this, so be sure to watch those when they are up and running.
The Zoom rooms where participants gathered for the live sessions were restricted to 500 attendees. The YouTube number of views does not include the number of live viewers, so you’ll need to add an additional number, up to 500.
When you see a number before the session name, whether recorded or live, that means that the session is part of a series. RootsTech required speakers to divide longer sessions into a series of shorter sessions no longer than 15-20 minutes each. The goal was for viewers to be able to watch the sessions one after the other, as one class, or separately, and still make sense of the content. Let’s just say this was the most challenging thing I’ve ever done as a presenter.
For recorded series sessions, these are posted as 1, 2 and 3, as you can see below with Diahan Southard’s sessions. However, with my live session series, that didn’t happen. It looks like my sessions are a series, but when you watch them, parts 1, 2 and 3 are recorded and presented as one session. Personally, I’m fine with this, because I think the information makes a lot more sense this way. However, it makes comparisons difficult.
This was only the second year for RootsTech to be virtual and the conference is absolutely HUGE, so live and learn. Next year will be smoother and hopefully, at least partially in-person too.
When I “arrived” to present my live session, “Associating Autosomal DNA Segments With Ancestors,” my lovely moderator, Rhett, told me that they were going to livestream my session to the RootsTech page on Facebook as well because they realized that the 500 Zoom seat limit had been a problem the day before with some popular sessions. I have about 9000 views for that session and more than 7,400 of them are on the RootsTech Facebook page – and that was WITHOUT any advance notice or advertising. I know that the Zoom room was full in addition. I felt kind of strange about including my results in the top ten because I had that advantage, but I didn’t know quite how to otherwise count my session. As it turns out, all sessions with more than 1000 views made it into the top ten so mine would have been there one way or another. A big thank you to everyone who watched!
I hope that the RootsTech team notices that the most viewed session is the one that was NOT constrained by the 500-seat limited AND was live-streamed on Facebook. Seems like this might be a great way to increase session views for everyone next year. Hint, hint!!!
I also want to say a huge thank you to all of the presenters for producing outstanding content. The sessions were challenging to find, plus RootsTech is always hectic, even virtually. So, I know a LOT of people will want to view these informative sessions, now that you know where to look and have more time. Please remember to “like” the session on YouTube as a way of thanking your presenter.
With 140 DNA-focused sessions available, you can watch a new session, and put it to use, every other day for the next year! How fun is that! You can use this article as your own playlist.
Please feel free to share this article with your friends and genealogy groups so everyone can learn more about using DNA for genealogy.
Ok, let’s look at the top 10. Drum roll please…
Top 10 Most Viewed RootsTech Sessions
| Session Title | Presenter | YouTube Link | Views | |
| 1 | 1. Associating Autosomal DNA Segments With Ancestors | Roberta Estes (live) | https://www.youtube.com/watch?v=_IHSCkNnX48
|
~9000: 1019 + 500 live viewers + 7,400+ Facebook |
| 2 | 1. What to Do with Your DNA Test Results in 2022 (part 1 of 3) | Diahan Southard | https://www.youtube.com/watch?v=FENAKAYLXX4 | 7428 |
| 3 | Who Is FamilyTreeDNA? | FamilyTreeDNA – Bennett Greenspan | https://www.youtube.com/watch?v=MHFtwoatJ-A | 2946 |
| 4 | 2. What to Do with Your DNA Test Results in 2022 (part 2 of 3) | Diahan Southard | https://www.youtube.com/watch?v=mIllhtONhlI | 2448 |
| 5 | Latest DNA Painter Releases | DNAPainter Jonny Perl (live) | https://www.youtube.com/watch?v=iLBThU8l33o | 2230 + live viewers |
| 6 | DNA Painter Introduction | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=Rpe5LMPNmf0 | 1983 |
| 7 | 3. What to Do with Your DNA Test Results in 2022 (part 3 of 3) | Diahan Southard | https://www.youtube.com/watch?v=hemY5TuLmGI | 1780 |
| 8 | The Tree of Mankind Age Estimates | Paul Maier | https://www.youtube.com/watch?v=jjkL8PWAEwk | 1638 |
| 9 | A Sneak Peek at FamilyTreeDNA Coming Attractions | FamilyTreeDNA (live) | https://www.youtube.com/watch?v=K9sKqNScvnE | 1270 + live viewers
|
| 10 | Extending Time Horizons with DNA | Rob Spencer (live) | https://www.youtube.com/watch?v=wppXD1Zz2sQ | 1037 + live viewers |
All DNA-Focused Sessions
I know you’ll find LOTS of goodies here. Which ones are your favorites?
| Session | Presenter | YouTube Link | Views | |
| 1 | Estimating Relationships by Combining DNA from Multiple Siblings | Amy Williams | https://www.youtube.com/watch?v=xs1U0ohpKSA | 201 |
| 2 | Overview of HAPI-DNA.org | Amy Williams | https://www.youtube.com/watch?v=FjNiJgWaBeQ | 126 |
| 3 | How do AncestryDNA® Communities help tell your story? | Ancestry® | Ancestry | https://www.youtube.com/watch?v=EQNpUxonQO4 | 183
|
| 4 | AncestryDNA® 201 | Ancestry – Crista Cowan | https://www.youtube.com/watch?v=lbqpnXloM5s
|
494 |
| 5 | Genealogy in a Minute: Increase Discoveries by Attaching AncestryDNA® Results to Family Tree | Ancestry – Crista Cowan | https://www.youtube.com/watch?v=iAqwSCO8Pvw | 369 |
| 6 | AncestryDNA® 101: Beginner’s Guide to AncestryDNA® | Ancestry® | Ancestry – Lisa Elzey | https://www.youtube.com/watch?v=-N2usCR86sY | 909 |
| 7 | Hidden in Plain Sight: Free People of Color in Your Family Tree | Cheri Daniels | https://www.youtube.com/watch?v=FUOcdhO3uDM | 179 |
| 8 | Finding Relatives to Prevent Hereditary Cancer | ConnectMyVariant – Dr. Brian Shirts | https://www.youtube.com/watch?v=LpwLGgEp2IE | 63 |
| 9 | Piling on the chromosomes | Debbie Kennett | https://www.youtube.com/watch?v=e14lMsS3rcY | 465 |
| 10 | Linking Families With Rare Genetic Condition Using Genealogy | Deborah Neklason | https://www.youtube.com/watch?v=b94lUfeAw9k | 43 |
| 11 | 1. What to Do with Your DNA Test Results in 2022 | Diahan Southard | https://www.youtube.com/watch?v=FENAKAYLXX4 | 7428 |
| 12 | 1. What to Do with Your DNA Test Results in 2022 | Diahan Southard | https://www.youtube.com/watch?v=hemY5TuLmGI | 1780 |
| 13 | 2. What to Do with Your DNA Test Results in 2022 | Diahan Southard | https://www.youtube.com/watch?v=mIllhtONhlI | 2448 |
| 14 | DNA Testing For Family History | Diahan Southard | https://www.youtube.com/watch?v=kCLuOCC924s | 84
|
| 15 | Understanding Your DNA Ethnicity Estimate at 23andMe | Diana Elder
|
https://www.youtube.com/watch?v=xT1OtyvbVHE | 66 |
| 16 | Understanding Your Ethnicity Estimate at FamilyTreeDNA | Diana Elder | https://www.youtube.com/watch?v=XosjViloVE0 | 73 |
| 17 | DNA Monkey Wrenches | Katherine Borges | https://www.youtube.com/watch?v=Thv79pmII5M | 245 |
| 18 | Advanced Features in your Ancestral Tree and Fan Chart | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=4u5Vf13ZoAc | 425 |
| 19 | DNA Painter Introduction | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=Rpe5LMPNmf0 | 1983 |
| 20 | Getting Segment Data from 23andMe DNA Matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=8EBRI85P3KQ | 134 |
| 21 | Getting segment data from FamilyTreeDNA DNA matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=rWnxK86a12U | 169 |
| 22 | Getting segment data from Gedmatch DNA matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=WF11HEL8Apk | 163 |
| 23 | Getting segment data from Geneanet DNA Matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=eclj8Ap0uK4 | 38 |
| 24 | Getting segment data from MyHeritage DNA matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=9rGwOtqbg5E | 160 |
| 25 | Inferred Chromosome Mapping: Maximize your DNA Matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=tzd5arHkv64 | 688 |
| 26 | Keeping track of your genetic family tree in a fan chart | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=W3Hcno7en94 | 806
|
| 27 | Mapping a DNA Match in a Chromosome Map | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=A61zQFBWaiY | 423 |
| 28 | Setting up an Ancestral Tree and Fan Chart and Exploring Tree Completeness | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=lkJp5Xk1thg | 77 |
| 29 | Using the Shared cM Project Tool to Evaluate DNA Matches | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=vxhn9l3Dxg4 | 763 |
| 30 | Your First Chromosome Map: Using your DNA Matches to Link Segments to Ancestors | DNAPainter – Jonny Perl | https://www.youtube.com/watch?v=tzd5arHkv64 | 688 |
| 31 | DNA Painter for absolute beginners | DNAPainter (Jonny Perl) | https://www.youtube.com/watch?v=JwUWW4WHwhk | 1196 |
| 32 | Latest DNA Painter Releases | DNAPainter (live) | https://www.youtube.com/watch?v=iLBThU8l33o | 2230 + live viewers |
| 33 | Unraveling your genealogy with DNA segment networks using AutoSegment from Genetic Affairs | Evert-Jan Blom | https://www.youtube.com/watch?v=rVpsJSqOJZI
|
162 |
| 34 | Unraveling your genealogy with genetic networks using AutoCluster | Evert-Jan Blom | https://www.youtube.com/watch?v=ZTKSz_X7_zs | 201
|
| 35 | Unraveling your genealogy with reconstructed trees using AutoTree & AutoKinship from Genetic Affairs | Evert-Jan Blom | https://www.youtube.com/watch?v=OmDQoAn9tVw | 143 |
| 36 | Research Like a Pro with DNA – A Genealogist’s Guide to Finding and Confirming Ancestors with DNA | Family Locket Genealogists | https://www.youtube.com/watch?v=NYpLscJJQyk | 183 |
| 37 | How to Interpret a DNA Network Graph | Family Locket Genealogists – Diana Elder | https://www.youtube.com/watch?v=i83WRl1uLWY | 393 |
| 38 | Find and Confirm Ancestors with DNA Evidence | Family Locket Genealogists – Nicole Dyer | https://www.youtube.com/watch?v=DGLpV3aNuZI | 144 |
| 39 | How To Make A DNA Network Graph | Family Locket Genealogists – Nicole Dyer | https://www.youtube.com/watch?v=MLm_dVK2kAA | 201 |
| 40 | Create A Family Tree With Your DNA Matches-Use Lucidchart To Create A Picture Worth A Thousand Words | Family Locket Genealogists – Robin Wirthlin | https://www.youtube.com/watch?v=RlRIzcW-JI4 | 270 |
| 41 | Charting Companion 7 – DNA Edition | Family Tree Maker | https://www.youtube.com/watch?v=k2r9rkk22nU | 316
|
| 42 | Family Finder Chromosome Browser: How to Use | FamilyTreeDNA | https://www.youtube.com/watch?v=w0_tgopBn_o | 750
|
| 43 | FamilyTreeDNA: 22 Years of Breaking Down Brick Walls | FamilyTreeDNA | https://www.familysearch.org/rootstech/session/familytreedna-22-years-of-breaking-down-brick-walls | Not available |
| 44 | Review of Autosomal DNA, Y-DNA, & mtDNA | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=EJoQVKxgaVY | 77 |
| 45 | Who Is FamilyTreeDNA? | FamilyTreeDNA – Bennett Greenspan | https://www.youtube.com/watch?v=MHFtwoatJ-A | 2946 |
| 46 | Part 1: How to Interpret Y-DNA Results, A Walk Through the Big Y | FamilyTreeDNA – Casimir Roman | https://www.youtube.com/watch?v=ra1cjGgvhRw | 684
|
| 47 | Part 2: How to Interpret Y-DNA Results, A Walk Through the Big Y | FamilyTreeDNA – Casimir Roman | https://www.youtube.com/watch?v=CgqcjBD6N8Y
|
259 |
| 48 | Big Y-700: A Brief Overview | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=IefUipZcLCQ | 96 |
| 49 | Mitochondrial DNA & The Million Mito Project | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=5Zppv2uAa6I | 179 |
| 50 | Mitochondrial DNA: What is a Heteroplasmy | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=ZeGTyUDKySk | 57 |
| 51 | Y-DNA Big Y: A Lifetime Analysis | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=E6NEU92rpiM | 154 |
| 52 | Y-DNA: How SNPs Are Added to the Y Haplotree | FamilyTreeDNA – Janine Cloud | https://www.youtube.com/watch?v=CGQaYcroRwY | 220 |
| 53 | Family Finder myOrigins: Beginner’s Guide | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=VrJNpSv8nlA | 88 |
| 54 | Mitochondrial DNA: Matches Map & Results for mtDNA | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=YtA1j01MOvs | 190 |
| 55 | Mitochondrial DNA: mtDNA Mutations Explained | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=awPs0cmZApE | 340
|
| 56 | Y-DNA: Haplotree and SNPs Page Overview | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=FOuVhoMD-hw | 432 |
| 57 | Y-DNA: Understanding the Y-STR Results Page | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=gCeZz1rQplI | 148 |
| 58 | Y-DNA: What Is Genetic Distance? | FamilyTreeDNA – Katy Rowe | https://www.youtube.com/watch?v=qJ6wY6ILhfg | 149 |
| 59 | DNA Tools: myOrigins 3.0 Explained, Part 1 | FamilyTreeDNA – Paul Maier | https://www.youtube.com/watch?v=ACgY3F4-w78 | 74
|
| 60 | DNA Tools: myOrigins 3.0 Explained, Part 2 | FamilyTreeDNA – Paul Maier | https://www.youtube.com/watch?v=h7qU36bIFg0 | 50 |
| 61 | DNA Tools: myOrigins 3.0 Explained, Part 3 | FamilyTreeDNA – Paul Maier | https://www.youtube.com/watch?v=SWlGPm8BGyU | 36 |
| 62 | African American Genealogy Research Tips | FamilyTreeDNA – Sherman McRae | https://www.youtube.com/watch?v=XdbkM58rXIQ | 153
|
| 63 | Connecting With My Ancestors Through Y-DNA | FamilyTreeDNA – Sherman McRae | https://www.youtube.com/watch?v=xbo1XnLkuQU | 200 |
| 64 | Join The Million Mito Project | FamilyTreeDNA (Join link) | https://www.familysearch.org/rootstech/session/join-the-million-mito-project | link |
| 65 | View the World’s Largest mtDNA Haplotree | FamilyTreeDNA (Link to mtDNA tree) | https://www.familytreedna.com/public/mt-dna-haplotree/L | n/a |
| 66 | View the World’s Largest Y Haplotree | FamilyTreeDNA (Link to Y tree) | https://www.familytreedna.com/public/y-dna-haplotree/A | link |
| 67 | A Sneak Peek at FamilyTreeDNA Coming Attractions | FamilyTreeDNA (live) | https://www.youtube.com/watch?v=K9sKqNScvnE | 1270 + live viewers
|
| 68 | DNA Upload: How to Transfer Your Autosomal DNA Data | FamilyTreeDNA -Katy Rowe | https://www.youtube.com/watch?v=CS-rH_HrGlo | 303 |
| 69 | Family Finder myOrigins: How to Compare Origins With Your DNA Matches | FamilyTreeDNA -Katy Rowe | https://www.youtube.com/watch?v=7mBmWhM4j9Y | 145 |
| 70 | Join Group Projects at FamilyTreeDNA | FamilyTreeDNA link to learning center article) | https://www.familysearch.org/rootstech/session/join-group-projects-at-familytreedna | link
|
| 71 | Product Demo – Unraveling your genealogy with reconstructed trees using AutoKinship | GEDmatch | https://www.youtube.com/watch?v=R7_W0FM5U7c | 803 |
| 72 | Towards a Genetic Genealogy Driven Irish Reference Genome | Gerard Corcoran | https://www.youtube.com/watch?v=6Kx8qeNiVmo | 155
|
| 73 | Discovering Biological Origins in Chile With DNA: Simple Triangulation | Gonzalo Alexis Luengo Orellana | https://www.youtube.com/watch?v=WcVby54Uigc | 40 |
| 74 | Cousin Lynne: An Adoption Story | International Association of Jewish Genealogical Societies | https://www.youtube.com/watch?v=AptMcV4_B4o | 111 |
| 75 | Using DNA Testing to Uncover Native Ancestry | Janine Cloud | https://www.youtube.com/watch?v=edzebJXepMA | 205 |
| 76 | 1. Forensic Genetic Genealogy | Jarrett Ross | https://www.youtube.com/watch?v=0euIDZTmx5g | 58 |
| 77 | Reunited and it Feels so Good | Jennifer Mendelsohn | https://www.youtube.com/watch?v=X-hxjm7grBE | 57
|
| 78 | Genealogical Research and DNA Testing: The Perfect Companions | Kimberly Brown | https://www.youtube.com/watch?v=X82jA3xUVXk | 80 |
| 79 | Finding a Jewish Sperm Donor | Kitty Munson Cooper | https://www.youtube.com/watch?v=iKRjFfNcpug | 164 |
| 80 | Using DNA in South African Genealogy | Linda Farrell | https://www.youtube.com/watch?v=HXkbBWmORM0 | 141 |
| 81 | Using DNA Group Projects In Your Family History Research | Mags Gaulden | https://www.youtube.com/watch?v=0tX7QDib4Cw | 165 |
| 82 | 2. The Expansion of Genealogy Into Forensics | Marybeth Sciaretta | https://www.youtube.com/watch?v=HcEO-rMe3Xo | 35
|
| 83 | DNA Interest Groups That Keep ’em Coming Back | McKell Keeney (live) | https://www.youtube.com/watch?v=HFwpmtA_QbE | 180 plus live viewers |
| 84 | Searching for Close Relatives with Your DNA Results | Mckell Keeney (live) | https://www.familysearch.org/rootstech/session/searching-for-close-relatives-with-your-dna-results | Not yet available |
| 85 | Top Ten Reasons To DNA Test For Family History | Michelle Leonard | https://www.youtube.com/watch?v=1B9hEeu_dic | 181 |
| 86 | Top Tips For Identifying DNA Matches | Michelle Leonard | https://www.youtube.com/watch?v=-3Oay_btNAI | 306 |
| 87 | Maximising Messages | Michelle Patient | https://www.youtube.com/watch?v=4TRmn0qzHik | 442 |
| 88 | How to Filter and Sort Your DNA Matches | MyHeritage | https://www.youtube.com/watch?v=fmIgamFDvc8 | 88 |
| 89 | How to Get Started with Your DNA Matches | MyHeritage | https://www.youtube.com/watch?v=JPOzhTxhU0E | 447
|
| 90 | How to Track DNA Kits in MyHeritage` | MyHeritage | https://www.youtube.com/watch?v=2W0zBbkBJ5w | 28
|
| 91 | How to Upload Your DNA Data to MyHeritage | MyHeritage | https://www.youtube.com/watch?v=nJ4RoZOQafY | 82 |
| 92 | How to Use Genetic Groups | MyHeritage | https://www.youtube.com/watch?v=PtDAUHN-3-4 | 62 |
| My Story: Hope | MyHeritage | https://www.youtube.com/watch?v=qjyggKZEXYA | 133 | |
| 93 | MyHeritage Keynote, RootsTech 2022 | MyHeritage | https://www.familysearch.org/rootstech/session/myheritage-keynote-rootstech-2022 | Not available |
| 94 | Using Labels to Name Your DNA Match List | MyHeritage | https://www.youtube.com/watch?v=enJjdw1xlsk | 139
|
| 95 | An Introduction to DNA on MyHeritage | MyHeritage – Daniel Horowitz | https://www.youtube.com/watch?v=1I6LHezMkgc | 60 |
| 96 | Using MyHeritage’s Advanced DNA Tools to Shed Light on Your DNA Matches | MyHeritage – Daniel Horowitz | https://www.youtube.com/watch?v=Pez46Xw20b4 | 110 |
| 97 | You’ve Got DNA Matches! Now What? | MyHeritage – Daniel Horowitz | https://www.youtube.com/watch?v=gl3UVksA-2E | 260 |
| 98 | My Story: Lizzie and Ayla | MyHeritage – Elizbeth Shaltz | https://www.youtube.com/watch?v=NQv6C8G39Kw | 147 |
| 99 | My Story: Fernando and Iwen | MyHeritage – Fernando Hermansson | https://www.youtube.com/watch?v=98-AR0M7fFE | 165
|
| 100 | Using the Autocluster and the Chromosome Browser to Explore Your DNA Matches | MyHeritage – Gal Zruhen | https://www.youtube.com/watch?v=a7aQbfP7lWU | 115
|
| 101 | My Story : Kara Ashby Utah Wedding | MyHeritage – Kara Ashby | https://www.youtube.com/watch?v=Qbr_gg1sDRo | 200 |
| 102 | When Harry Met Dotty – using DNA to break down brick walls | Nick David Barratt | https://www.youtube.com/watch?v=8SdnLuwWpJs | 679 |
| 103 | How to Add a DNA Match to Airtable | Nicole Dyer | https://www.youtube.com/watch?v=oKxizWIOKC0 | 161 |
| 104 | How to Download DNA Match Lists with DNAGedcom Client | Nicole Dyer | https://www.youtube.com/watch?v=t9zTWnwl98E | 124 |
| 105 | How to Know if a Matching DNA Segment is Maternal or Paternal | Nicole Dyer | https://www.youtube.com/watch?v=-zd5iat7pmg | 161 |
| 106 | DNA Basics Part I Centimorgans and Family Relationships | Origins International, Inc. dba Origins Genealogy | https://www.youtube.com/watch?v=SI1yUdnSpHA | 372 |
| 107 | DNA Basics Part II Clustering and Connecting Your DNA Matches | Origins International, Inc. dba Origins Genealogy | https://www.youtube.com/watch?v=ECs4a1hwGcs | 333 |
| 108 | DNA Basics Part III Charting Your DNA Matches to Get Answers | Origins International, Inc. dba Origins Genealogy | https://www.youtube.com/watch?v=qzybjN0JBGY | 270 |
| 109 | 2. Using Cluster Auto Painter | Patricia Coleman | https://www.youtube.com/watch?v=-nfLixwxKN4 | 691 |
| 110 | 3. Using Online Irish Records | Patricia Coleman | https://www.youtube.com/watch?v=mZsB0l4z4os | 802 |
| 111 | Exploring Different Types of Clusters | Patricia Coleman | https://www.youtube.com/watch?v=eEZBFPC8aL4 | 972
|
| 112 | The Million Mito Project: Growing the Family Tree of Womankind | Paul Maier | https://www.youtube.com/watch?v=cpctoeKb0Kw | 541 |
| 113 | The Tree of Mankind Age Estimates | Paul Maier | https://www.youtube.com/watch?v=jjkL8PWAEwk | 1638 |
| 114 | Y-DNA and Mitochondrial DNA Testing Plans | Paul Woodbury | https://www.youtube.com/watch?v=akymSm0QKaY | 168 |
| 115 | Finding Biological Family | Price Genealogy | https://www.youtube.com/watch?v=4xh-r3hZ6Hw | 137 |
| 116 | What Y-DNA Testing Can Do for You | Richard Hill | https://www.youtube.com/watch?v=a094YhIY4HU | 191 |
| 117 | Extending Time Horizons with DNA | Rob Spencer (live) | https://www.youtube.com/watch?v=wppXD1Zz2sQ | 1037 + live viewers |
| 118 | DNA for Native American Ancestry by Roberta Estes | Roberta Estes | https://www.youtube.com/watch?v=EbNyXCFfp4M | 212 |
| 119 | 1. Associating Autosomal DNA Segments With Ancestors | Roberta Estes (live) | https://www.youtube.com/watch?v=_IHSCkNnX48
|
~9000: 1019 + 500 live viewers + 7,400+ Facebook |
| 120 | 1. What Can I Do With Ancestral DNA Segments? | Roberta Estes (live) | https://www.youtube.com/watch?v=Suv3l4iZYAQ | 325 plus live viewers
|
| 121 | Native American DNA – Ancient and Contemporary Maps | Roberta Estes (live) | https://www.youtube.com/watch?v=dFTl2vXUz_0 | 212 plus 483 live viewers
|
| 122 | How Can DNA Enhance My Family History Research? | Robin Wirthlin | https://www.youtube.com/watch?v=f3KKW-U2P6w | 102 |
| 123 | How to Analyze a DNA Match | Robin Wirthlin | https://www.youtube.com/watch?v=LTL8NbpROwM | 367 |
| 124 | 1. Jewish Ethnicity & DNA: History, Migration, Genetics | Schelly Talalay Dardashti | https://www.youtube.com/watch?v=AIJyphGEZTA | 82
|
| 125 | 2. Jewish Ethnicity & DNA: History, Migration, Genetics | Schelly Talalay Dardashti | https://www.youtube.com/watch?v=VM3MCYM0hkI | 72 |
| 126 | Ask us about DNA | Talking Family History (live) | https://www.youtube.com/watch?v=kv_RfR6OPpU | 96 plus live viewers |
| 127 | 1. An Introduction to Visual Phasing | Tanner Blair Tolman | https://www.youtube.com/watch?v=WNhErW5UVKU
|
183 |
| 128 | 2. An Introduction to Visual Phasing | Tanner Blair Tolman | https://www.youtube.com/watch?v=CRpQ8EVOShI | 110
|
| 129 | Common Problems When Doing Visual Phasing | Tanner Blair Tolman | https://www.youtube.com/watch?v=hzFxtBS5a8Y | 68 |
| 130 | Cross Visual Phasing to Go Back Another Generation | Tanner Blair Tolman | https://www.youtube.com/watch?v=MrrMqhfiwbs | 64 |
| 131 | DNA Basics | Tanner Blair Tolman | https://www.youtube.com/watch?v=OCMUz-kXNZc | 155 |
| 132 | DNA Painter and Visual Phasing | Tanner Blair Tolman | https://www.youtube.com/watch?v=2-eh1L4wOmQ | 155 |
| 133 | DNA Painter Part 2: Chromosome Mapping | Tanner Blair Tolman | https://www.youtube.com/watch?v=zgOJDRG7hJc | 172 |
| 134 | DNA Painter Part 3: The Inferred Segment Generator | Tanner Blair Tolman | https://www.youtube.com/watch?v=96ai8nM4lzo
|
100 |
| 135 | DNA Painter Part 4: The Distinct Segment Generator | Tanner Blair Tolman | https://www.youtube.com/watch?v=Pu-WIEQ_8vc | 83 |
| 136 | DNA Painter Part 5: Ancestral Trees | Tanner Blair Tolman | https://www.youtube.com/watch?v=dkYDeFLduKA | 73 |
| 137 | Understanding Your DNA Ethnicity Results | Tanner Blair Tolman | https://www.youtube.com/watch?v=4tAd8jK6Bgw | 518 |
| 138 | What’s New at GEDmatch | Tim Janzen | https://www.youtube.com/watch?v=AjA59BG_cF4
|
515 |
| 139 | What Does it Mean to Have Neanderthal Ancestry? | Ugo Perego | https://www.youtube.com/watch?v=DshCKDW07so | 190 |
| 140 | Big Y-700 | Your DNA Guide | https://www.youtube.com/watch?v=rIFC69qswiA | 143 |
| 141 | Next Steps with Your DNA | Your DNA Guide – Diahan Southard (live) | https://www.familysearch.org/rootstech/session/next-steps-with-your-dna | Not yet available |
Additions:
142 Adventures of an Amateur Genetic Genealogist – Geoff Nelson https://www.familysearch.org/rootstech/session/adventures-of-an-amateur-genetic-genealogist 291 views
____________________________________________________________
Sign Up Now – It’s Free!
If you enjoyed this article, subscribe to DNAeXplain for free, to automatically receive new articles by email each week.
Here’s the link. Just look for the little grey “follow” button on the right-hand side on your computer screen below the black title bar, enter your e-mail address, and you’re good to go!
In case you were wondering, I never have nor ever will share or use your e-mail outside of the intended purpose.
_____________________________________________________________
Follow DNAexplain on Facebook, here or follow me on Twitter, here.
Share the Love!
You’re always welcome to forward articles or links to friends and share on social media.
If you haven’t already subscribed (it’s free,) you can receive an email whenever I publish by clicking the “follow” button on the main blog page, here.
You Can Help Keep This Blog Free
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Uploads
- FamilyTreeDNA – Y, mitochondrial, and autosomal DNA testing
- MyHeritage DNA – Autosomal DNA test
- MyHeritage FREE DNA file upload – Upload your DNA file from other vendors free
- AncestryDNA – Autosomal DNA test
- 23andMe Ancestry – Autosomal DNA only, no Health
- 23andMe Ancestry Plus Health
Genealogy Products and Services
- MyHeritage FREE Tree Builder – Genealogy software for your computer
- MyHeritage Subscription with Free Trial
- Legacy Family Tree Webinars – Genealogy and DNA classes, subscription-based, some free
- Legacy Family Tree Software – Genealogy software for your computer
- Newspapers.com – Search newspapers for your ancestors
- NewspaperArchive – Search different newspapers for your ancestors
My Book
- DNA for Native American Genealogy – by Roberta Estes, for those ordering within the United States
- DNA for Native American Genealogy – for those ordering outside the US
Genealogy Books
- Genealogical.com – Lots of wonderful genealogy research books
Genealogy Research
- Legacy Tree Genealogists – Professional genealogy research