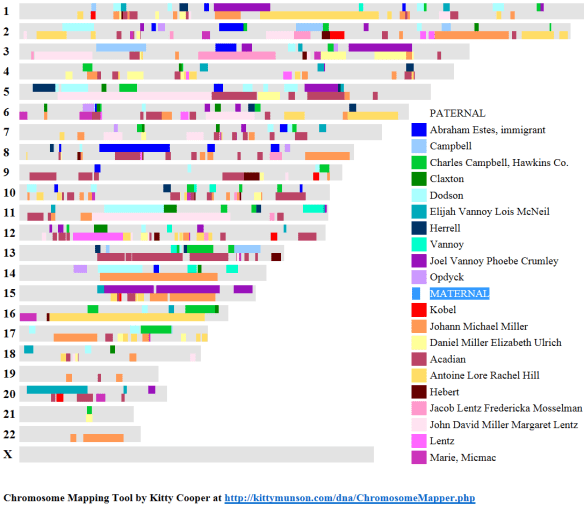
A few days after I published the article, Concepts – Segment Size, Legitimate and False Matches, Philip Gammon, a statistician who lives in Australia, posted a comment to my blog.
Great post Roberta! I’m a statistician so my eyes light up as soon as I see numbers. That table you have produced showing by segment length the percentage that are IBD is one of the most useful pieces of information that I have seen. Two days to do the analysis!!! I’m sure that I could write a formula that would identify the IBD segments and considerably reduce this time.
By this time, my eyes were lighting up too, because the work for the original article had taken me two days to complete manually, just using segments 3 cM and above. Using smaller segments would have taken days longer. By manually, I mean comparing the child’s matches with that of both parents’ matches to see which, if either, parent the child’s match also matches on the same segment.
In the simplest terms, the Segment Size article explained how to copy the child’s and both parents’ matches to a spreadsheet and then manually compare the child’s matches to those of the parents. In the example above, you can see that both the child and the mother have matches to Cecelia. As it turns out, the exact same segment of DNA was passed in its entirety to the child from the mother, who is shown in pink – so Cecelia matches both the child and the parent on exactly the same segment.
That’s not always the case, and the Segment Size article went into much greater detail.
For the past month or so, Philip and I have been working back and forth, along with some kind volunteers who tested Philip’s new tool, in order to create something so that you too can do this comparison and in much less than two days.
Foundation
Here’s the underlying principle for this tool – if a child has a match that does NOT match either parent on the same segment, then the match is not a legitimate match. It’s a false match, identical by chance, and it is NOT genealogically relevant.
If the child’s match also matches either parent on the same segment, it is most likely a match by descent and is genealogically relevant.
For those of you who noticed the words “most likely,” yes, it is possible for someone to match a parent and child both and still not phase (or match) to the next higher generation, but it’s unusual and so far, only found in smaller segments. I wrote about multiple generation phasing in the article, “Concepts – Segment Survival – 3 and 4 Generation Phasing.” Once a segment phases, it tends to continue phasing, especially with segments above about 3.5 cM.
For those who have both parents available to test, phased matching is a HUGE benefit.
But I Have Only One Parent Available
You can still use the tool to identify matches to that one parent, but you CANNOT presume that matches that DON’T match that parent are from the other (missing) parent. Matches matching the child but not matching the tested parent can be due to:
- A match to the missing parent
- A false match that is not genealogically relevant
According to the statistics generated from Philip’s Match-Maker-Breaker tool, shown below, segments 9 cM and above tend to match one or the other parent 90% or more of the time. Segments 12 cM and over match 97% of the time or more, so, in general, one could “assume” (dangerous word, I know) that segments of this size that don’t match to the tested parent would match to the other parent if the other parent was available. You can also see that the reliability of that assumption drops rapidly as the segment sizes get smaller.
Platform
This tool was written utilizing Microsoft Excel and only works reliably on that platform.
If you are using Excel and are NOT attempting to use MAC Numbers, skip this section. If you want to attempt to use Numbers, read this section.
I tried, along with a MAC person, to try to coax Numbers (free MAC spreadsheet) into working. If you have any other option other than using Numbers, so do. Microsoft Excel for MAC seemed to work fine, but it was only tested on one MAC.
Here’s what I discovered when trying to make Numbers work:
- You must first launch numbers and then select the various spreadsheets.
- The tabs are not at the bottom and are instead at the top without color.
- The instructions for copying the formulas in cells H2-K2 throughout the spreadsheet must be done manually with a copy/paste.
- After the above step, the calculations literally took a couple hours (MacBook Air) instead of a couple minutes on the PC platform. The older MAC desktop still took significantly longer than on a Microsoft PC, but less time than the solid state MacBook Air.
- After the calculations complete, the rows on the child’s spreadsheet are not colored, which is one of the major features of the Match-Maker-Breaker tool, as Numbers reports that “Conditional highlighting rules using formulas are not supported and were removed.”
- Surprisingly, the statistical Reports page seems to function correctly.
How Long Does Running Match-Maker-Breaker Tool on a PC Take?
The first time I ran this tool, which included reading Philip’s instructions for the first time, the entire process took me about 10 minutes after I downloaded the files from Family Tree DNA.
Vendors
This tool only works with matches downloaded from Family Tree DNA.
Transfer Kits
It’s strongly suggested that all 3 individuals being compared have tested at Family Tree DNA or on the same chip version imported into Family Tree DNA.
Matches not run on the same chip as Family Tree DNA testers can only provide a portion of the matches that the same person’s results run on the FTDNA chip can provide. You can run the matching tool with transferred results, but the results will only provide a subset of the results that will be provided by having all parties that are being compared, meaning the child and both parents, test at Family Tree DNA.
The following products versions CAN be all be compared successfully at Family Tree DNA, as they all utilize the same Illumina chip:
- All Family Finder tests
- Ancestry V1 (before May 2016)
- 23andMe V3 (before November 2013)
- MyHeritage
The following tests do NOT utilize the same Illumina testing platform and cannot be compared successfully with Family Finder tests from Family Tree DNA, or the list above. Cross platform testing results cannot be reliably compared. Those that DO match will be accurate, but many will not match that would match if all 3 testers were utilizing the same platform, therefore leading you to inaccurate conclusions.
- Ancestry V2 (beginning in May 2016 to present)
- 23andMe V4 (beginning November 2013 to present)
The child and two parents should not be compared utilizing mixed platforms – meaning, for example, that the child should not have been tested at FTDNA and the parents transferred from Ancestry on the V2 platform since May 2016.
If any of the three family members, being the child or either parent, have tested on an incompatible platform, they should retest at Family Tree DNA before using this tool.
What You Need
- You will need to download the chromosome match lists from the child and both parents, AT THE SAME TIME. I can’t stress this enough, because any matches that have been added for either of the three people at a later time than the others will skew the matching and the statistics. Matches are being added all the time.
- You will also need a relatively current version of Excel on your computer to run this tool. No, I did not do version compatibility testing so I don’t know how old is too old. I am running MSOffice 2013.
- You will need to know how to copy and paste data from and to a spreadsheet.
Instructions for Downloading Match Files
My recommendation is that you download your matches just before utilizing this tool.
To download your matches, sign on to each account. On your main page, you will see the Family Finder section, and the Chromosome Browser. Click on that link.
At the top of the chromosome browser page, below, you’ll see the image of chromosomes 1 through X. At the top right, you’ll see the option to “Download all matches to Excel (CSV Format). Click on that link.
Next, you’ll receive a prompt to open or save the file. Save it to a file name that includes the name of the person plus the date you did the download. I created a separate folder so there would be no confusion about which files are which and whether or not they are current.
Your match file includes all of your matches and the chromosome matching locations like the example shown below.
These files of matches are what you’ll need to copy into the Match-Maker-Breaker spreadsheet.
Do not delete any information from your match spreadsheets. If you normally delete small segments, don’t. You may cause a non-match situation if the parent carries a larger portion of the same segment.
You can rerun the Match-Maker-Breaker tool at will, and it only takes a very few minutes.
The Match-Maker-Breaker Tool
The Match-Maker-Breaker Tool has 5 sheets when you open the spreadsheet:
- Instructions – Please read entirely before beginning.
- Results – The page where your statistical results will be placed.
- Child – The page where you will paste the child’s matches and then look at the match results after processing.
- Father – The page where you will paste the father’s matches.
- Mother – The page where you will paste the mother’s matches.
Download
Download the free Match-Maker-Breaker tool which is a spreadsheet by clicking on this link: Match-Maker-Breaker Tool V2
Please don’t start using the tool before reading the instructions completely and reading the rest of this article.
Make a Copy
After you download the tool, make a copy on your system. You’ll want to save the Match-Maker-Breaker spreadsheet file for each trio of people individually, and you’ll want a fresh Match-Maker-Breaker spreadsheet copy to run with each new set of download files.
Instructions
I’m not going to repeat Philip’s instructions here, but please read them entirely before beginning and please follow them exactly. Philip has included graphic illustrations of each step to the right of the instruction box. The spreadsheet opens to the Instructions page. You can print the instruction page as well.
Copy/Pasting Data
When copying the parents’ and child’s data into the spreadsheets, do NOT copy and paste the entire page by selecting the page. Select and copy the relevant columns by highlighting columns A through G by touching your cursor to the A-G across the top, as shown below. After they are selected, then click on “copy.” In the child’s chromosome browser download spreadsheet, position the curser in the first cell in row 1 in the child’s page of the Match-Maker-Breaker spreadsheet and click on “paste.”
Do NOT select columns H-K when highlighting and copying, or your paste will wipe out Philip’s formulas to do calculations on the child’s tab on the spreadsheet.
The example above, assuming that Annie is the last entry on the spreadsheet, shows that I’ve highlighted all of the cells in columns A-G, prior to executing the copy command. Your spreadsheets of course will be much longer.
I wrote a very quick and dirty article about using Excel here
The Match Making Breaking Part
After you copy the formulas from rows H2 to K2 through the rest of the spreadsheet by following Philip’s instructions, you’ll see the results populating in the status bar at the bottom. You’ll also see colors being added to the matches on the left hand side of the spreadsheet page and counts accruing in the 4 right columns. Be patient and wait. It may take a few minutes. When it’s finished, you can verify by scrolling to the last row on the child’s page and you’ll see something like the example below, where every row has been assigned a color and every match that matches the child and the father, mother, both or is found in the HLA region is counted as 1 in the right 4 columns.
In this example, 5 segments, shown in grey, don’t match anyone, one, shown in tan is found in the HLA region, and three match the father, in blue.
Output
After you run the Match-Maker-Breaker tool, the child’s matches on the Child tab will be identified as follows:
This means that segment of the child that matches that individual also matches the father, the mother, both parents, the HLA region, or none of the above on all or part of that same segment.
What is a Match?
Philip and I worked to answer the question, “what is a match?” In the Concepts article, I discussed the various kinds of matches.
- Full match: The child’s match and parent’s match share the same exact segment, meaning same start and end points and same number of SNPs within that segment.
- Partial match: The child’s match matches a portion of the segment from the parent – meaning that the child inherited part of the segment, but not the entire segment.
- Overhanging match: The child’s match matches part or all of the parent’s segment, but either the beginning or end extends further than the parents match. This means that the overlapping portion is legitimate, meaning identical by descent (IBD), but the overhanging portion is identical by chance (IBC.)
- Nested match: The child’s match is smaller than the match to the parent, but fully within the parent’s match, indicating a legitimate match.
- No match: The person matches the child, but neither parent, meaning that this match is not legitimate. It’s identical by chance (IBC).
Full matches and no matches are easy.
However, partial matches, overlapping matches and nested matches are not as straightforward.
What, exactly, is a match? Let’s look at some different scenarios.
If someone matches a parent on a large segment, say 20cM, and only matches the child on 2cM, fully within the parent’s segment, is this match genealogically relevant, or could the match be matching the child by chance on a part of the same segment that they match the parents by descent? We have no way to know for sure, just utilizing this tool. Hopefully, in this case, the fact that the person matches the parent on a large segment would answer any genealogical questions through triangulation.
If the person matches the parent but only matches the child on a small portion of the same segment plus an overhanging region, is that a valid match? Because they do match on an overhanging region, we know that match is partly identical by chance, but is the entire match IBC or is the overlapping part legitimate? We don’t know. Partly, how strongly I would consider this a valid match would be the size of the matching portion of the segment.
One of the purposes of phasing and then looking at matches is to, hopefully, learn more about which matches are legitimate, which are not, and predictors of false versus legitimate matches.
Relative to this tool, no editing has been done, meaning that matches are presented exactly as that, regardless of their size or the type of match. A match is a match if any portion of the match’s DNA to the child overlaps any portion of either or both parent’s DNA, with the exception of part of chromosome 6. It’s up to you, as the genealogist, to figure out by utilizing triangulation and other tools whether the match is relevant or not to your genealogy.
If you are not familiar with identical by descent (meaning a legitimate match), identical by population (IBP) meaning identical by descent but because the population as a whole carries that segment and identical by chance (IBC) meaning a false match, the article Identical by…Descent, State, Population and Chance explains the terms and the concepts so that you can apply them usefully.
About Chromosome 6
After analyzing the results of several people, the area of chromosome 6 that includes the HLA region has been excluded from the analysis. Long known to be a pileup region where people carry significant segments of the same DNA that is not genealogically relevant (meaning IBP or identical by population,) this region has found to be often unreliable genealogically, and falls outside the norm as compared to the rest of the segments. This area has been annotated separately and excluded from match results. This was the only region found to universally have this effect.
This does not mean that a match in this region is positively invalid or false, but matches in the HLA region should be viewed very skeptically.
The Results Tab – Statistics
Now that you’ve populated the spreadsheet and you can see on the Child tab which matches also match either or both parents, or neither, or the HLA region, go to the Results tab of the spreadsheet.
This tab gives you some very interesting statistics.
First, you’ll see the number and percent of matches by chromosome.
The person compared was a female, so she would have X matches to both parents. However, notice that X matching is significantly lower than any of the other chromosomes.
Frankly, I’ve suspected for a long time that there was a dramatic difference in matching with the X chromosome, and wrote about it here. It was suggested by some at the time that I was only reporting my personal observations that would not hold beyond a few results (ascertainment bias), but this proves that there is something different about X chromosome matching. I don’t know what or why, but according to this data that is consistent between all of the beta testers, matching to the X chromosome is much less reliable.
The second statistics box you will see are statistics for the matches to the child that also match the parents. The actual matches of the child to the parents are shown as the 23 shown under “excluded from calculations.”
The next group of statistics on your page will be your own, but for this example, Philip has combined the results from several beta testers and provided summary information, so that the statistics are not skewed by any one individual.
Next, the match results by segment size for chromosomes 1-22. Philip has separated out segments with less than 500 SNPs and reports them separately.
You will note that 90% or more of the segments 9 cM and above match one of the two parents, and 97% or more of segments 12cM or above.
The X chromosome follows, analyzed separately. You’ll notice that while 27% of the matches on chromosomes 1-22 match one or both parents, only 14% of the X matches do.
Even with larger segments, not all X segments match both the child and the parents, suggesting that skepticism is warranted when evaluating X chromosome matches.
Philip then calculated a nice graph for showing matching autosomal segments by cM size, excluding the X.
The next set of charts shows matches by SNP density. Many people neglect SNP count when evaluating results, but the higher the SNP count, the more robust the match.
Note that SNP density above 2,200 almost always matched, but not always, while SNP density of 2,800 reaches the 97% threshold..
The X chromosome, by SNP count, below.
X segment reach the 100% threshold about 1600, however, we really need more results to be predictive at the same level as the results for chromosomes 1-22. Two data samples really isn’t adequate.
Once again, Philip prepared a nice chart showing percentage of matching segments by SNP count, below.
Predictive
In the Segment Survival – 3 and 4 Generation Phasing article, one can see that phased matches are predictive, meaning that a child/parent match is highly suggestive that the segment is a valid segment match and that it will hold in generations further upstream.
Several years ago, Dr. Tim Janzen, one of the early phasing pioneers, suggested that people test their children, even if both parents had already tested. For the life of me, I couldn’t understand how that would be the least bit productive, genealogically, since people were more likely to match the parents than the children, and children only carry a subset of their parent’s DNA.
However, the predictive nature of a segment being legitimate with a child/parent match to a third party means that even in situations where your own parent isn’t available, a match by a third party on the same segment with your child suggests that the match is legitimate, not IBC.
In the article, I showed both 3 and 4 generations of phased comparisons between generations of the same family and a known cousin. The results of the 5 different family comparisons are shown below, where the red segments did not phase or lost phasing between generations, and the green segments did phase through multiple generations.
Very, very few segments lost phasing in upper (older) generations after matching between a parent and a child. In the five 4-generation examples above, only a total of 7 groups of segments lost phasing. The largest segment that lost phasing in upper generations was 3.69 cM. In two examples, no segments were lost due to not phasing in upper generations.
The net-net of this is that you can benefit by testing your children if your parents aren’t available, because the matches on the segment to both you and the child are most likely to be legitimate. Of course, there will be segments where someone matches you and not your child, because your child did not inherit that segment of your DNA, and those may be legitimate matches as well. However, the segments where you and your child both match the same person will likely be legitimate matches, especially over about 3.5 cM. Please read the Segment Survival article for more details.
If you want to order additional Family Finder tests for more family members, you can click here.
Group Analysis
Philip has performed a group analysis which has produced some expected results along with some surprising revelations. I’d prefer to let people get their feet wet with this tool and the results it provides before publishing the results, with one exception.
In case you’re wondering if the comparisons used as examples, above, are representative of typical results, Philip analyzed 10 of our beta testers and says the following:
The results are remarkably consistent between all 10 participants. Summing it up in words: with each person that you match you will have an average of 11 matching segments. Three will be genuine and will add to [a total of] 21 cM. Eight will be false and add to [a total of] 19 cM.
Philip compiled the following chart summarizing 10 beta testers’ results. Please note that you can click to enlarge the images.
The X, being far less consistent, is shown below.
We Still Need Endogamous Parent-Child Trios
When I asked for volunteer testers, we were not able to obtain a trio of fully endogamous individuals. Specifically, we would like to see how the statistics for groups of non-endogamous individuals compare to the statistics for endogamous individuals.
Endogamous groups include people who are 100% Jewish, Amish, Mennonite, or have a significant amount of first or second cousin marriages in recent generations.
Of these, Jewish families prove to be the most highly endogamous, so if you are Jewish and have both Jewish parents’ DNA results, please run this tool and send either Philip or me the resulting spreadsheet. Your results won’t be personally identified, only the statistics used in conjunction with others, similar to the group analysis shown above. Your results will be entirely anonymous.
Philip’s e-mail is philip.gammon@optusnet.com.au and you can reach me at roberta@dnaexplain.com.
Caveat
Philip has created the Match-Maker-Breaker tool which is free to everyone. He has included some wonderful diagnostics, but Philip is not providing individual support for the tooI. In other words, this is a “what you see is what you get” gift.
Thank You and Acknowledgements
Of course, a very big thank you to Philip for creating this tool, and also to people who volunteered as alpha and beta testers and provided feedback. Also thanks to Jim Kvochick for trying to coax Numbers into working.
Match-Maker-Breaker Author Bio:
Philip’s official tagline reads: Philip Gammon, BEng(ManSysEng) RMIT, GradDipSc(AppStatistics) Swinburne
I asked Philip to describe himself.
I’d describe myself as a business analyst with a statistics degree plus an enthusiastic genetic genealogist with an interest in the mathematical and statistical aspects of inheritance and cousinship.
The important aspect of Philip’s resume is that he is applying his skills to genetic genealogy where they can benefit everyone. Thank you so much Philip.
Watch for some upcoming guest articles from Philip.
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research