Last year I wrote a column at the end of the year titled “2012 Top 10 Genetic Genealogy Happenings.” It’s amazing the changes in this industry in just one year. It certainly makes me wonder what the landscape a year from now will look like.
I’ve done the same thing this year, except we have a dozen. I couldn’t whittle it down to 10, partly because there has been so much more going on and so much change – or in the case of Ancestry, who is noteworthy because they had so little positive movement.
If I were to characterize this year of genetic genealogy, I would call it The Year of the SNP, because that applies to both Y DNA and autosomal. Maybe I’d call it The Legal SNP, because it is also the year of law, court decisions, lawsuits and FDA intervention. To say it has been interesting is like calling the Eiffel Tower an oversized coat hanger.
I’ll say one thing…it has kept those of us who work and play in this industry hopping busy! I guarantee you, the words “I’m bored” have come out of the mouth of no one in this industry this past year.
I’ve put these events in what I consider to be relatively accurate order. We could debate all day about whether the SNP Tsunami or the 23andMe mess is more important or relevant – and there would be lots of arguing points and counterpoints…see…I told you lawyers were involved….but in reality, we don’t know yet, and in the end….it doesn’t matter what order they are in on the list:)
Y Chromosome SNP Tsunami Begins
The SNP tsumani began as a ripple a few years ago with the introduction at Family Tree DNA of the Walk the Y program in 2007. This was an intensively manual process of SNP discovery, but it was effective.
By the time that the Geno 2.0 chip was introduced in 2012, 12,000+ SNPs would be included on that chip, including many that were always presumed to be equivalent and not regularly tested. However, the Nat Geo chip tested them and indeed, the Y tree became massively shuffled. The resolution to this tree shuffling hasn’t yet come out in the wash. Family Tree DNA can’t really update their Y tree until a publication comes out with the new tree defined. That publication has been discussed and anticipated for some time now, but it has yet to materialize. In the mean time, the volunteers who maintain the ISOGG tree are swamped, to say the least.
Another similar test is the Chromo2 introduced this year by Britain’s DNA which scans 15,000 SNPs, many of them S SNPs not on the tree nor academically published, adding to the difficulty of figuring out where they fit on the Y tree. While there are some very happy campers with their Chromo2 results, there is also a great deal of sloppy science, reporting and interpretation of “facts” through this company. Kind of like Jekyll and Hyde. See the Sloppy Science section.
But Walk the Y, Chromo2 and Geno 2.0, are only the tip of the iceburg. The new “full Y” sequencing tests brought into the marketspace quietly in early 2013 by Full Genomes and then with a bang by Family Tree DNA with the their Big Y in November promise to revolutionize what we know about the Y chromosome by discovering thousands of previously unknown SNPs. This will in effect swamp the Y tree whose branches we thought were already pretty robust, with thousands and thousands of leaves.
In essence, the promise of the “fully” sequenced Y is that what we might term personal or family SNPs will make SNP testing as useful as STR testing and give us yet another genealogy tool with which to separate various lines of one genetic family and to ratchet down on the time that the most common recent ancestor lived.
http://dna-explained.com/2013/03/31/new-y-dna-haplogroup-naming-convention/
http://dna-explained.com/2013/11/10/family-tree-dna-announces-the-big-y/
http://dna-explained.com/2013/11/16/what-about-the-big-y/
http://www.yourgeneticgenealogist.com/2013/11/first-look-at-full-genomes-y-sequencing.html
http://cruwys.blogspot.com/2013/12/a-first-look-at-britainsdna-chromo-2-y.html
http://cruwys.blogspot.com/2013/11/yseqnet-new-company-offering-single-snp.html
http://cruwys.blogspot.com/2013/11/the-y-chromosome-sequence.html
http://cruwys.blogspot.com/2013/11/a-confusion-of-snps.html
http://cruwys.blogspot.com/2013/11/a-simplified-y-tree-and-common-standard.html
23andMe Comes Unraveled
The story of 23andMe began as the consummate American dotcom fairy tale, but sadly, has deteriorated into a saga with all of the components of a soap opera. A wealthy wife starts what could be viewed as an upscale hobby business, followed by a messy divorce and a mystery run-in with the powerful overlording evil-step-mother FDA. One of the founders of 23andMe is/was married to the founder of Google, so funding, at least initially wasn’t an issue, giving 23andMe the opportunity to make an unprecedented contribution in the genetic, health care and genetic genealogy world.
Another way of looking at this is that 23andMe is the epitome of the American Dream business, a startup, with altruism and good health, both thrown in for good measure, well intentioned, but poorly managed. And as customers, be it for health or genealogy or both, we all bought into the altruistic “feel good” culture of helping find cures for dread diseases, like Parkinson’s, Alzheimer’s and cancer by contributing our DNA and responding to surveys.
The genetic genealogy community’s love affair with 23andMe began in 2009 when 23andMe started focusing on genealogy reporting for their tests, meaning cousin matches. We, as a community, suddenly woke up and started ordering these tests in droves. A few months later, Family Tree DNA also began offering this type of testing as well. The defining difference being that 23andMe’s primary focus has always been on health and medical information with Family Tree DNA focused on genetic genealogy. To 23andMe, the genetic genealogy community was an afterthought and genetic genealogy was just another marketing avenue to obtain more people for their health research data base. For us, that wasn’t necessarily a bad thing.
For awhile, this love affair went along swimmingly, but then, in 2012, 23andMe obtained a patent for Parkinson’s Disease. That act caused a lot of people to begin to question the corporate focus of 23andMe in the larger quagmire of the ethics of patenting genes as a whole. Judy Russell, the Legal Genealogist, discussed this here. It’s difficult to defend 23andMe’s Parkinson’s patent while flaying alive Myriad for their BRCA patent. Was 23andMe really as altruistic as they would have us believe?
Personally, this event made me very nervous, but I withheld judgment. But clearly, that was not the purpose for which I thought my DNA, and others, was being used.
But then came the Designer Baby patent in 2013. This made me decidedly uncomfortable. Yes, I know, some people said this really can’t be done, today, while others said that it’s being done anyway in some aspects…but the fact that this has been the corporate focus of 23andMe with their research, using our data, bothered me a great deal. I have absolutely no issue with using this information to assure or select for healthy offspring – but I have a personal issue with technology to enable parents who would select a “beauty child,” one with blonde hair and blue eyes and who has the correct muscles to be a star athlete, or cheerleader, or whatever their vision of their as-yet-unconceived “perfect” child would be. And clearly, based on 23andMe’s own patent submission, that is the focus of their patent.
Upon the issuance of the patent, 23andMe then said they have no intention of using it. They did not say they won’t sell it. This also makes absolutely no business sense, to focus valuable corporate resources on something you have no intention of using? So either they weren’t being truthful, they lack effective management or they’ve changed their mind, but didn’t state such.
What came next, in late 2013 certainly points towards a lack of responsible management.
23andMe had been working with the FDA for approval the health and medical aspect of their product (which they were already providing to consumers prior to the November 22nd cease and desist order) for several years. The FDA wants assurances that what 23andMe is telling consumers is accurate. Based on the letter issued to 23andMe on November 22nd, and subsequent commentary, it appears that both entities were jointly working towards that common goal…until earlier this year when 23andMe mysteriously “somehow forgot” about the FDA, the information they owed them, their submissions, etc. They also forgot their phone number and their e-mail addresses apparently as well, because the FDA said they had heard nothing from them in 6 months, which backdates to May of 2013.
It may be relevant that 23andMe added the executive position of President and filled it in June of 2013, and there was a lot of corporate housecleaning that went on at that time. However, regardless of who got housecleaned, the responsibility for working with the FDA falls squarely on the shoulders of the founders, owners and executives of the company. Period. No excuses. Something that critically important should be on the agenda of every executive management meeting. Why? In terms of corporate risk, this was obviously a very high risk item, perhaps the highest risk item, because the FDA can literally shut their doors and destroy them. There is little they can do to control or affect the FDA situation, except to work with the FDA, meet deadlines and engender goodwill and a spirit of cooperation. The risk of not doing that is exactly what happened.
It’s unknown at this time if 23andMe is really that corporately arrogant to think they could simply ignore the FDA, or blatantly corporately negligent or maybe simply corporately stupid, but they surely betrayed the trust and confidence of their customers by failing to meet their commitments with and to the FDA, or even communicate with them. I mean, really, what were they thinking?
There has been an outpouring of sympathy for 23andme and negative backlash towards the FDA for their letter forcing 23andMe to stop selling their offending medical product, meaning the health portion of their testing. However, in reality, the FDA was only meting out the consequences that 23andMe asked for. My teenage kids knew this would happen. If you do what you’re not supposed to….X, Y and Z will, or won’t, happen. It’s called accountability. Just ask my son about his prom….he remembers vividly. Now why my kids, or 23andMe, would push an authority figure to that point, knowing full well the consequences, utterly mystifies me. It did when my son was a teenager and it does with 23andMe as well.
Some people think that the FDA is trying to stand between consumers and their health information. I don’t think so, at least not in this case. Why I think that is because the FDA left the raw data files alone and they left the genetic genealogy aspect alone. The FDA knows full well you can download your raw data and for $5 process it at a third party site, obtaining health related genetic information. The difference is that Promethease is not interpreting any data for you, only providing information.
There is some good news in this and that is that from a genetic genealogy perspective, we seem to be safe, at least for now, from government interference with the testing that has been so productive for genetic genealogy. The FDA had the perfect opportunity to squish us like a bug (thanks to the opening provided by 23andMe,) and they didn’t.
The really frustrating aspect of this is that 23andMe was a company who, with their deep pockets in Silicon Valley and other investors, could actually afford to wage a fight with the FDA, if need be. The other companies who received the original 2010 FDA letter all went elsewhere and focused on something else. But 23andMe didn’t, they decided to fight the fight, and we all supported their decision. But they let us all down. The fight they are fighting now is not the battle we anticipated, but one brought upon themselves by their own negligence. This battle didn’t have to happen, and it may impair them financially to such a degree that if they need to fight the big fight, they won’t be able to.
Right now, 23andMe is selling their kits, but only as an ancestry product as they work through whatever process they are working through with the FDA. Unfortunately, 23andMe is currently having some difficulties where the majority of matches are disappearing from some testers records. In other cases, segments that previously matched are disappearing. One would think, with their only revenue stream for now being the genetic genealogy marketspace that they would be wearing kid gloves and being extremely careful, but apparently not. They might even consider making some of the changes and enhancements we’ve requested for so long that have fallen on deaf ears.
One thing is for sure, it will be extremely interesting to see where 23andMe is this time next year. The soap opera continues.
I hope for the sake of all of the health consumers, both current and (potentially) future, that this dotcom fairy tale has a happy ending.
Also, see the Autosomal DNA Comes of Age section.
http://dna-explained.com/2013/10/05/23andme-patents-technology-for-designer-babies/
http://www.thegeneticgenealogist.com/2013/10/07/a-new-patent-for-23andme-creates-controversy/
http://dna-explained.com/2013/11/13/genomics-law-review-discusses-designing-children/
http://www.thegeneticgenealogist.com/2013/06/11/andy-page-fills-new-president-position-at-23andme/
http://dna-explained.com/2013/11/25/fda-orders-23andme-to-discontinue-testing/
http://dna-explained.com/2013/11/26/now-what-23andme-and-the-fda/
http://dna-explained.com/2013/12/06/23andme-suspends-health-related-genetic-tests/
http://www.legalgenealogist.com/blog/2013/11/26/fooling-with-fda/
Supreme Court Decision – Genes Can’t Be Patented – Followed by Lawsuits
In a landmark decision, the Supreme Court determined that genes cannot be patented. Myriad Genetics held patents on two BRCA genes that predisposed people to cancer. The cost for the tests through Myriad was about $3000. Six hours after the Supreme Court decision, Gene By Gene announced that same test for $995. Other firms followed suit, and all were subsequently sued by Myriad for patent infringement. I was shocked by this, but as one of my lawyer friends clearly pointed out, you can sue anyone for anything. Making it stick is yet another matter. Many firms settle to avoid long and very expensive legal battles. Clearly, this issue is not yet resolved, although one would think a Supreme Court decision would be pretty definitive. It potentially won’t be settled for a long time.
http://dna-explained.com/2013/06/13/supreme-court-decision-genes-cant-be-patented/
http://www.legalgenealogist.com/blog/2013/06/14/our-dna-cant-be-patented/
http://dna-explained.com/2013/09/07/message-from-bennett-greenspan-free-my-genes/
http://www.thegeneticgenealogist.com/2013/06/13/new-press-release-from-dnatraits-regarding-the-supreme-courts-holding-in-myriad/
http://www.legalgenealogist.com/blog/2013/08/18/testing-firms-land-counterpunch/
http://www.legalgenealogist.com/blog/2013/07/11/myriad-sues-genetic-testing-firms/
Gene By Gene Steps Up, Ramps Up and Produces
As 23andMe comes unraveled and Ancestry languishes in its mediocrity, Gene by Gene, the parent company of Family Tree DNA has stepped up to the plate, committed to do “whatever it takes,” ramped up the staff both through hiring and acquisitions, and is producing results. This is, indeed, a breath of fresh air for genetic genealogists, as well as a welcome relief.
http://dna-explained.com/2013/08/07/gene-by-gene-acquires-arpeggi/
http://dna-explained.com/2013/12/05/family-tree-dna-listens-and-acts/
http://dna-explained.com/2013/12/10/family-tree-dnas-family-finder-match-matrix-released/
http://www.haplogroup.org/ftdna-family-finder-matches-get-new-look/
http://www.haplogroup.org/ftdna-family-finder-new-look-2/
http://www.haplogroup.org/ftdna-family-finder-matches-new-look-3/
Autosomal DNA Comes of Age
Autosomal DNA testing and analysis has simply exploded this past year. More and more people are testing, in part, because Ancestry.com has a captive audience in their subscription data base and more than a quarter million of those subscribers have purchased autosomal DNA tests. That’s a good thing, in general, but there are some negative aspects relative to Ancestry, which are in the Ancestry section.
Another boon to autosomal testing was the 23andMe push to obtain a million records. Of course, the operative word here is “was” but that may revive when the FDA issue is resolved. One of the down sides to the 23andMe data base, aside from the fact that it’s not genealogist friendly, is that so many people, about 90%, don’t communicate. They aren’t interested in genealogy.
A third factor is that Family Tree DNA has provided transfer ability for files from both 23andMe and Ancestry into their data base.
Fourth is the site, GedMatch, at www.gedmatch.com which provides additional matching and admixture tools and the ability to match below thresholds set by the testing companies. This is sometimes critically important, especially when comparing to known cousins who just don’t happen to match at the higher thresholds, for example. Unfortunately, not enough people know about GedMatch, or are willing to download their files. Also unfortunate is that GedMatch has struggled for the past few months to keep up with the demand placed on their site and resources.
A great deal of time this year has been spent by those of us in the education aspect of genetic genealogy, in whatever our capacity, teaching about how to utilize autosomal results. It’s not necessarily straightforward. For example, I wrote a 9 part series titled “The Autosomal Me” which detailed how to utilize chromosome mapping for finding minority ethnic admixture, which was, in my case, both Native and African American.
As the year ends, we have Family Tree DNA, 23andMe and Ancestry who offer the autosomal test which includes the relative-matching aspect. Fortunately, we also have third party tools like www.GedMatch.com and www.DNAGedcom.com, without which we would be significantly hamstrung. In the case of DNAGedcom, we would be unable to perform chromosome segment matching and triangulation with 23andMe data without Rob Warthen’s invaluable tool.
http://dna-explained.com/2013/06/21/triangulation-for-autosomal-dna/
http://dna-explained.com/2013/07/13/combining-tools-autosomal-plus-y-dna-mtdna-and-the-x-chromosome/
http://dna-explained.com/2013/07/26/family-tree-dna-levels-the-playing-field-sort-of/
http://dna-explained.com/2013/08/03/kitty-coopers-chromsome-mapping-tool-released/
http://dna-explained.com/2013/09/29/why-dont-i-match-my-cousin/
http://dna-explained.com/2013/10/03/family-tree-dna-updates-family-finder-and-adds-triangulation/
http://dna-explained.com/2013/10/21/why-are-my-predicted-cousin-relationships-wrong/
http://dna-explained.com/2013/12/05/family-tree-dna-listens-and-acts/
http://dna-explained.com/2013/12/09/chromosome-mapping-aka-ancestor-mapping/
http://dna-explained.com/2013/12/10/family-tree-dnas-family-finder-match-matrix-released/
http://dna-explained.com/2013/12/15/one-chromosome-two-sides-no-zipper-icw-and-the-matrix/
http://dna-explained.com/2013/06/02/the-autosomal-me-summary-and-pdf-file/
DNAGedcom – Indispensable Third Party Tool
While this tool, www.dnagedcom.com, falls into the Autosomal grouping, I have separated it out for individual mention because without this tool, the progress made this year in autosomal DNA ancestor and chromosomal mapping would have been impossible. Family Tree DNA has always provided segment matching boundaries through their chromosome browser tool, but until recently, you could only download 5 matches at a time. This is no longer the case, but for most of the year, Rob’s tool saved us massive amounts of time.
23andMe does not provide those chromosome boundaries, but utilizing Rob’s tool, you can obtain each of your matches in one download, and then you can obtain the list of who your matches match that is also on your match list by requesting each of those files separately. Multiple steps? Yes, but it’s the only way to obtain this information, and chromosome mapping without the segment data is impossible
A special hats off to Rob. Please remember that Rob’s site is free, meaning it’s donation based. So, please donate if you use the tool.
http://www.yourgeneticgenealogist.com/2013/01/brought-to-you-by-adoptiondna.html
I covered www.Gedmatch.com in the “Best of 2012” list, but they have struggled this year, beginning when Ancestry announced that raw data file downloads were available. GedMatch consists of two individuals, volunteers, who are still struggling to keep up with the required processing and the tools. They too are donation based, so don’t forget about them if you utilize their tools.
Ancestry – How Great Thou Aren’t
Ancestry is only on this list because of what they haven’t done. When they initially introduced their autosomal product, they didn’t have any search capability, they didn’t have a chromosome browser and they didn’t have raw data file download capability, all of which their competitors had upon first release. All they did have was a list of your matches, with their trees listed, with shakey leaves if you shared a common ancestor on your tree. The implication, was, and is, of course, that if you have a DNA match and a shakey leaf, that IS your link, your genetic link, to each other. Unfortunately, that is NOT the case, as CeCe Moore documented in her blog from Rootstech (starting just below the pictures) as an illustration of WHY we so desperately need a chromosome browser tool.
In a nutshell, Ancestry showed the wrong shakey leaf as the DNA connection – as proven by the fact that both of CeCe’s parents have tested at Ancestry and the shakey leaf person doesn’t match the requisite parent. And there wasn’t just one, not two, but three instances of this. What this means is, of course, that the DNA match and the shakey leaf match are entirely independent of each other. In fact, you could have several common ancestors, but the DNA at any particular location comes only from one on either Mom or Dad’s side – any maybe not even the shakey leaf person.
So what Ancestry customers are receiving is a list of people they match and possible links, but most of them have no idea that this is the case, and blissfully believe they have found their genetic connection. They have found a genealogical cousin, and it MIGHT be the genetic connection. But then again, they could have found that cousin simply by searching for the same ancestor in Ancestry’s data base. No DNA needed.
Ancestry has added a search feature, allowed raw data file downloads (thank you) and they have updated their ethnicity predictions. The ethnicity predictions are certainly different, dramatically different, but equally as unrealistic. See the Ethnicity Makeovers section for more on this. The search function helps, but what we really need is the chromosome browser, which they have steadfastly avoided promising. Instead, they have said that they will give us “something better,” but nothing has materialized.
I want to take this opportunity, to say, as loudly as possible, that TRUST ME IS NOT ACCEPTABLE in any way, shape or form when it comes to genetic matching. I’m not sure what Ancestry has in mind by the way of “better,” but it if it’s anything like the mediocrity with which their existing DNA products have been rolled out, neither I nor any other serious genetic genealogist will be interested, satisfied or placated.
Regardless, it’s been nearly 2 years now. Ancestry has the funds to do development. They are not a small company. This is obviously not a priority because they don’t need to develop this feature. Why is this? Because they can continue to sell tests and to give shakey leaves to customers, most of whom don’t understand the subtle “untruth” inherent in that leaf match – so are quite blissfully happy.
In years past, I worked in the computer industry when IBM was the Big Dog against whom everyone else competed. I’m reminded of an old joke. The IBM sales rep got married, and on his wedding night, he sat on the edge of the bed all night long regaling his bride in glorious detail with stories about just how good it was going to be….
You can sign a petition asking Ancestry to provide a chromosome browser here, and you can submit your request directly to Ancestry as well, although to date, this has not been effective.
The most frustrating aspect of this situation is that Ancestry, with their plethora of trees, savvy marketing and captive audience testers really was positioned to “do it right,” and hasn’t, at least not yet. They seem to be more interested in selling kits and providing shakey leaves that are misleading in terms of what they mean than providing true tools. One wonders if they are afraid that their customers will be “less happy” when they discover the truth and not developing a chromosome browser is a way to keep their customers blissfully in the dark.
http://dna-explained.com/2013/03/21/downloading-ancestrys-autosomal-dna-raw-data-file/
http://dna-explained.com/2013/03/24/ancestry-needs-another-push-chromosome-browser/
http://dna-explained.com/2013/10/17/ancestrys-updated-v2-ethnicity-summary/
http://www.thegeneticgenealogist.com/2013/06/21/new-search-features-at-ancestrydna-and-a-sneak-peek-at-new-ethnicity-estimates/
http://www.yourgeneticgenealogist.com/2013/03/ancestrydna-raw-data-and-rootstech.html
http://www.legalgenealogist.com/blog/2013/09/15/dna-disappointment/
http://www.legalgenealogist.com/blog/2013/09/13/ancestrydna-begins-rollout-of-update/
Ancient DNA
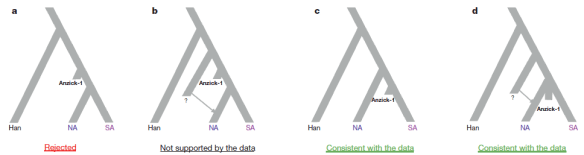
This has been a huge year for advances in sequencing ancient DNA, something once thought unachievable. We have learned a great deal, and there are many more skeletal remains just begging to be sequenced. One absolutely fascinating find is that all people not African (and some who are African through backmigration) carry Neanderthal and Denisovan DNA. Just this week, evidence of yet another archaic hominid line has been found in Neanderthal DNA and on Christmas Day, yet another article stating that type 2 Diabetes found in Native Americans has roots in their Neanderthal ancestors. Wow!
Closer to home, by several thousand years is the suggestion that haplogroup R did not exist in Europe after the ice age, and only later, replaced most of the population which, for males, appears to have been primarily haplogroup G. It will be very interesting as the data bases of fully sequenced skeletons are built and compared. The history of our ancestors is held in those precious bones.
http://dna-explained.com/2013/01/10/decoding-and-rethinking-neanderthals/
http://dna-explained.com/2013/07/04/ancient-dna-analysis-from-canada/
http://dna-explained.com/2013/07/10/5500-year-old-grandmother-found-using-dna/
http://dna-explained.com/2013/10/25/ancestor-of-native-americans-in-asia-was-30-western-eurasian/
http://dna-explained.com/2013/11/12/2013-family-tree-dna-conference-day-2/
http://dna-explained.com/2013/11/22/native-american-gene-flow-europe-asia-and-the-americas/
http://dna-explained.com/2013/12/05/400000-year-old-dna-from-spain-sequenced/
http://www.thegeneticgenealogist.com/2013/10/16/identifying-otzi-the-icemans-relatives/
http://cruwys.blogspot.com/2013/12/recordings-of-royal-societys-ancient.html
http://cruwys.blogspot.com/2013/02/richard-iii-king-is-found.html
http://dna-explained.com/2013/12/22/sequencing-of-neanderthal-toe-bone-reveals-unknown-hominin-line/
http://dna-explained.com/2013/12/26/native-americans-neanderthal-and-denisova-admixture/
http://dienekes.blogspot.com/2013/12/ancient-dna-what-2013-has-brought.html
Sloppy Science and Sensationalist Reporting
Unfortunately, as DNA becomes more mainstream, it becomes a target for both sloppy science or intentional misinterpretation, and possibly both. Unfortunately, without academic publication, we can’t see results or have the sense of security that comes from the peer review process, so we don’t know if the science and conclusions stand up to muster.
The race to the buck in some instances is the catalyst for this. In other cases, and not in the links below, some people intentionally skew interpretations and results in order to either fulfill their own belief agenda or to sell “products and services” that invariably report specific findings.
It’s equally as unfortunate that much of these misconstrued and sensationalized results are coming from a testing company that goes by the names of BritainsDNA, ScotlandsDNA, IrelandsDNA and YorkshiresDNA. It certainly does nothing for their credibility in the eyes of people who are familiar with the topics at hand, but it does garner a lot of press and probably sells a lot of kits to the unwary.
I hope they publish their findings so we can remove the “sloppy science” aspect of this. Sensationalist reporting, while irritating, can be dealt with if the science is sound. However, until the results are published in a peer-reviewed academic journal, we have no way of knowing.
Thankfully, Debbie Kennett has been keeping her thumb on this situation, occurring primarily in the British Isles.
http://dna-explained.com/2013/08/24/you-might-be-a-pict-if/
http://cruwys.blogspot.com/2013/12/the-british-genetic-muddle-by-alistair.html
http://cruwys.blogspot.com/2013/12/setting-record-straight-about-sara.html
http://cruwys.blogspot.com/2013/09/private-eye-on-britainsdna.html
http://cruwys.blogspot.com/2013/07/private-eye-on-prince-williams-indian.html
http://cruwys.blogspot.com/2013/06/britainsdna-times-and-prince-william.html
http://cruwys.blogspot.com/2013/03/sense-about-genealogical-dna-testing.html
http://cruwys.blogspot.com/2013/03/sense-about-genetic-ancestry-testing.html
Citizen Science is Coming of Age
Citizen science has been slowing coming of age over the past few years. By this, I mean when citizen scientists work as part of a team on a significant discovery or paper. Bill Hurst comes to mind with his work with Dr. Doron Behar on his paper, A Copernican Reassessment of the Human Mitochondrial DNA from its Root or what know as the RSRS model. As the years have progressed, more and more discoveries have been made or assisted by citizen scientists, sometimes through our projects and other times through individual research. JOGG, the Journal of Genetic Genealogy, which is currently on hiatus waiting for Dr. Turi King, the new editor, to become available, was a great avenue for peer reviewed publication. Recently, research projects have been set up by citizen scientists, sometimes crowd-funded, for specific areas of research. This is a very new aspect to scientific research, and one not before utilized.
The first paper below includes the Family Tree DNA Lab, Thomas and Astrid Krahn, then with Family Tree DNA and Bonnie Schrack, genetic genealogist and citizen scientist, along with Dr. Michael Hammer from the University of Arizona and others.
http://dna-explained.com/2013/03/26/family-tree-dna-research-center-facilitates-discovery-of-ancient-root-to-y-tree/
http://dna-explained.com/2013/04/10/diy-dna-analysis-genomeweb-and-citizen-scientist-2-0/
http://dna-explained.com/2013/06/27/big-news-probable-native-american-haplogroup-breakthrough/
http://dna-explained.com/2013/07/22/citizen-science-strikes-again-this-time-in-cameroon/
http://dna-explained.com/2013/11/30/native-american-haplogroups-q-c-and-the-big-y-test/
http://www.yourgeneticgenealogist.com/2013/03/citizen-science-helps-to-rewrite-y.html
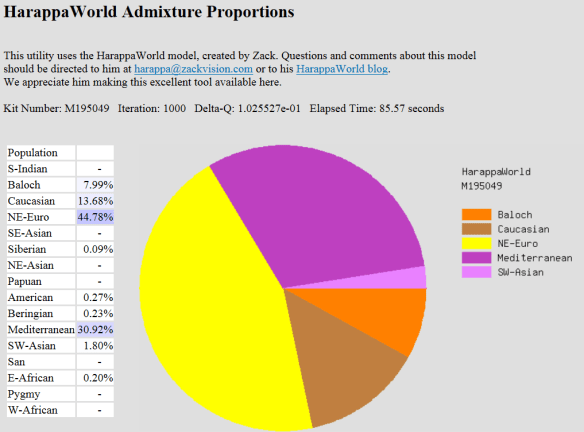
Ethnicity Makeovers – Still Not Soup
Unfortunately, ethnicity percentages, as provided by the major testing companies still disappoint more than thrill, at least for those who have either tested at more than one lab or who pretty well know their ethnicity via an extensive pedigree chart.
Ancestry.com is by far the worse example, swinging like a pendulum from one extreme to the other. But I have to hand it to them, their marketing is amazing. When I signed in, about to discover that my results had literally almost reversed, I was greeted with the banner “a new you.” Yea, a new me, based on Ancestry’s erroneous interpretation. And by reversed, I’m serious. I went from 80% British Isles to 6% and then from 0% Western Europe to 79%. So now, I have an old wrong one and a new wrong one – and indeed they are very different. Of course, neither one is correct…..but those are just pesky details…
23andMe updated their ethnicity product this year as well, and fine tuned it yet another time. My results at 23andMe are relatively accurate. I saw very little change, but others saw more. Some were pleased, some not.
The bottom line is that ethnicity tools are not well understood by consumers in terms of the timeframe that is being revealed, and it’s not consistent between vendors, nor are the results. In some cases, they are flat out wrong, as with Ancestry, and can be proven. This does not engender a great deal of confidence. I only view these results as “interesting” or utilize them in very specific situations and then only using the individual admixture tools at www.Gedmatch.com on individual chromosome segments.
As Judy Russell says, “it’s not soup yet.” That doesn’t mean it’s not interesting though, so long as you understand the difference between interesting and gospel.
http://dna-explained.com/2013/08/05/autosomal-dna-ancient-ancestors-ethnicity-and-the-dandelion/
http://dna-explained.com/2013/10/04/ethnicity-results-true-or-not/
http://www.legalgenealogist.com/blog/2013/09/15/dna-disappointment/
http://cruwys.blogspot.com/2013/09/my-updated-ethnicity-results-from.html?utm_source=feedburner&utm_medium=email&utm_campaign=Feed%3A+Cruwysnews+%28Cruwys+news%29
http://dna-explained.com/2013/10/17/ancestrys-updated-v2-ethnicity-summary/
http://dna-explained.com/2013/10/19/determining-ethnicity-percentages/
http://www.thegeneticgenealogist.com/2013/09/12/ancestrydna-launches-new-ethnicity-estimate/
http://cruwys.blogspot.com/2013/12/a-first-look-at-chromo-2-all-my.html
Genetic Genealogy Education Goes Mainstream
With the explosion of genetic genealogy testing, as one might expect, the demand for education, and in particular, basic education has exploded as well.
I’ve written a 101 series, Kelly Wheaton wrote a series of lessons and CeCe Moore did as well. Recently Family Tree DNA has also sponsored a series of free Webinars. I know that at least one book is in process and very near publication, hopefully right after the first of the year. We saw several conferences this year that provided a focus on Genetic Genealogy and I know several are planned for 2014. Genetic genealogy is going mainstream!!! Let’s hope that 2014 is equally as successful and that all these folks asking for training and education become avid genetic genealogists.
http://dna-explained.com/2013/08/10/ngs-series-on-dna-basics-all-4-parts/
https://sites.google.com/site/wheatonsurname/home
http://www.yourgeneticgenealogist.com/2012/08/getting-started-in-dna-testing-for.html
http://dna-explained.com/2013/12/17/free-webinars-from-family-tree-dna/
http://www.thegeneticgenealogist.com/2013/06/09/the-first-dna-day-at-the-southern-california-genealogy-society-jamboree/
http://www.yourgeneticgenealogist.com/2013/06/the-first-ever-independent-genetic.html
http://cruwys.blogspot.com/2013/10/genetic-genealogy-comes-to-ireland.html
http://cruwys.blogspot.com/2013/03/wdytya-live-day-3-part-2-new-ancient.html
http://cruwys.blogspot.com/2013/03/who-do-you-think-you-are-live-day-3.html
http://cruwys.blogspot.com/2013/03/who-do-you-think-you-are-live-2013-days.html
http://genealem-geneticgenealogy.blogspot.com/2013/03/the-surnames-handbook-guide-to-family.html
http://www.isogg.org/wiki/Beginners%27_guides_to_genetic_genealogy
A Thank You in Closing
I want to close by taking a minute to thank the thousands of volunteers who make such a difference. All of the project administrators at Family Tree DNA are volunteers, and according to their website, there are 7829 projects, all of which have at least one administrator, and many have multiple administrators. In addition, everyone who answers questions on a list or board or on Facebook is a volunteer. Many donate their time to coordinate events, groups, or moderate online facilities. Many speak at events or for groups. Many more write articles for publications from blogs to family newsletters. Additionally, there are countless websites today that include DNA results…all created and run by volunteers, not the least of which is the ISOGG site with the invaluable ISOGG wiki. Without our volunteer army, there would be no genetic genealogy community. Thank you, one and all.
2013 has been a banner year, and 2014 holds a great deal of promise, even without any surprises. And if there is one thing this industry is well known for….it’s surprises. I can’t wait to see what 2014 has in store for us!!! All I can say is hold on tight….
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Services
Genealogy Research
Like this:
Like Loading...