Recently, as a comment to one of my blog postings, someone asked how the testing companies can reach so far back in time and tell you about your ancestors. Great question.
The tests that reliably reach the furthest back, of course, are the direct line Y-Line and mitochondrial DNA tests, but the commenter was really asking about the ethnicity predictions. Those tests are known as BGA, or biogeographical ancestry tests, but most people just think of them or refer to them as the ethnicity tests.
Currently, Family Tree DNA, 23andMe and Ancestry.com all provide this function as a part of their autosomal product along with the Genographic 2.0 test. In addition, third party tools available at www.gedmatch.com don’t provide testing, but allow you to expand what you can learn with their admixture tools if you upload your raw data files to their site. I wrote about how to use these ethnicity tools in “The Autosomal Me” series. I’ve also written about how accurate ethnicity predictions from testing companies are, or aren’t, here, here and here.
But today, I’d like to just briefly review the 3 steps in ethnicity prediction, and how those steps are accomplished. It’s simple, really, in concept, but like everything else, the devil is in the details.
There are three fundamental steps.
- Creation of the underlying population data base.
- Individual DNA extraction.
- Comparison to the underlying population data base.
Step 1: Creation of the underlying population data base.
Don’t we wish this was as simple as it sounds. It isn’t. In fact, this step is the underpinnings of the accuracy of the ethnicity predictions. The old GIGO (garbage in, garbage out) concept applies here.
How do researchers today obtain samples of what ancestral populations looked like, genetically? Of course, the evident answer is through burials, but burials are not only few and far between, the DNA often does not amplify, or isn’t obtainable at all, and when it is, we really don’t have any way to know if we have a representative sample of the indigenous population (at that point in time) or a group of travelers passing through. So, by and large, with few exceptions, ancient DNA isn’t a readily available option.
The second way to obtain this type of information is to sample current populations, preferably ones in isolated regions, not prone to in-movement, like small villages in mountain valleys, for example, that have been stable “forever.” This is the approach the National Geographic Society takes and a good part of what the Genograpic Geno 2.0 project funding does. Indigenous populations are in most cases our most reliable link to the past. These resources, combined with what we know about population movement and history are very telling. In fact, National Geographic included over 75,000 AIMs (Ancestrally Informative Markers) on the Geno 2.0 chip when it was released.
The third way to obtain this type of information is by inference. Both Ancestry.com and 23andMe do some of this. Ancestry released its V2 ethnicity updates this week, and as a part of that update, they included a white paper available to DNA participants. In that paper, Ancestry discusses their process for utilizing contributed pedigree charts and states that, aside from immigrant locations, such as the United States and Canada, a common location for 4 grandparents is sufficient information to include that individuals DNA as “native” to that location. Ancestry used 3000 samples in their new ethnicity predictions to cover 26 geographic locations. That’s only 115 samples, on average, per location to represent all of that population. That’s pretty slim pickins. Their most highly represented area is Eastern Europe with 432 samples and the least represented is Mali with 16. The regions they cover are shown below.
Survey Monkey, a widely utilized web survey company, in their FAQ about Survey Size For Accuracy provides guidelines for obtaining a representative sample. Take a look. No matter which calculations you use relative to acceptable Margin of Error and Confidence Level, Ancestry’s sample size is extremely light.
23andMe states in their FAQ that their ethnicity prediction, called Ancestry Composition covers 22 reference populations and that they utilize public reference datasets in addition to their clients’ with known ancestry.
23andMe asks geographic ancestry questions of their customers in the “where are you from” survey, then incorporates the results of individuals with all 4 grandparents from a particular country. One of the ways they utilize this data is to show you where on your chromosomes you match people whose 4 grandparents are from the same country. In their tutorial, they do caution that just because a grandparent was born in a particular location doesn’t necessarily mean that they were originally from that location. This is particularly true in the past few generations, since the industrial revolution. However, it may still be a useful tool, when taken with the requisite grain of salt.
The third way of creating the underlying population data base is to utilize academically published information or information otherwise available. For example, the Human Genome Diversity Project (HGDP) information which represents 1050 individuals from 52 world populations is available for scrutiny. Ancestry, in their paper, states that they utilized the HGDP data in addition to their own customer database as well as the Sorenson data, which they recently purchased.
Academically published articles are available as well. Family Tree DNA utilizes 52 different populations in their reference data base. They utilize published academic papers and the specific list is provided in their FAQ.
As you can see, there are different approaches and tools. Depending on which of these tools are utilized, the underlying data base may look dramatically different, and the information held in the underlying data base will assuredly affect the results.
Step 2: Your Individual DNA Extraction
This is actually the easy part – where you send your swab or spit off to the lab and have it processed. All three of the main players utilize chip technology today. For example, 23andMe focuses on and therefore utilizes medical SNPs, where Family Tree DNA actively avoids anything that reports medical information, and does not utilize those SNPs.
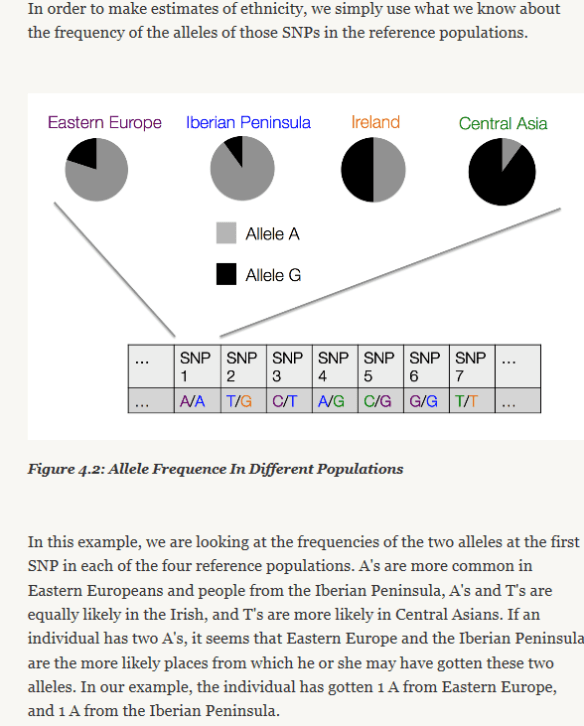
In Ancestry’s white paper, they provide an excellent graphic of how, at the molecular level, your DNA begins to provide information about the geographic location of your ancestors. At each DNA location, or address, you have two alleles, one from each parent. These alleles can have one of 4 values, or nucleotides, at each location, represented by the abbreviations T, A, C and G, short for Thymine, Adenine, Cytosine and Guanine. Based on their values, and how frequently those values are found in comparison populations, we begin to fine correlations in geography, which takes us to the next step.
Step 3: Comparison to Underlying Population Data Base
Now that we have the two individual components in our recipe for ethnicity, a population reference set and your DNA results, we need to combine them.
After DNA extraction, your individual results are compared to the underlying data base. Of course, the accuracy will depend on the quality, diversity, coverage and quantity of the underlying data base, and it will also depend on how many markers are being utilized or compared.
For example, Family Tree DNA utilizes about 295,000 out of 710,000 autosomal SNPs tested for ethnicity prediction. Ancestry’s V1 product utilized about 30,000, but that has increased now to about 300,000 in the 2.0 version.
When comparing your alleles to the underlying data set one by one, patterns emerge, and it’s the patterns that are important. To begin with, T, A, C and G are not absent entirely in any population, so looking at the results, it then becomes a statistics game. This means that, as Ancestry’s graphic, above, shows, it becomes a matter of relativity (pardon the pun), and a matter of percentages.
For example, if the A allele above is shown is high frequencies in Eastern Europe, but in lower frequencies elsewhere, that’s good data, but may not by itself be relevant. However if an entire segment of locations, like a street of DNA addresses, are found in high percentages in Eastern Europe, then that begins to be a pattern. If you have several streets in the city of You that are from Eastern Europe, then that suggests strongly that some of your ancestors were from that region.
To show this in more detailed format, I’m shifting to the third party tool, GedMatch and one of their admixture tools. I utilized this when writing the series, “The Autosomal Me” and in Part 2, “The Ancestor’s Speak,” I showed this example segment of DNA.
On the graph below, which is my chromosome painting of one a small part of one of my chromosomes on the top, and my mother’s showing the exact same segment on the bottom, the various types of ethnicity are colored, or painted.
The grid shows location, or address, 120 on the chromosome and each tick mark is another number, so 121, 122, etc. It’s numbered so we can keep track of where we are on the chromosome.
You can readily see that both of us have a primary ethnicity of North European, shown by the teal. This means that for this entire segment, the results are that our alleles are found in the highest frequencies in that region.
However, notice the South Asian, East Asian, Caucus, and North Amerindian. The important part to notice here, other than I didn’t inherit much of that segment at 123-127 from her, except for a small part of East Asian, is that these minority ethnicities tend to nest together. Of course, this makes sense if you think about it. Native Americans would carry Asian DNA, because that is where their ancestors lived. By the same token, so would Germans and Polish people, given the history of invasion by the Mongols. Well, now, that’s kind of a monkey-wrench isn’t it???
This illustrates why the results may sometimes be confusing as well as how difficult it is to “identify” an ethnicity. Furthermore, small segments such as this are often “not reported” by the testing companies because they fall under the “noise” threshold of between about 5 and 7cM, depending on the company, unless there are a lot of them and together they add up to be substantial.
In Summary
In an ideal world, we would have one resource that combines all of these tools. Of course, these companies are “for profit,” except for National Geographic, and they are not going to be sharing their resources anytime soon.
I think it’s clear that the underlying data bases need to be expanded substantially. The reliability of utilizing contributed pedigrees as representative of a population indigenous to an area is also questionable, especially pedigrees that only reach back two generations.
All of these tools are still in their infancy. Both Ancestry and Family Tree DNA’s ethnicity tools are labeled as Beta. There is useful information to be gleaned, but don’t take the results too seriously. Look at them more as establishing a pattern. If you want to take a deeper dive by utilizing your raw data and downloading it to GedMatch, you can certainly do so. The Autosomal Me series shows you how.
Just keep in mind that with ethnicity predictions, with all of the vendors, as is particularly evident when comparing results from multiple vendors, “your mileage may vary.” Now you know why!
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research
Discover more from DNAeXplained - Genetic Genealogy
Subscribe to get the latest posts sent to your email.





How difficult would it be for one of these firms to withhold 1 million dollars from advertising over two years, and instead allocate those funds to send a researcher/extraction specialist to Poland or Mali or Uruguay to get more samples for the underlying population data set? I understand the process but I’m confused about the rationale for the specific ethnic groups and total population samples being so small. Clearly I’m missing something…. I can’t envision any other US biz entity setting price or any other forecast on such scant data…. So I must be missing why it’s so hard to get more population same. Pls illuminate… Thanks
That’s what Nat Geo is doing – but remember, they are nonprofit. However, the real answer to your question is because there is no profit motivation to do so. The ethnicity percentages are just part of the product and few people are going to purchase it because of that functionality.
Remember, the people in Poland today, and much of Europe are not the same as the people there 100 years ago. There were 2 world wars and most of the Jewish population was wiped out or moved to a different location than before the wars. In addition, many non Jews were displaced and or moved during and after the wars. Some genetic material was wiped out of the gene pool by the holocaust and others were moved to different continents from the Americas to South Africa. This is a much more complicated issue than it appears.
Thanks for another thorough explanation on biogeographical ancestry tests, or autosomal admixture. As we’ve discussed before, I question the paucity and veracity of sample sizes used in all of the current tests, including Geno 2.0.
Geno 2.0, as well as FTDNA uses, as you mention, the Human Genome Diversity Project data from 2005. First one would think that with the incredible progress in genomics in the last eight years, there would be some better samples to use, but I guess funding for these academic projects is limited. HGDP’s samples appear to reflect the interests of its leading researcher, the doyen of population genetics, and thus includes 14 samples from Northern Italy, 8 samples from Tuscany, and 28 samples from Sardinia, the clear favorite of the doyen. He has called Sardinia a unique isolated genetic population; how, from 28 samples out of a population of several million can one claim uniqueness I can’t understand, since no one knows who these 28 people were. No one has obtained their pedigrees to determine ancestry; for all we know they could have just come off the cruise ship from Naples.
Besides these samples form Italy and Sardinia, HGDP includes 24 French, 24 Basque, and my favorite 16 Orcadian, from the Orkney Islands, from whence we get FTDNA’s Orcadian heritage percentages. That’s it for Europe !! No German, no Spanish, no Polish or other Central European, no Scandinavian (except if you really understand the heritage of Orcadian), no British, Irish or Scottish for those of us who would like to fine-tune our heritage. And no North American Native, either.
This is why these autosomal tests, including Geno 2.0, sorry to say, have so little meaning, and are, in my humble opinion, worthless or worse.
Thank you for this article and the imbedded articles and postings. Helps me obtain a better grasp and perspective as someone new to all this. Very nice of you to provide such information.
Jim Stritzel
Pingback: 2013’s Dynamic Dozen – Top Genetic Genealogy Happenings | DNAeXplained – Genetic Genealogy
My question is how can a Black American (Myself) have Near Eastern and Southwest and South Asian.
Various runs of my Raw Dna from ancestry.com shows North African,South Asian,Mediterranean and General Middle Eastern ancestry which i was excited about because i love middle eastern and East Indian culture.
I get results like 5.95% Moroccan , Gedrosia 1.04% , Baloch 1.99% , East African
Could this come from my Saharan- African or European ancestry ?
My European ancestry comes from my maternal great grandfather which is mostly West European Irish British. I know at on point Britain had colonies in India.
My Paternal 3Greatgranfather was from South Carolina my dad spoke about American Indian Ancestry in our family but never specified what tribe or state it came from. I theorized my paternal line may have some connection to the Turks of South Carolina who married American Indians who married Black Americans.My Paternal great grandmother was listed in the census as Basim Read but later it was Betsy and then Bessie.
Gedmatch Oracle Calculator gave me results of 5.95% Lumbee and 6.04% Aleut.
But other calculations listing for American Indian are 1.96% Athabask , 1.09% Pima , and 2.82% Mayan.
I also on test my Ancestry.Com got 1% Polynesian.
But on Gedmatch i get tiny percentages of East Asian, South Asian,Australian, Melanesian,Siberian and South East Asian.
I also Prometheus report which said i carried a rare Gene that is responsible of Native American Myopathy
Many black Americans have admixture from both European and Native American ancestors. Native Americans were enslaved alongside African people. You’re doing all of the right things to understand your heritage.
The whole point of this article is that the sample pool is too small to be reliable. That is why you get all kinds of results.
You are totally correct and you have the last word on this!
Pingback: Ethnicity Percentages – Second Generation Report Card | DNAeXplained – Genetic Genealogy
Pingback: Ethnicity Testing and Results | DNAeXplained – Genetic Genealogy
Pingback: DNAeXplain Archives – Introductory DNA | DNAeXplained – Genetic Genealogy
Pingback: Ethnicity Testing – A Conundrum | DNAeXplained – Genetic Genealogy
Thank you for such a clear explanation of a topic which is confusing to me as a newbie. I worked in genetics in the days before automation, so I have a very basic knowledge, and I was frustrated by the assumptions being made by people in various forums on what their results told them. The detail just didn’t quite ring true with me. I have been searching the internet for a simple, factual explanation of how the process of determining ethnicity works and how confident you should be, and your blog has answered all my questions.
if I have 0.2 black, how many generations back was my black ancestor, assuming it was just 1 person was African?
I recently received my DNA results from Ancestry.com. The results line up with my Family Tree; however, there was a 7% Italy/Greece result that does not line up. I can not find any ancestors from that area in my over 7,000 names database. First question, how many generations back would that 7% cover? Second, could this be an error?
Thank you for your time.
This article will help you I think. https://dna-explained.com/2016/02/10/ethnicity-testing-a-conundrum/
Hello, if my DNA results show that I have less then < .1% of a particular ethnicity is that likely to be an error?
It could be noise but there is no way to tell.
My DNA results show that I have Africa < 1% and Asia < 1% and West Asia < 1% Can you explain to me what that means? I would really appreciate it!
Thank You.
Pingback: Ethnicity and Physical Features are NOT Accurate Predictors of Parentage or Heritage | DNAeXplained – Genetic Genealogy
I am showing I am 95% African. My parents are deceased. But my remaining Paternal Uncle is showing as 96% African. What percentage would my grandparents African percentage possibly be?
They would most likely show something similar.
“African” is very vague though.
I recently got DNA Ethnicity results back from My Heritage DNA. My results come back 49.9% Irish,Scottish, Welsh, 43.7 % Scandinavian and a couple of minor others. My results show No English ethnicity at all.
I can understand having some Scandinavian as ancestors were from Caithness at the top of Scotland, however 43.7 % and no English puzzles me as reliable family trees have relatives from Wolverhampton, Yorkshire, Bury St Edmund and others….from both parents.
I rang My Heritage DNA and they said it was because these genes were not carried down to me, but does not mean I have no English heritage.
Could a 0% result be possible with some many English ancestors ? Would appreciate your help .
Partly, it depends on how far back your English heritage is in your tree. However, ethnicity estimates, from all companies, are just that, estimates. MyHeritage it no different.