Pre-release information from the paper, “Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans” which included results and analysis of DNA sequencing of 24,000 year old skeletal remains of a 4 year old Siberian boy caused quite a stir. Unfortunately, it was also misconstrued and incorrectly extrapolated in some articles. Some people misunderstood, either unintentionally or intentionally, and suggested that people with haplogroups U and R are Native American. That is not what either the prerelease or the paper itself says. Not only is that information and interpretation incorrect, the paper itself with the detailed information wasn’t published until November 20th, in Nature.
The paper is currently behind a paywall, so I’m going to discuss parts of it here, along with some additional information from other sources. To help with geography, the following google map shows the following locations: A=the Altai Republic, in Russia, B=Mal’ta, the location of the 24,000 year old skeletal remains and C=Lake Baikal, the region from where the Native American population originated in Asia.
Nature did publish an article preview. That information is in bold, italics and I will be commenting in nonbold, nonitalics.
The origins of the First Americans remain contentious. Although Native Americans seem to be genetically most closely related to east Asians1, 2, 3, there is no consensus with regard to which specific Old World populations they are closest to4, 5, 6, 7, 8. Here we sequence the draft genome of an approximately 24,000-year-old individual (MA-1), from Mal’ta in south-central Siberia9, to an average depth of 1×. To our knowledge this is the oldest anatomically modern human genome reported to date.
Within the paper, the authors also compare the MA-1 sequence to that of another 40,000 year old individual from Tianyuan Cave, China whose genome has been partially sequenced. This Chinese individual has been shown to be ancestral to both modern-day Asians and Native Americans. This comparison was particularly useful, because it showed that MA-1 is not closely related to the Tianyuan Cave individual, and is more closely related to Native Americans. This means that MA-1’s line and Tianyuan Cave’s line had not yet met and admixed into the population that would become the Native Americans. That occurred sometime later than 24,000 years ago and probably before crossing Beringia into North America sometime between about 18,000 and 20,000 years ago.
The MA-1 mitochondrial genome belongs to haplogroup U, which has also been found at high frequency among Upper Palaeolithic and Mesolithic European hunter-gatherers10, 11, 12, and the Y chromosome of MA-1 is basal to modern-day western Eurasians and near the root of most Native American lineages5.
The paper goes on to say that MA-1 is a member of mitochondrial (maternal) haplogroup U, very near the base of that haplogroup, but without affiliation to any known subclade, implying either that the subclade is rare or extinct in modern populations. In other words, this particular line of haplogroup U has NOT been found in any population, anyplace. According to the landmark paper, “A ‘‘Copernican’’ Reassessment of the Human Mitochondrial DNA Tree from its Root,” by Behar et al, 2012, haplogroup U itself was born about 46,500 years ago (plus or minus 3.200 years) and today has 9 major subclades (plus haplogroup K) and about 300 branching clades from those 9 subclades, excluding haplogroup K.
The map below, from the supplemental material included with the paper shows the distribution of haplogroup U, the black dots showing locations of haplogroup U comparison DNA.
In a recent paper, “Ancient DNA Reveals Key Stages in the Formation of Central European Mitochondrial Genetic Diversity” by Brandt et al (including the National Geographic Consortium) released in October 2013, the authors report that in the 198 ancient DNA samples collected from 25 German sites and compared to almost 68,000 current results, all of the ancient Hunter-Gatherer cultural results were haplogroup U, U4, U5 and U8. No other haplogroups were represented. In addition, those haplogroups disappeared from the region entirely with the advent of farming, shown on the chart below.
So, if someone who carries haplogroup U wants to say that they are distantly related to MA-1 who lived 24,000 years ago who was also related to their common ancestor who lived sometime prior to that, between 24,000 and 50,000 years ago, probably someplace between the Middle East where U was born, Mal’ta, Siberia and Western Europe, they would be correct. They are also distantly related to every other person in the world who carries haplogroup U, and many much more closely that MA-1 whose mitochondrial DNA line is either rare as chicken’s teeth (i.e. never found) or has gone extinct.
Let me be very clear about this, there is no evidence, none, that mitochondrial haplogroup U is found in the Native American population today that is NOT a result of post-contact admixture. In other words, in the burials that have been DNA tested, there is not one example in either North or South America of a burial carrying mitochondrial haplogroup U, or for that matter, male Y haplogroup R. Native American haplogroups found in the Americas remain subsets of mitochondrial haplogroups A, B, C, D and X and Y DNA haplogroups C and Q. Mitochondrial haplogroup M has potentially been found in one Canadian burial. No other haplogroups have been found. Until pre-contact remains are found with base haplogroups other than the ones listed above, no one can ethically claim that other haplogroups are of Native American origin. Finding any haplogroup in a contemporary Native population does not mean that it was originally Native, or that it should be counted as such. Admixture and adoption have been commonplace since Europeans first set foot on the soil of the Americas.
Now let’s talk about the Y DNA of MA-1.
The authors state that MA-1’s results are found very near the base of haplogroup R. They note that the sister lineage of haplogroup R, haplogroup Q, is the most common haplogroup in Native Americans and that the closest Eurasian Q results to Native Americans come from the Altai region.
The testing of the MA-1 Y chromosome was much more extensive than the typical STR genealogy tests taken by consumers today. MA-1’s Y chromosome was sequenced at 5.8 million base pairs at a coverage of 1.5X.
The resulting haplotree is shown below, again from the supplementary material.
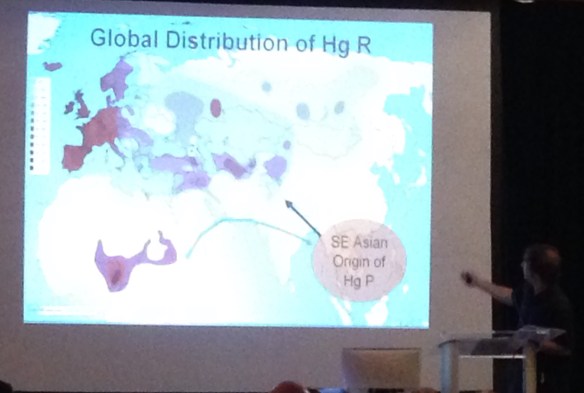
The current haplogroup distribution range for haplogroup R is shown below, again with comparison points as black dots.
The current distribution range for Eurasian haplogroup Q is shown on the map below. Haplogroup Q is the most common haplogroup in Native Americans.
Similarly, we find autosomal evidence that MA-1 is basal to modern-day western Eurasians and genetically closely related to modern-day Native Americans, with no close affinity to east Asians. This suggests that populations related to contemporary western Eurasians had a more north-easterly distribution 24,000 years ago than commonly thought. Furthermore, we estimate that 14 to 38% of Native American ancestry may originate through gene flow from this ancient population. This is likely to have occurred after the divergence of Native American ancestors from east Asian ancestors, but before the diversification of Native American populations in the New World. Gene flow from the MA-1 lineage into Native American ancestors could explain why several crania from the First Americans have been reported as bearing morphological characteristics that do not resemble those of east Asians2, 13.
Kennewick Man is probably the most famous of the skeletal remains that don’t neatly fit into their preconceived box. Kennewick man was discovered on the bank of the Columbia River in Kennewick, Washington in 1996 and is believed to be from 7300 to 7600 years old. His anatomical features were quite different from today’s Native Americans and his relationship to ancient people is unknown. An initial evaluation and a 2010 reevaluation of Kennewick Man let to the conclusion by Doug Owsley, a forensic anthropologist, that Kennewick Man most closely resembles the Ainu people of Japan who themselves are a bit of an enigma, appearing much more Caucasoid than Asian. Unfortunately, DNA sequencing of Kennewick Man originally was ussuccessful and now, due to ongoing legal issues, more technologically advanced DNA testing has not been allowed. Nova sponsored a facial reconstruction of Kennewick Man which you can see here.
Sequencing of another south-central Siberian, Afontova Gora-2 dating to approximately 17,000 years ago14, revealed similar autosomal genetic signatures as MA-1, suggesting that the region was continuously occupied by humans throughout the Last Glacial Maximum. Our findings reveal that western Eurasian genetic signatures in modern-day Native Americans derive not only from post-Columbian admixture, as commonly thought, but also from a mixed ancestry of the First Americans.
In addition to the sequencing they set forth above, the authors compared the phenotype information obtainable from MA-1 to the Tyrolean Iceman, typically called Otzi. You can see Otzi’s facial reconstruction along with more information here. This is particularly interesting in light of the pigmentation change from darker skin in Africa to lighter skin in Eurasia, and the question of when this appearance change occurred. MA-1 shows a genetic affinity with the contemporary people of northern Europe, the population today with the highest frequency of light pigmentation phenotypes. The authors compared the DNA of MA-1 with a set of 124 SNPs identified in 2001 by Cerquira as informative on skin, hair and eye pigmentation color, although they also caution that this method has limited prediction accuracy. Given that, they say that MA-1 had dark hair, skin and eyes, but they were not able to sequence the full set of SNPs. MA-1 also had the SNP value associated with a high risk of male pattern baldness, a trait seldom found in Native American people and was not lactose tolerant, a trait found in western Eurasians. MA-1 also does not carry the mutation associated with hair thickness and shovel shaped incisors in Asians.
The chart below from the supplemental material shows the comparison with MA-1 and the Tyrolean Iceman.
The Tarim Mummies, found in the Tarim Basin in present-day Xinjiang, China are another example of remains that seem out of place. The earliest Tarim mummies, found at Qäwrighul and dated to 1800 BCE, are of a Europoid physical type whose closest affiliation is to the Bronze Age populations of southern Siberia, Kazakhstan, Central Asia, and the Lower Volga.
The cemetery at Yanbulaq contained 29 mummies which date from 1100–500 BCE, 21 of which are Mongoloid—the earliest Mongoloid mummies found in the Tarim Basin—and eight of which are of the same Europoid physical type found at Qäwrighul.
Notable mummies are the tall, red-haired “Chärchän man” or the “Ur-David” (1000 BCE); his son (1000 BCE), a small 1-year-old baby with brown hair protruding from under a red and blue felt cap, with two stones positioned over its eyes; the “Hami Mummy” (c. 1400–800 BCE), a “red-headed beauty” found in Qizilchoqa; and the “Witches of Subeshi” (4th or 3rd century BCE), who wore 2-foot-long (0.61 m) black felt conical hats with a flat brim. Also found at Subeshi was a man with traces of a surgical operation on his neck; the incision is sewn up with sutures made of horsehair.
Their costumes, and especially textiles, may indicate a common origin with Indo-European neolithic clothing techniques or a common low-level textile technology. Chärchän man wore a red twill tunic and tartan leggings. Textile expert Elizabeth Wayland Barber, who examined the tartan-style cloth, discusses similarities between it and fragments recovered from salt mines associated with the Hallstatt culture.
DNA testing revealed that the maternal lineages were predominantly East Eurasian haplogroup C with smaller numbers of H and K, while the paternal lines were all R1a1a. The geographic location of where this admixing took place is unknown, although south Siberia is likely. You can view some photographs of the mummies here.
In closing, the authors of the MA-1 paper state that the study has four important implications.
First, we find evidence that contemporary Native Americans and western Eurasians shareancestry through gene flow from a Siberian Upper Palaeolithic population into First Americans.
Second, our findings may provide an explanation for the presence of mtDNA haplogroup X in Native Americans, which is related to western Eurasians but not found in east Asian populations.
Third, such an easterly presence in Asia of a population related to contemporary western Eurasians provides a possibility that non-east Asian cranial characteristics of the First Americans derived from the Old World via migration through Beringia, rather than by a trans-Atlantic voyage from Iberia as proposed by the Solutrean hypothesis.
Fourth, the presence of an ancient western Eurasian genomic signature in the Baikal area before and after the LGM suggests that parts of south-central Siberia were occupied by humans throughout the coldest stages of the last ice age.
The times, they are a changin’.
Dr. Michael Hammer’s presentation at the 9th Annual International Conference on Genetic Genealogy may shed some light on all of this seeming confusing and somewhat conflicting information.
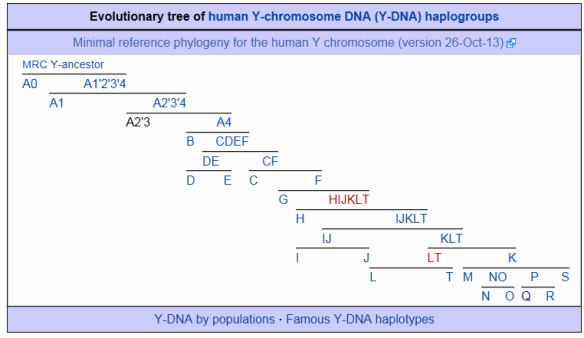
The graphic below shows the Y haplogroup base tree as documented by van Oven.
You can see, in the lower right corner, that Y haplogroup K (not to be confused with mtDNA haplogroup K discussed in conjunction with mtDNA haplogroup U) was the parent of haplogroup P which is the parent of both haplogroups Q and R.
It has always been believed that haplogroup R made its way into Europe before the arrival of Neolithic farmers about 10,000 years ago. However, that conclusion has been called into question, also by the use of Ancient DNA results. You can view additional information about Hammer’s presentation here, but in a nutshell, he said that there is no early evidence in burials, at all, for haplogroup R being in Europe at an early age. In about 40 burials from several location, haplogroup R has never been found. If it were present, especially in the numbers expected given that it represents more than half of the haplogroups of the men of Europe today, it should be represented in these burials, but it is not. Hammer concludes that evidence supports a recent spread of haplogroup R into Europe about 5000 years ago. Where was haplogroup R before spreading into Europe? In Asia.
It appears that haplogroup K diversified in Southeast Asian, giving birth to haplogroups P, Q and R. Dr. Hammer said that this new information, combined with new cluster information and newly discovered SNP information over the past two years requires that haplogroup K be significantly revised. Between the revision of haplogroup K, the parent of both haplogroup R, previously believed to be European, and haplogroup Q, known to be Asian, European and Native, we may be in for a paradigm shift in terms of what we know about ancient migrations and who is whom. This path for haplogroup R into Europe really shouldn’t be surprising. It’s the exact same distribution as haplogroup Q, except haplogroup Q is much less frequently found in Europe than haplogroup R.
What Can We Say About MA-1?
In essence, we can’t label MA-1 as paternally European because of Y haplogroup R which now looks to have had an Asian genesis and was not known to have been in Europe 24,000 years ago, only arriving about 5,000 years ago. We can’t label haplogroup R as Native American, because it has never been found in a pre-Columbian New World burial.
We can say that mitochondrial haplogroup U is found in Europe in Hunter-Gatherer groups six thousand years ago (R was not) but we really don’t know if haplogroup U was in Europe 24,000 years ago. We cannot label haplogroup U as Native because it has never been found in a pre-Columbian New World burial.
We can determine that MA-1 did have ancestors who eventually became European due to autosomal analysis, but we don’t know that those people lived in what is now Europe 24,000 years ago. So the migration might have been into Europe, not out of Europe. MA-1, his ancestors and descendants, may have lived in Asia and subsequently settled in Europe or lived someplace inbetween. We can determine that MA-1’s line of people eventually admixed with people from East Asia, probably in Siberia, and became today’s First People of North and South America.
We can say that MA-1 appears to have been about 30% what is today Western Eurasian and that he is closely related to modern day Native Americans, but not eastern Asians. The authors estimate that between 14% and 38% of Native American ancestry comes from MA-1’s ancient population.
Whoever thought we could learn so much from a 4 year old?
For anyone seriously interested in Native American population genetics, “Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans” is a must read.
It’s been a great month for ancient DNA. Additional recent articles which pertain to this topic include:
http://www.sciencedaily.com/releases/2013/11/131120143631.htm
http://dienekes.blogspot.com/2013/11/ancient-dna-from-upper-paleolithic-lake.html
http://blogs.discovermagazine.com/gnxp/2013/11/long-first-age-mankind/#.Uo0eOcSkrIU
http://cruwys.blogspot.com/2013/11/day-1-at-royal-societys-2013-ancient.html
http://cruwys.blogspot.co.uk/2013/11/day-2-at-royal-societys-2013-ancient.html
http://www.sciencedaily.com/releases/2013/11/131118081251.htm
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research
Discover more from DNAeXplained - Genetic Genealogy
Subscribe to get the latest posts sent to your email.











Roberta, you are absolutely awesome to have put this together! What a labor of love.
Absolutely excellent article! I can always count on DNAeXPLAINED to delve into the details! Learning more and more about Haplo U continues to amaze. ISOGG’S former article on the Cheddar Man and his U Haplogroup (sample taken from a tooth) really had me interested. I told my husband that there is a reason why I love Cheddar Cheese so much. Of course I was kidding! But then again?
Thank you for keeping us so well read, Roberta!
I read one time that the Zuni language origins are unknown. The article also stated that the root words more closely matched Phoenician.
WOW! To find an ancestral Y-DNA haplogroup R*, no subclade, from this era. To think that my R1b1a2a1a1b5 descends from this 4-year-old’s extended family, so to speak, is, as the kids say today, awesome. It appears to push back the time when the haplogroups separated, and implies that there was a lot more population migration around the time of the LGM than what we had assumed. Excellent review Roberta!
Excellent Roberta …….
Pingback: Cradle of Civilization
Pingback: 2013’s Dynamic Dozen – Top Genetic Genealogy Happenings | DNAeXplained – Genetic Genealogy
I’m a bit late in seeing this but it was quite interesting to read due to my own background in linguistics.
Several years ago I had the opportunity to see a talk by Edward Vajda on the Dene-Yeneseian language family – a proposed (though controversial) language family linking a large family of Native American languages (The Dene family which includes Tlingit languages, Eyak languages and Athabaskan languages like Navajo, Plains Apache, etc) with an almost extinct family of Siberian languages (The Yeneseian languages, all of which are extinct except for Ket, which is highly endangered). Many times non-linguists people will purport to link languages with a small sampling of words that seem similar, but Vajda has a large list of corresponding matches between Ket words and words in Dene languages that have undergone regular, systematic sound changes (which is expected in language family). In addition he has posited that Ket and Dene languages share similar morphology and grammatical tendencies, which is more persuasive than a small list of words. There isn’t a consensus in linguistics yet on the existence of the family and there may never be – there’s a huge time scale separating the languages and the Yeneseian languages aren’t well attested. If there were more surviving languages in the family there’d be more data to process. But despite this, the idea isn’t crazy; it’s not pseudo-science (as opposed to the idea that Zuni and Phoenician are related, mentioned by another commenter). This is closest linguists have come to connecting a North American language with one in the Old World (aside from Yupik languages spoken on both sides of the Bering Strait). It’s interesting that the Yeneseain languages were spoken in an area somewhat west of where MA-1 was found.
This isn’t, of course, proof that MA-1 spoke some Proto-Dene-Yeneseian language or anything. But I thought it simply interesting to note the general geographic overlap between linguistic and genetic studies.
I am not an expert in this area, but have lived and worked with the Navajo, Apache, and Hopi people for 21 years. There is a source documentation at the end that links it to Linguistics University of Arizona so hopefully it is valid. I just find this interesting since the Dine people say they are related to the Chinese.
The Tartar Chinese speak the dialect of the Apaches. The Apaches bear a striking resemblance to the Tartar. In about the year 1885, W. B. Horton, who had served as County Superintendent of Schools, at Tucson, was appointed Post Trader at Camp Apache, and went to San Francisco to purchase his stock, where he hired a Chinese cook. His kitchen adjoined his sleeping apartment, and one evening while in his room he heard in the kitchen some Indians talking. Wondering what they were doing there at that hour of the night, he opened the door and found his cook conversing with an Apache. He asked his cook where he had acquired the Indian language. The cook said: “He speak all same me. I Tartar Chinese; he speak same me, little different, not much.” At Williams, in Navajo County, is another Tartar Chinaman, Gee Jim, who converses freely with the Apaches in his native language. From these facts it would seem that the Apache is of Tartar origin. From the fact that the Apache language was practically the same as that of the Tartar Chinese, colour is given to the theory advanced by Bancroft in his “Native Races,” Volume 5, p. 33, et seq., that Western America was “originally peopled by the Chinese, or, at least, that the greater part of the new world civilization may be attributed to these people…” Reference Source: The University of Arizona Library “Books of the South West” Chapter 1, Indians of Arizona:
http://southwest.library.arizona.edu/hav7/body.1_div.1.html
Cecil Lyew(c) Linguistics – Peru
As a linguist, I found the location in Siberia to be interesting. MA-1 was found somewhat East of where the Yeneseian language family was spoken during historical times. The only Yeneseian language still spoken is Ket, which a number of linguists have tried to the Dene languages of North America (Tlingit, Navajo, Apache, and many more). This proposed language family, Dene-Yeneseian, would be the first connection between New World and Old World languages. There have been pseudo-scientific theories in the past, where people have tried to connect disparate languages through short word lists of similar sounding words (with shared/similar meanings) in each language. In linguistics, though, you want lots of corresponding words, with regular sound changes between languages (English: Water German: Wasser, English: Nut German: Nuss), as well as shared grammatical features. Edward Vajda, a proponent of Dene-Yeniseian, has word lists, regular sound correspondences, shared morphological features, and similar grammatical tendencies. The case for Dene-Yeneseian is far from settled, and it may never be fully proven, as there’s only one living Yeneseian language left. If there were data from five or ten languages in Siberia, the added data would make it easier to say “yes” or “no”. That said, there is some evidence. It’s not an unfounded, crackpot theory. And it’s the closest we’ve seen to connecting a Native American language family to an Old World language (aside from Yupik languages spoken on both sides of the Bering Strait). So, I was intrigued that MA-1 should have been located so close to the historical heartland of the Yeniseian languages. This, of course, isn’t to say that MA-1 definitely spoke some form of Proto-Dene-Yeneseian (if that ever existed), but the geographic overlap between the linguistic clues and the genetic clues is interesting.
I lived on the Navajo Reservation for 21 years and one of the common elements in their history is some kind of a disaster and then regrouping in small bands in the area near Aztec, New Mexico. They came together based upon common language. I know in one case a group was stranded do to an ice melt and not being able to go back to there home. There are different stories based up the clans.
There have been attempts to link Yupik to Indo-European but Iroquoian is a far better match. For instance, the PIE word for ‘water’ is ‘akwa’ while PIro is ‘ahwa’. In fact, there are numerous very specific points of simularity between Western Indo-Iranian and Iroquoian languages. Take word for ‘two’ in Mohawk and Nottaway, ‘tekeni’ and ‘dekeni’ respectively. The d-/t- prefix indicates dual case in Iroquoian and so it does in IE laguages. ‘Ek/yek’ is ‘one’ in Persia and Kurdish.
I would also take exception to the statement that it cannot ethically be claimed that YDNA R1 is pre-contact. The truth is that ancient YDNA data is lacking for eastern North American populations. Mainstream genetics researchers such as Schurr maintain that R1a is a founding Native American lineage. My analysis of Hammer’s 2005 data appears to indicate that R1b is twice as abundant in modern NA populations as would predicted from the incidence of YDNA HG I and therefore may not be entirely due to admixture.
Hi Gary,
Could you please provide the citation for where Tad Schurr states that R1a is a founding Native American lineage? Thanks.
Roberta
The estimated age of haplogroup R1a1-M17 is also intriguing. Using the SNP
evolution rate from Underhill et al. (2000), one obtains an age of 13,800 cal yr BP
for this lineage, which falls toward the end of the LGM. These haplotypes also
constitute a distinct branch within R1a and are not especially common in Siberian
populations, although they do occur across a broad geographic area (Lell et al.
2002). Such data suggest that R1a1-M17 haplotypes did not emerge in Siberia
until after the Americas had already been settled, and they, along with C-M130
and P-M45b haplotypes, were brought to the New World through a secondary
expansion of ancient Asian populations (Lell et al. 2002).
THE PEOPLING OF THE NEWWORLD:
Perspectives from Molecular Anthropology
Theodore G. Schurr
Department of Anthropology, University of Pennsylvania, Philadelphia,
Pennsylvania 19104-6398; email: tgschurr@sas.upenn.edu
Hope this helps.
This is really interesting because in 2008, Ripan Malhi also discusses this and suggests that R1a1 is a result of later European DNA introduction. http://www.ncbi.nlm.nih.gov/pubmed/18618732
I know, I read through Theodore Schurr’s findings a couple of times to make sure I was understanding what he meant. It looks like he is including the R1a1 with the second wave for those that were disbursed across North America.
The problem is that we don’t find any in the western or southern areas, just in the northeast, which certainly doesn’t suggest an Asian source. I just dropped Tad and Ripan both notes to see if there is anything new and unpublished. I have never seen R1a or any R in burials, and if they were part of the early Native landscape, you would think they would be found in burials.
Should be interesting.
Since NAGPRA is in place old bones may be difficult to examine. With all of the movement of tribal people post Columbus there has to be a great deal of mixing among the tribes. At one time the Cherokee were supposed to be part of the tribe more to the north.
Yes, the Cherokee are Iroquoian.
Roberta, do you think the NA tribes will ever become enlightened or generous enough to allow testing on some of their NA remains. Is it for a purely religious reasons, or a power thing?? Aren’t they interested in their true origins? Have you ever had the opportunity to talk to a tribal leader, re same??
DNA testing is tied up with a lot of various factors, not the least of which is politics.
Their true origins are in the Americas obviously. And in case you haven’t noticed Native Americans have no power, let alone political power . The Natives already know their origins, and they all arose in the Americas!
What % chance, in your opinion, will they allow testing in the next 20 years? What would it take to give them the incentive? No more questions, I promise……Thanks, as always.
Pingback: Clovis People Are Native Americans, and from Asia, not Europe | DNAeXplained – Genetic Genealogy
Pingback: Clovis People Are Native Americans, and from Asia, not Europe | Native Heritage Project
Pingback: Finding Native American Ethnic Results in Germanic People | DNAeXplained – Genetic Genealogy
Pingback: Ancient DNA Matches – What Do They Mean? | DNAeXplained – Genetic Genealogy
Pingback: More Ancient DNA Samples For Comparison | DNAeXplained – Genetic Genealogy
Pingback: Peopling of Europe 2014 – Identifying the Ghost Population | DNAeXplained – Genetic Genealogy
There was a huge Asian presence in North American as early as the 1500’s:
Asians first arrived in post-Colombian North America aboard the Manila Galleons in the 1580s. Some worked on the Spanish Treasure Fleet from Veracruz which plied the waters of the Caribbean and Gulf of Mexico on the way to Spain.
Starting in 1763, Filipinos sometimes deserted the ships in the Gulf of Mexico to escape the brutalities meted out by the Spanish. These Manilamen (also called “Tagalas”) formed communities in the Louisiana Bayou, building their houses on stilts and kept themselves apart from the rest of Louisiana society. Very few women joined these ships, so the men formed families with Cajun, Native American or Creole women. Saint Malo, one of their fishing villages, was continuously occupied until a hurricane wiped it out in 1915.
In the early 1800s a trickling of Asians entered the South. These included the Siamese Twins, Chang and Eng Bunker, ethnic Chinese from Thailand, who entered the US in 1829. In 1839, they each married white women and settled in North Carolina and became naturalized citizens (requiring their classification as “white”). They even owned black slaves, and their sons fought on the Confederate side of the Civil War.
In the middle 1800s Chinese-American immigration exploded. Over 300,000 entered California between the California Gold Rush (1848) and the Chinese Exclusion Act (1882).
In the late 1860s they started coming South after:
the Civil War freed the slaves (1865),
the Burlingame Treaty (1868) expanded Chinese immigration, and
the completion of the Transpacific Railroad (1869) was followed by increasing racial violence in the West.
the completion of the Suez Canal (1869) facilitated trans-Atlantic travel from Asia.
Southern plantation owners, hearing of how effective Chinese labour was in building the railroads in the West and working on plantations in the Caribbean, devised schemes to lure Chinese to come to the South to replace black slaves. They believed non-citizens who could not vote could be controlled more easily than freed slaves. They even sent delegations to China to recruit labour.
By the 1870s, thousands of Chinese were working on plantations in Mississippi, Arkansas and Louisiana, in the port city of New Orleans, and even in the cotton fields of South Carolina and Georgia.
However, the Burlingame treaty required employers to engage the Chinese with labour contracts, which they soon learned the white plantation owners had no intention to honour. They were treated as nothing more than slaves. They went on strike. When that did not work, they fled the plantations. The Chinese Exclusion Act then abruptly halted Southerners’ ability to recruit additional Chinese labour after 1882.
By the late 1880s, few if any Chinese were still working on southern plantations. Whites viewed the labour importation scheme as a complete failure.
Some Chinese left the South, particularly to cities in the East and Midwest, but some remained and went onto other occupations, such as grocers to black sharecroppers.
Enjoyed the article and thank you for posting it. One question; what other explanation would account for R1 Y-DNA being in such large percentage of (for example) Cherokee paternal ancestry. If this was of a post Columbian origin, surely it would be R1b and sub clades there of, or perhaps a small bit of R1a. Hope you post your comments, would benefit from your insight.
Pingback: DNAeXplain Archives – General Information Articles | DNAeXplained – Genetic Genealogy
Pingback: Ethnicity Testing and Results | DNAeXplained – Genetic Genealogy
Pingback: Native American Haplogroup X2a – Solutrean, Hebrew or Beringian? | DNAeXplained – Genetic Genealogy
Pingback: Native American Haplogroup X2a – Solutrean, Hebrew or Beringian? | Native Heritage Project
Roberta, do you know what the Tianyuan Cave’s individual’s haplogroups were?
I don’t, off the top of my head. I would have to look that up.
Hi, Just found your blog. THANK YOU!!
I just did mtDNA with Family Tree because we have a maternal great great gMom who supposedly was Native Am (born Texas +- 1875). We show as U4, specifically U4b1b1. Family Tree shows for HVR1 – U4 for US Native Am, 8 matches, and U4b1b1, 2 matches; the HVR1 and HVR2 give 1 match for U4b1b1 US Native Am. The HVR1, HVR2, and Coding matches for U4b1b1 = 1.
What I understand you to say: the U4, my maternal lineage, came into the US east to west over the Atlantic via a European woman (women?) who co-mixed with Native Americans which is how I ended up (supposedly) with a maternal gg gMother who is Native American but with U4 (U4b1b1) mtDNA haplogroup?? Or, why are there Native Americans with the U4, specifically U4b1b1 haplogroup? [What’s confusing me, the HVR 1, HVR 2, and Coding Matches, are they current, for me paired to those listed for my results?? I’m not sure what this means as I have HVR 1 U4b1b1 matches in England, Ireland, Germany, France, etc.??
My sister points out – the fluidly of cultures in the US 1600-1800. I recommend The Unredeemed Captive by John Demos: an extraordinary read about race, culture, religion, and international politics (England, France), 1680-1720, between Boston and Montreal. Yes, people were raiding, killing each other, but there were a lot of people who intermarried into tribes or who carried on trading relations, etc. Thinking about people caught up in their own lives through this 300 year lense of time – biases give way to nuanced thought.
I do Personalized DNA Reports for people, if you’re interested, where I explain about haplogroups and such. http://www.dnaxplain.com/shop/features.aspx