During the Family Tree DNA Conference, Dr. Miguel Vilar, the Scientific Data Manager for the Explorer Programs was kind enough to give us an update on the Genographic project. One of the things that he mentioned was that no overarching paper had been written about the completed Geno 1.0 phase of the project, although that has been discussed. He did say that a total of 42 papers have been written by the Genographic Consortium as the result of the Genographic project, to date, and that there are several more in the pipeline.
During the Family Tree DNA Conference, Dr. Miguel Vilar, the Scientific Data Manager for the Explorer Programs was kind enough to give us an update on the Genographic project. One of the things that he mentioned was that no overarching paper had been written about the completed Geno 1.0 phase of the project, although that has been discussed. He did say that a total of 42 papers have been written by the Genographic Consortium as the result of the Genographic project, to date, and that there are several more in the pipeline.
As follow-up to that comment, Dr. Vilar was kind enough to provide a list of the papers along with a short description of the findings in each one. Thank you to both Dr. Vilar and National Geographic for sharing.
Personally, I take a great deal of pleasure and satisfaction in knowing that I was (and am) in a cumulative way a small part of this amazing, ongoing project. For anyone who has not yet, but would like to participate in testing, the Genographic 2.0 project is ongoing.
The Genographic Consortium has published 42 scientific papers, and other manuscripts are in advanced stages of preparation. Below are the titles and references plus short descriptions of the major findings, compliments of Dr. Vilar.
2007
1. Behar, D. M., Rosset, S., Blue-Smith, J., Balanovsky, O., Tzur, S., Comas, D., Mitchell, R. J., Quintana-Murci, L., Tyler-Smith, C., Wells, R. S., and The Genographic Consortium. 2007. The Genographic Project public participation mitochondrial DNA database. PLoS Genetics 3: 1083-1095.
- This paper establishes Genographic’s database as the new standard mtDNA data repository and reports a new “Nearest Neighbor” statistical method for improved haplogroup classification, presenting learned experience from the public part of the project. It also makes publicly available a portion of the Genographic database, a process that will continue throughout project duration. This technical paper has been crucial in establishing the project’s importance in the scientific community.
2008
2. Gan, R. J., Pan, S. L., Mustavich, L. F., Qin, Z. D., Cai, X. Y., Qian, J., Liu, C. W., Peng, J. H., Li, S. L., Xu, J. S., Jin, L., Li, H., and The Genographic Consortium. 2008. Pinghua population as an exception of Han Chinese’s coherent genetic structure. Journal of Human Genetics 53: 303-313.
- The Han Chinese are the largest ethnic group in the world with more than 1.3 billion people, comprising 19 percent of the world population. Chinese is the language spoken by this ethnic group, which can be classified into 10 major dialects. This paper focuses on studying the genetic structure of the people speaking one of these dialects, the Pinghua people. When the genetic structure of Pinghua people was compared to the rest of the Han Chinese populations, it was observed that Pinghua populations did not directly descend from Han Chinese, who originated in the north, but from other southern populations. Thus, from a genetic point of view, the Pinghua populations represent an exception to the rest of Han Chinese populations. These results can be explained if ancestral populations of Pinghua people were not replaced by Han Chinese population, but if they assimilated the Han Chinese language and culture.
3. Zalloua, P. A., Xue, Y., Khalife, J., Makhoul, N., Debiane, L., Platt, D. E., Royyuru, A. K., Herrera, R. J., Soria Hernanz, D. F., Blue-Smith, J., Wells, R. S., Comas, D., Bertranpetit, J., Tyler-Smith, C., and The Genographic Consortium. 2008. Y-chromosomal diversity in Lebanon is structured by recent historical events. American Journal of Human Genetics 82: 873-882.
- Lebanon is a small country in the Middle East inhabited by almost 4 million people from a wide variety of ethnicities and religions. The results of this paper indicate that male genetic variation within Lebanon is strongly structured by religion. This unusual situation can be accounted for by two major known historical migrations into Lebanon. The Islamic expansion from the Arabian Peninsula beginning in the 7th century introduced genetic lineages typical of the Arabian peninsula into Lebanese Muslims, while the crusader activity in the 11th-13th centuries introduced Western European lineages into Lebanese Christians.
4. Behar, D. M., Villems, R., Soodyall, H., Blue-Smith, J., Pereira, L., Metspalu, E., Scozzari, R., Makkan, H., Tzur, S., Comas, D., Bertranpetit, J., Quintana-Murci, L., Tyler-Smith, C., Wells, R. S., Rosset, S., and The Genographic Consortium. 2008. The dawn of human matrilineal diversity. American Journal of Human Genetics 82: 1130-1140.
- African genetic diversity is unlike that found anywhere else in the world. This paper seeks to make sense of some of the most fundamental questions surrounding our earliest ancestors on the continent. Where specifically did we originate in Africa? Was it from a single group or the result of many? When do we first see African lineages appear outside of Africa? About 350 novel mitochondrial whole-genome sequences were included — doubling the existing published dataset — and the paper presented a new tree of African mtDNA diversity, reporting many novel African lineages for the first time. This paper provides an age estimate for the earliest split of humans in East Africa as one group headed south and was subsequently isolated. It explains that all humans came from a single population that split into two groups, shows that more than 99 percent of all living humans descend from one of these two groups, and suggests historical reasons for why genetic mixture did not exist between these ancient populations. It also presents evidence for the emergence of these early lineages into the Middle East and the origins of the two major non-African groups, M and N, respectively. The paper received considerable media attention — approximately 275 articles — including substantial pieces in the Economist and on CNN/BBC online.
5. Behar, D. M., Blue-Smith, J., Soria-Hernanz, D. F., Tzur, S., Hadid, Y., Bormans, C., Moen, A., Tyler-Smith, C., Quintana-Murci, L., Wells, R. S., and The Genographic Consortium. 2008. A novel 154-bp deletion in the human mitochondrial DNA control region in healthy
individuals. Human Mutation 29: 1387-1391.
- This paper describes a novel deletion of 154 base pairs within the control region of the human mitochondrial genome that was originally identified in an anonymous Japanese public participant. It was demonstrated that this deletion is a heritable character since it was transmitted from the participant’s mother to her two sons. This is the first time that such a large deletion located in this specific portion of the control region has been observed to not have negative effects in the health of the carriers. The identification of this large heritable deletion in healthy individuals challenges the current view of the control region as playing a crucial role in the replication and regulation of the mitochondrial genome. It is anticipated that this finding will lead to further research on the reported samples in an attempt to increase our understanding of the role of specific sequences within the control region for mtDNA replication. Finally, this paper illustrates the importance of creating a large database of human genetic variation in order to discover rare genetic variants that otherwise would remain unidentified. The discovery of such rare mtDNA haplotypes will be important to identifying the relative power of adaptive and non-adaptive forces acting on the evolution of the mtDNA genome.
6. Parida, L., Melé, M., Calafell, F., Bertranpetit, J., and The Genographic Consortium. 2008. Estimating the ancestral recombinations graph (ARG) as compatible networks of SNP patterns. Journal of Computational Biology 15: 1133-1153.
- Traditionally the nonrecombinant, maternally inherited (mtDNA) and paternally inherited (Y chromosome) genomes have been widely used for phylogenetic and evolutionary studies in humans. However, these two genomes only represent 1 percent of the total genetic variation within an individual, and sampling just these two loci is inadequate to reconstruct with any precision the time-depth and pattern of human evolution. The scope of this paper is to elaborate on a mathematical algorithm that includes recombination patterns among human populations. This approach will allow us to use the rest of the recombining genome to reconstruct more accurately the patterns of human migration.
7. Rossett, S., Wells, R. S., Soria-Hernanz, D. F., Tyler-Smith, C., Royyuru, A. K., Behar, D. M., and The Genographic Consortium. 2008. Maximum-likelihood estimation of site-specific mutation rates in human mitochondrial DNA from partial phylogenetic classification. Genetics 180: 1511-1524.
- This paper presents novel algorithms to estimate how frequently each base pair of the hypervariable region of the mtDNA changes. Implementations of these algorithms will help to better investigate functionality in the mtDNA and improve current classification of mtDNA haplogroups.
8. Zalloua, P. A., Platt, D. E., El Sibai, M., Khalife, J., Makhoul, N., Haber, M., Xue, Y., Izaabel, H., Bosch, E., Adams, S. M., Arroyo, E., López-Parra, A. M., Aler, M., Picornell, A., Ramon, M., Jobling, M. A., Comas, D., Bertranpetit, J., Wells, R. S., Tyler-Smith, C., and The Genographic Consortium. 2008. Identifying genetic traces of historical expansions: Phoenician footprints in the Mediterranean. American Journal of Human Genetics 83: 633-642.
- The Phoenicians gave the world the alphabet and a love of the color purple, and this study shows that they left some of their genes as well. The paper shows that as many as one in 17 men in the Mediterranean basin may have a Phoenician as a direct male-line ancestor, using a novel analytical method for detecting the subtle genetic impact of historical population migrations. Its first application has been to reveal the genetic legacy of the Phoenicians, an intriguing and mysterious first-millennium B.C. trading empire. From their base in present-day Lebanon, the Phoenicians expanded by sea throughout the Mediterranean, founding colonies as far as Spain and North Africa, where their most powerful city, Carthage, was located. The world’s first “global capitalists,” the Phoenicians controlled trade throughout the Mediterranean basin for nearly a thousand years until their conquest by Rome in the 2nd century B.C. Over the ensuing centuries, much of what was known about this enigmatic people was lost or destroyed. This paper received substantial international and domestic press coverage, including an article in The New York Times.
2009
9. Parida, L., Javed, A., Melé, M., Calafell, F., Bertranpetit, J., and The Genographic Consortium. 2009. Minimizing recombinations in consensus networks for phylogeographic studies. BMC Bioinformatics 10: Article S72.
- This paper implements a new mathematical model to identify recombination spots in human populations to infer ancient recombination and population-specific recombination on a portion of the X chromosome. The results support the widely accepted out-of-Africa model of human dispersal, and the recombination patterns were capable of detecting both continental and population differences. This is the first characterization of human populations based on recombination patterns.
10. El-Sibai, M., Platt, D. E., Haber, M., Xue, Y., Youhanna, S. C., Wells, R. S., Izaabel, H., Sanyoura, M. F., Harmanani, H., Ashrafian Bonab, M., Behbehani, J., Hashwa, F., Tyler-Smith, C., Zalloua, P. A., and The Genographic Consortium. 2009. Geographical structure of the Y-chromosomal genetic landscape of the Levant: A coastal-inland contrast. Annals of Human Genetics 73: 568-581.
- This paper examines the male-specific phylogeography of the Levant and its surroundings. The Levant lies in the eastern Mediterranean region, south of the mountains of south Turkey and north of the Sinai Peninsula. It was found that the Levantine populations cluster together when considered against a broad Middle-East and North African background. However, within Lebanon there is a coastal-inland (east-west) pattern in the diversity and frequency of several Y haplogroups. This pattern is likely to have arisen from differential migrations, with different lineages introduced from the east and west.
2010
11. Haak, W., Balanovsky, O., Sanchez, J. J., Koshel, S., Zaporozhchenko, V., Adler, C. J., Der Sarkissian, C. S. I., Brandt, G., Schwarz, C., Nicklisch, N., Dresely, V., Fritsch, B., Balanovska, E., Villems, R., Meller, H., Alt, K. W., Cooper, A., and The Genographic Consortium. 2010. Ancient DNA from European Early Neolithic farmers reveals their Near Eastern affinities. PLoS Biology 8: Article e1000536.
- The nature and speed of the Neolithic transition in Europe is a matter of continuing debate. In this paper, new genetic analyses based on ancient human remains from the earliest farming culture in Central Europe known as the Linear Pottery Culture (5,500-4,900 years ago) indicate a shared genetic maternal affinity with modern-day Near East and Anatolia, and therefore they likely came from the Middle East. However, these lineages from the earliest agriculturalists were also distinct from the current genetic lineages observed in European populations, indicating that major demographic events continued in Europe during the Neolithic. These results point out the importance of using ancient DNA to better understand past demographic events.
12. Melé, M., Javed, A., Pybus, M., Calafell, F., Parida, L., Bertranpetit, J., and The Genographic Consortium. 2010. A new method to reconstruct recombination events at a genomic scale. PLoS Computational Biology 6: Article e1001010.
- A chromosomal recombination event creates a junction between two parental sequences. These recombinant sequences are transmitted to subsequent generations, and recombination is one of the main forces molding human genetic diversity. However, the information about genetic relationships among populations given by these events is usually overlooked due to the analytical difficulty of identifying the history of recombination events. This paper validates and calibrates the IRiS software for inferring the history of recombination events, allowing the creation of novel recombinational “markers” known as recotypes, which can be analyzed in a similar way to standard mutational markers.
13. Qin, Z., Yang, Y., Kang, L., Yan, S., Cho, K., Cai, X., Lu, Y., Zheng, H., Zhu, D., Fei, D., Li, S., Jin, L., Li, H., and The Genographic Consortium. 2010. A mitochondrial revelation of early human migrations to the Tibetan Plateau before and after the Last Glacial Maximum. American Journal of Physical Anthropology 143: 555-569.
- The Tibetan Plateau was long considered one of the last areas to be populated by modern humans. Recent archaeological, linguistic and genetic findings have challenged this view. In this paper, maternal lineages of 562 individuals from nine different regions within Tibet have been analyzed to further investigate the timing and routes of entry of humans into the plateau. The maternal diversity in Tibet primarily reflects northern East Asian ancestry, likely reflecting a population expansion from this region into the plateau prior to the Last Glacial Maximum (LGM) ~18,000 years ago. In addition, the highest diversity was concentrated in the southern part of the plateau, indicating that this region probably acted as a population refugium during the LGM and the source of a post-LGM expansion within the plateau.
14. Zhadanov, S. I., Dulik, M. C., Markley, M., Jennings, G. W., Gaieski, J. B., Elias, G., Schurr, T. G., and The Genographic Project Consortium. 2010. Genetic heritage and native identity of the Seaconke Wampanoag tribe of Massachusetts. American Journal of Physical Anthropology 142: 579-589.
- The biological ancestry of the Seaconke Wampanoag tribe, a group of Native American clans in southern Massachusetts, reflects the genetic consequences of epidemics and conflicts during the 16th century that decimated their population, reducing them from an estimated 12,000 individuals at the beginning of the century to less than 400 at the end. The majority of the paternal and maternal lineages in present-day Seaconke Wampanoag, however, belong to West Eurasian and African lineages, revealing the extensive interactions with people from different ancestries that settled the region during the past four centuries.
2011
15. Adler, C. J., Haak, W., Donlon, D., Cooper, A., and The Genographic Consortium. 2011. Survival and recovery of DNA from ancient teeth and bones. Journal of Archaeological Science 38: 956-964.
- The recovery of genetic material from ancient human remains depends on the sampling methods used as well as the environment where the human material was preserved. The results presented in this study quantify the damage caused to ancient DNA by various methods of sampling teeth and bones. The negative impact is minimized if very low drill speeds are used during DNA extraction, increasing both the quantity and quality of material recovered. In addition, the mtDNA content of tooth cementum was five times higher than other commonly used methods, making this component the best place to sample ancient DNA. These conclusions will help to guide future sampling of DNA from ancient material.
16. Haber, M., Platt, D. E., Badro, D. A., Xue, Y., El-Sibai, M., Ashrafian Bonab, M., Youhanna, S. C., Saade, S., Soria-Hernanz, D. F., Royyuru, A., Wells, R. S., Tyler-Smith, C., Zalloua, P. A., and The Genographic Consortium. 2011. Influences of history, geography, and religion on genetic structure: The Maronites in Lebanon. European Journal of Human Genetics 19: 334-340.
- Cultural patterns frequently leave genetic traces. The aim of this study was to explore the genetic signature of the establishment of religious communities in a region where some of the most influential world religions originated, using the Y chromosome as an informative male-lineage marker. The analysis shows that the religions in Lebanon were adopted within already distinguishable communities. Differentiation appears to have begun before the establishment of Islam and Christianity, dating to the Phoenician period, and isolation continued during the period of Persian domination. Religious affiliation served to reinforce the genetic signatures of pre-existing population differentiation.
17. Martínez-Cruz, B., Ziegle, J., Sanz, P., Sotelo, G., Anglada, R., Plaza, S., Comas, D., and The Genographic Consortium. 2011. Multiplex single-nucleotide polymorphism typing of the human Y chromosome using TaqMan probes. Investigative Genetics 2: Article 13.
- This paper presents a robust and accurate Y-chromosome multiplex assay that can genotype in a single reaction 121 markers distinguishing most of the haplogroups and subhaplogroups observed in European populations. The assay was >99 percent accurate in assigning haplogroups, minimizing sample handling errors that can occur with several independent TaqMan reactions.
18. Jota, M. S., Lacerda, D. R., Sandoval, J. R., Vieira, P. P. R., Santos-Lopes, S. S., Bisso-Machado, R., Paixão-Cortes, V. R., Revollo, S., Paz-y-Miño, C., Fujita, R., Salzano, F. M., Bonatto, S. L., Bortolini, M. C., Tyler-Smith, C., Santos, F. R., and The Genographic Consortium. 2011. A new subhaplogroup of Native American Y-chromosomes from the Andes. American Journal of Physical Anthropology (published online Sept. 13, 2011.)
- Almost all Y chromosomes in South America fall into a single haplogroup, Q1a3a. This paper presents a new single nucleotide polymorphism (SNP) in the Q1a3a lineage that is specific to Andean populations, allowing more accurate inferences of the population history of this region. This novel marker is estimated to be ~5,000 years old, consistent with an ancient settlement of the Andean highlands.
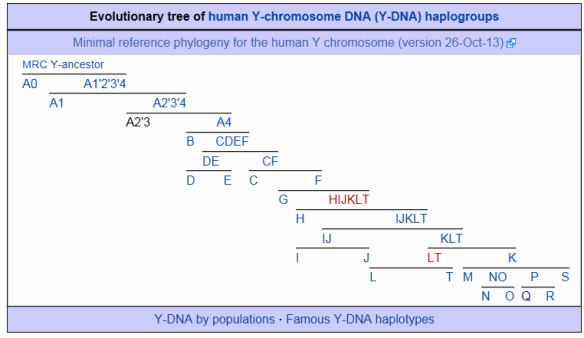
19. Yan, S., Wang, C. C., Li, H., Li, S. L., Jin, L., and The Genographic Consortium. 2011. An updated tree of Y-chromosome Haplogroup O and revised phylogenetic positions of mutations P164 and PK4. European Journal of Human Genetics 19: 1013-1015.
- Y-chromosome Haplogroup O is the dominant Y-chromosome lineage in East Asians, carried by more than a quarter of all males on the world. This study revises the haplogroup O phylogeny, using several recently discovered markers. The newly generated tree for this haplogroup will lead to a more detailed understanding of the population history of East Asia.
20. Yang, K., Zheng, H., Qin, Z., Lu, Y., Farina, S. E., Li, S., Jin, L., Li, D., Li, H., and The Genographic Consortium. 2011. Positive selection on mitochondrial M7 lineages among the Gelong people in Hainan. Journal of Human Genetics 56: 253-256.
- The Gelong people migrated in the last 1,000 years from Guizhou province in southern China to Hainan island (the hottest province in China). The genetic structure of the Gelong people showed a clearly sex-biased pattern of admixture with the indigenous Hainan population (Hlai people), with 30.7 percent of the maternal lineages being of Hainan origin in contrast to 4.9 percent of the paternal lineages. This striking pattern is partially explained through the action of selection on the M7 Hainan autochthonous maternal lineages, leading to their expansion in the admixed population. This may be due to some selective advantage provided by the M7 lineages in the tropical Hainan climate. Future whole mtDNA genome sequencing of these M7 lineages may reveal their functional relevance and the mechanism involved in human adaptation to tropical climates.
21. Balanovsky, O., Dibirova, K., Dybo, A., Mudrak, O., Frolova, S., Pocheshkhova, E., Haber, M., Platt, D., Schurr, T., Haak, W., Kuznetsova, M., Radzhabov, M., Balaganskaya, O., Druzhinina, E., Zakharova, T., Soria Hernanz, D. F., Zalloua, P., Koshel, S., Ruhlen, M., Renfrew, C., Wells, R. S., Tyler-Smith, C., Balanovska, E., and The Genographic Consortium. 2011. Parallel evolution of genes and languages in the Caucasus region. Molecular Biology and Evolution 28: 2905-2920.
- The Caucasus region harbors some of the highest linguistic diversity on Earth, leading to the moniker “The Mountain of Languages.” To investigate the forces that may have molded Caucasian linguistic patterns, the Genographic team studied Y-chromosome variation in 1,525 men from 14 populations in the Caucasus. The Y-chromosome lineages found in the Caucasus originated in the Near East and were introduced to the Caucasus in the late Upper Paleolithic or early Neolithic periods. This initial settlement was followed by a high degree of population isolation due to the mountainous terrain. Comparisons between the genetic and linguistic trees showed a striking correspondence between the topology and divergence times for the two, revealing a parallel evolution of genes and languages in the Caucasus in the past few millennia. This high degree of correspondence between genetic and linguistic patterns has not been seen in other regions of the world.
22. Gaieski, J. B., Owings, A. C., Vilar, M. G., Dulik, M. C., Gaieski, D. F., Gittelman, R. M., Lindo, J., Gau, L., Schurr, T. G., and The Genographic Consortium. 2011. Genetic ancestry and indigenous heritage in a Native American descendant community in Bermuda. American Journal of Physical Anthropology 146: 392-405.
- Bermuda is an isolated group of islands in the middle of the Atlantic settled during the 17th century by Western Europeans along with African and Native American slaves. The pattern of Y-chromosome and mitochondrial DNA diversity was studied in 111 members of a “native” community on St. David’s Island. Two-thirds of the paternal lineages are of European origin, while two-thirds of the mitochondrial DNA lineages are African. In contrast to other English-speaking communities in the Americas, however, the majority of St. David’s maternal lineages appear to derive from central and southern Africa, regions that historically were controlled by Portuguese slave traders. It is likely that the English settlers of Bermuda obtained slaves from these Portuguese sources. Despite genealogical records and oral traditions indicating significant arrivals of Native Americans as labor force, the proportion of Native American lineages was less than 2 percent on both the paternal and maternal sides. This study gives new insights into the complex history of colonization and migration in the Caribbean.
23. Cai, X., Qin, Z., Wen, B., Xu, S., Wang, Y., Lu, Y., Wei, L., Wang, C., Li, S., Huang, X., Jin, L., Li, H., and The Genographic Consortium. 2011. Human Migration through bottlenecks from Southeast Asia into East Asia during Last Glacial Maximum revealed by Y chromosomes. PLoS ONE 6: e24282. doi:10.1371/journal.pone.0024282
- The number and timing of the initial migrations to East Asia remain unresolved. This paper studied the Y-chromosome diversity in Mon-Khmer (MK)- and Hmong-Mien (HM)-speaking populations who are believed to be the source populations of other East Asians. The pattern of diversity for the O3a3b-M7 and O3a3c1-M117 lineages among MK, HM and other East Asian populations suggests an early unidirectional diffusion from Southeast Asia northward into East Asia around the time of the Last Glacial Maximum (~18,000 years ago). The ancestral population sizes of these first colonizers are believed to have gone through drastic reductions due to the barriers imposed by the geographic conditions (mountains and jungle) and the colder climate at the time of the migration. This “serial bottleneck” effect has left a distinctive genetic pattern in the present-day populations of East Asia, revealing their past demographic history.
24. Melé, M., Javed, A., Pybus, M., Zalloua, P., Haber, M., Comas, D., Netea, M. G., Balanovsky, O., Balanovska, E., Jin, L., Yang, Y., Pitchappan, R. M., Arunkumar, G., Parida, L., Calafell, F., Bertranpetit, J., and The Genographic Consortium. 2011. Recombination gives a new insight in the effective population size and the history of the Old World human populations. Molecular Biology and Evolution (published online Sept. 1, 2011.) doi:10.1093/molbev/msr213
- The IRiS method (described in paper 12) was used to assess the patterns of recombination on the X chromosome in 30 populations from Africa, Europe and Asia. The results suggest that the ancestors of non-African populations first left Africa in a single coastal migration across the Bad-el-Mandeb strait rather than through the Sinai Peninsula. The method allowed the team to estimate that sub-Saharan ancestral population sizes were four times greater than those in populations outside of Africa, while Indian ancestral sizes were the greatest among Eurasians. These results suggest that Indian populations played a major role in the expansions of modern humans to the rest of the world.
25. Javed, A., Melé, M., Pybus, M., Zalloua, P., Haber, M., Comas, D., Netea, M. G., Balanovsky, O., Balanovska, E., Jin, l., Yang, Y., Arunkumar, G., Pitchappan, R., Bertranpetit, J., Calafell, F., Parida, L., and The Genographic Consortium. 2011. Recombination networks as genetic markers in a human variation study of the Old World. Human Genetics (first published online Oct. 18, 2011.)
- An expanded analysis of the recombination dataset published in abbreviated form in paper 24, analyzing three additional populations. The conclusions outlined in paper 24 are bolstered through the more thorough presentation of the results.
2012
26. Behar DM, Harmant C, Manry J, van Oven M, Haak W, Martinez-Cruz B, Salaberria J, Oyharçabal B, Bauduer F, Comas D, Quintana-Murci L; Genographic Consortium. 2012. The Basque paradigm: genetic evidence of a maternal continuity in the Franco-Cantabrian region since pre-Neolithic times. American Journal of Human Genetics 9;90(3):486-93.
- This study focus on the maternal genetic diversity of Basques, the last European population to have kept a pre-Indo European language, to increase knowledge of the origins of the Basque people and, more generally, on the role of the Franco-Cantabrian refuge in the post-glacial repopulation of Europe. The maternal ancestry of 908 Basque and non-Basque individuals from the Great Basque Country and adjacent regions were studied plus 420 complete mtDNA genomes within haplogroup H. The results identified six mtDNAhaplogroups autochthonous to the Franco-Cantabrian region and, more specifically, to Basque-speaking populations. Further, expansion of these haplogroups were estimated at ~4,000 ybp with a separation from the general European gene pool to have happened ~8,000 ybp predating the Indo-European arrival to the region. Thus, the results clearly support the hypothesis of a partial genetic continuity of contemporary Basques with the indigenous Paleolithic settlers of their homeland.
27. Martínez-Cruz B, Harmant C, Platt DE, Haak W, Manry J, Ramos-Luis E, Soria-Hernanz DF, Bauduer F, Salaberria J, Oyharçabal B, Quintana-Murci L, Comas D; the Genographic Consortium. Evidence of pre-Roman tribal genetic structure in Basques from uniparentally inherited markers. Molecular Biology and Evolution (published online March 12, 2012) doi: 10.1093/molbev/mss091.
- Basques have received considerable attention from anthropologists, geneticists and linguists during the last century due to the singularity of their language and to other cultural and biological characteristics. Despite the multidisciplinary efforts performed to address the questions of the origin, uniqueness and heterogeneity of Basques, the genetic studies performed up to now have suffered from a weak study-design where populations are not analyzed in an adequate geographic and population context. To address the former questions and to overcome these design limitations, uniparental genomes (Y chromosome and mitochondrial DNA) of ~900 individuals from 18 populations were analyzed, including those where Basque is currently spoken and surrounding populations where Basque might have been spoken in historical times. Results situate Basques within the western European genetic landscape, although with less external influences than other Iberians and French populations. In addition, the genetic heterogeneity and structure observed in the Basque region results from pre-Roman tribal structure related to geography and is linked to the increased complexity of emerging societies during the Bronze Age. The rough overlap of tribal and current dialect limits supports the notion that the environmental diversity in the region has played a recurrent role in cultural differentiation and ethnogenesis at different time periods.
28. Kang, L., Lu, Y., Wang, C., Hu, K., Chen, F., Liu, K., Li, S., Jin, L., Li, H., and The Genographic Consortium. 2012. Y-chromosome O3 Haplogroup diversity in Sino-Tibetan populations reveals two migration routes into the Eastern Himalayas. Annals of Human Genetics 76: 92–99.
- This paper further explores the question of how Himalayas was populated by studying the genetic diversity of the paternal lineages of two ethnic groups from the eastern Himalayas: the Luoba and Deng. These two Sino-Tibetan speaking groups exhibited a distinct genetic composition indicating different genetic origins. The paternal diversity of the Louba people indicates past gene flow from Tibetans as well as from western and north Eurasian people. In contrast, Deng exhibited lineages similar to most of Sino-Tibetans from the east. The overall lowest diversity observed in the eastern Himalayas suggests that this area was the end point of two migratory routes of Sino-Tibetans from north China around 2,000-3,000 years ago. These date estimates also agrees with the historical records.
29. Lu, Y., Wang, C., Qin, Z., Wen, B., Farina, S. E., Jin, L., Li, H., and The Genographic Consortium. 2012. Mitochondrial origin of the matrilocal Mosuo people in China. Mitochondrial DNA 23: 13–19
- The Mosuo people currently live around the Lugu Lake on the border of the Yunan and Sichuan provinces of China and they are the last matrilocal population in the main land of the country. To investigate the maternal history of this ethnic group, partial genetic sequences of the mitochondria (a maternally inherited genome) were studied among Mosuo people and other larger surrounding ethnic groups. Groups with matrilocal traditions are expected to exhibited a lower mitochondrial genetic diversity because the movement of these genomes are reduced since woman remain within families after marriage. However, the results presented here did not reflect these expectations indicating that Mouso may have started practicing matrilocality long time ago, at least after the Paleolithic Age. In contrast to previous studies that showed a clear relationship between Mouso and Naxi people based on just mtDNA haplogroup frequencies, the network analyses presented here indicated clear clusters of individual sequences between Mouso and Pumi lineages. The genetic resemblance between these two group are concordant with other evidences from cultural and language studies. These results indicate that simply comparing haplogroups frequencies among ethnic groups may lead to erroneous conclusions and analyses comparing mtDNA sequences are better suitable for exploring genetic relationship among ethnic groups.
30. Haber M, Platt DE, Ashrafian Bonab M, Youhanna SC, Soria-Hernanz DF, Martínez-Cruz B, Douaihy B, Ghassibe-Sabbagh M, Rafatpanah H, Ghanbari M, Whale J, Balanovsky O, Wells RS, Comas D, Tyler-Smith C, Zalloua PA; The Genographic Consortium. 2012. Afghanistan’s Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events. PLoS ONE 7(3): e34288. doi:10.1371/journal.pone.0034288
- This study focus on how Afghanistan’s ethnic groups relate to each others and with other populations from neighboring countries. The results presented indicated that major genetic differences among Afghanistan’s ethnic groups are relatively recent. The different modern ethnic groups share a genetic heritage probably formed during the Neolithic in the founding of the early farming communities. However, differentiation among the ethnic groups likely started during the Bronze Age driven by the establishment of the first civilizations. Later migrations and invasions to the region, gave the Afghans a unique genetic diversity in Central Asia.
31. Schurr, T. G., Dulik, M. C., Owings, A. C., Zhadanov, S. I., Gaieski, J. B., Vilar, M. G., Ramos, J., Moss, M. B., Natkong, F. and The Genographic Consortium. 2012. Clan, language, and migration history has shaped genetic diversity in Haida and Tlingit populations from Southeast Alaska. American Journal of Physical Anthropology. (published online May 1, 2012) doi: 10.1002/ajpa.22068.
- This manuscript gives new insights about the genetics of the linguistically distinctive Haida and Tlingit tribes of Southeast Alaska. More espcifically, this paper study the role that Southeast Alaska may have played in the early colonization of the Americas; the genetic relationships of Haida and Tlingit to other indigenous groups in Alaska and Canada; the relationship between linguistic and genetic data for populations assigned to the Na-Dene linguistic family; the possible influence of matrilineal clan structure on patterns of genetic variation in Haida and Tlingit populations; and the impact of European entry into the region on the genetic diversity of these indigenous communities. The analysis indicates that, while sharing a ‘northern’ genetic profile, the Haida and the Tlingit are genetically distinctive from each other. In addition, Tlingit groups themselves differ across their geographic range, in part due to interactions of Tlingit tribes with Athapaskan and Eyak groups to the north. The data also reveal a strong influence of maternal clan identity on mtDNA variation in these groups, as well as the significant influence of non-native males on Y-chromosome diversity. These results yield new details about the histories of the Haida and Tlingit tribes in this region.
32. Dulik, M. C., Owings, A. C., Zhadanov, S. I., Gaieski, J. B., Vilar, M. G., Schurr, T. G., and The Genographic Consortium. 2012. Y-chromosome analysis of native North Americans reveals new paternal lineages and genetic differentiation between Eskimo-Aleut and Dene speaking populations. Accepted for publication in April in PNAS.
- The genetic origins of the linguistically diverse Native Americans and when they reached the Americas are questions that have been explored during the last several decades. This study provides new information to these questions by increasing the number of populations sampled and the genetic resolution used in the analyses Here, it is tested whether there is any correlation between genetic diversity from paternally inherited Y-chromosomes and native populations speaking the two distinctive linguistic families: Eskimo-Aleut and Na-Dene. The results indicate that the Y chromosome genetic diversity among the first Native American was greater than previously shown in other publications. In addition, the Eskimo-Aleut and Na-Dene speaking populations showed clear genetic differences between then. The disparities in language, culture and genetic diversity between these two populations likely reflect the outcome of two migrations that happened after the initial settlement of people into the Americas.
33. Martinez-Cruz B, Ioana M, Calafell F, Arauna LR, Sanz P, Ionescu R, Boengiu S, Kalaydjieva L, Pamjav H, Makukh H, Plantiga T, van der Meer JWM, Comas D, Netea M, The Genographic Consortium. 2012. Y-chromosome analysis in individuals bearing the Basarab name of the first dynasty of Wallachian kings. PLoS ONE 7(7): e41803
- The most famous Transylvanian prince is Vlad III from the Basarab royal dynasty, also commonly known as Dracula. The ethnic origins of the Basarab is intensively debated among historians and it is unclear of whether they are descendants of the Cuman people (an admixed Turkic people that reached Romania from the East in the 11th century) or of Vlach people (local Romanians). This paper investigated the Y chromosome of 29 Romanian men carrying the surname Basarab and in order to identify their genetic origin the data was compared with four Romanian and other surrounding populations. Different Y-chromosome haplogroups were found within the individuals bearing the Basarab name, indicating that not all these individuals can be direct biological descendants of the Basarab dynasty. In addition, all these haplogroups are common in Romania and other Central and Eastern European populations. The Basarab group exhibited closer genetic distances with other Romanian populations. These results together with the absence of Eastern Asian paternal lineages in the Basarab men can be interpreted as a lack of evidence for a Cuman origin of this royal dynasty, although it cannot be positively ruled out. As a final conclusion, it seems that the Basarab dynasty was successful in spreading its name beyond the spread of its genes.
34. Rebala K, Martínez-Cruz B, Tönjes A, Kovacs P, Stumvoll M, Lindner I, Büttner A, Wichmann H-E, Siváková D, Soták M, Quintana-Murci L, Szczerkowska Z, Comas D, The Genographic Consortium. 2012. Contemporary paternal genetic landscape of Polish and German populations: from early medieval Slavic expansion to post-World War II resettlements. European Journal of Human Genetics 21(4): 415-422
- One of the most outstanding phenomena in the Y-chromosomal diversity in Europe concerns the sharp genetic border identified between the ethnically /linguistically defined Slavic (from Poland) and German populations (from Germany). The Polish paternal lineages also reveal great degree of homogeneity in spite of a relatively large geographic area seized by the Polish state. Two main explanations have been proposed to explain the phenomena: (i) Massive human resettlements during and shortly after the World War II, and (ii) an early medieval Slavic migrations that displayed previous genetic heterogeneity. In order to answer these questions, 1,156 individuals from several Slavic and German populations were analyzed, including Polish pre-war regional populations and an autochthonous Slavic population from Germany. This study demonstrates for the first time that the Polish paternal lineages were unevenly distributed within the country before the forced resettlements of millions of people during and shortly after the WWII. Finally, the coalescence analyses support hypothesis that the early medieval Slavic expansion in Europe was a demographic event rather than solely a linguistic spread of the Slavic language.
35. Arunkumar G, Soria-Hernanz DF, Kavitha VJ, Arun VS, Syama A, Ashokan KS, Gandhirajan KT, Vijayakumar K, Narayanan M, Jayalakshmi M, Ziegle JS, Royyuru AK, Parida L, Wells RS, Renfrew C, Schurr TG, Smith CT, Platt DE, Pitchappan R; Genographic Consortium. 2012. Population differentiation of southern Indian male lineages correlates with agricultural expansions predating the caste system. PLoS ONE. 7(11): e50269
- Previous studies that pooled Indian populations from a wide variety of geographical locations, have obtained contradictory conclusions about the processes of the establishment of the Varna caste system. This study investigates the origin of the caste system by genotyping 1,680 Y chromosomes representing 12 tribal and 19 non-tribal (caste) populations from the Dravidian-speaking Tamil Nadu state in the southernmost part of India. 81% of Y chromosome were autochthonous Indian haplogroups (H-M69, F-M89, R1a1-M17, L1-M27, R2-M124, and C5-M356; 81% combined) with a shared genetic heritage dating back to the late Pleistocene (10-30 Kya). Results show a strong evidence for genetic structure, and coalescent analyses suggest that the stratification was established 4-6 thousand years ago, with little admixture took place during the last several millennia. The overall Y-chromosomal patterns, the time depth of population diversifications and the period of differentiation are best explained by the emergence of agricultural technology in South Asia. These results highlight the utility of detailed local genetic studies within India, without prior assumptions about the importance of Varna rank status for population grouping, to obtain new insights into the relative influences of past demographic events for the population structure of the whole of modern India.
2013
36. Badro DA, Douaihy B, Haber M, Youhanna SC, Salloum A, Ghassibe-Sabbagh M, Johnsrud B, Khazen G, Matisoo-Smith E, Soria-Hernanz DF, Wells RS, Tyler-Smith C, Platt DE, Zalloua PA, The Genographic Consortium. 2013. Y-chromosome and mtDNA genetics reveal significant contrasts in affinities of Modern Middle Eastern populations with European and African populations. PLoS ONE 8(1):e54616
- The Middle East was a funnel of human expansion out of Africa, a staging area for the Neolithic Agricultural Revolution, and the home to some of the earliest world empires. In addition, post LGM expansions into the region and subsequent population movements have created a striking genetic mosaic in the region. In this study 5,174 mtDNA and 4,658 Y-chromosome samples were investigated. Lebanon’s mtDNA showed a very strong association to Europe, while Yemen shows very strong affinity with Egypt and North and East Africa. Previous Y-chromosome results showed a Levantine coastal-inland contrast marked by Y-haplogroups J1 and J2, and a very strong North African component was evident throughout the Middle East. Neither of these patterns were observed in the mtDNA. While J2 has penetrated into Europe, the pattern of Y-chromosome diversity in Lebanon does not show the widespread affinities with Europe, as indicated by the mtDNA data. Lastly, while each population shows evidence of historic expansions that now define the Middle East, Africa, and Europe, most Middle Eastern populations show distinctive mtDNA and Y-haplogroup characteristics that suggest long standing settlements with relatively little impact from other populations.
37. Der Sarkissian C, Balanovsky O, Brandt G, Khartanovich V, Buzhilova A, Koshel S, Zaporozhchenko V, Gronenborn D, Moiseyev V, Kolpakov E, Shumkin V, Alt KW, Balanovska E, Cooper A, Haak W, The Genographic Consortium. 2013. Ancient DNA reveals prehistoric gene-flow from Siberia in the complex human population history of North East Europe. PLoS Genetics 9(2): e1003296
- Archaeological, anthropological, and genetic research of Northeastern European populations have revealed a series of influences from Western and Eastern Eurasia. While genetic data from modern-day populations is commonly used to make inferences about origins and past migrations, ancient DNA provides a powerful tool by giving a snapshot of the past genetic diversity. This study generated and analyzed 34 mitochondrial genotypes from the skeletal remains of three Mesolithic and the Early Metal Age (7,500 and 3,500 years ago) sites in northwest Russia. Comparisons of genetic data from ancient and modern-day populations revealed significant changes in the makeup of North East Europeans through time. Mesolithic foragers showed high frequencies and diversity of haplogroup U (U2e, U4, U5a), commonly observed in hunter-gatherers from Iberia to Scandinavia. In contrast, the presence of mitochondrial DNA haplogroups C, D, and Z in Early Metal Age individuals suggested genetic influx from central/eastern Siberia. This genetic dissimilarities between prehistoric and modern-day North East Europeans/Saami suggests a strong influence of post-Mesolithic migrations from Western Europe and subsequent population replacement/extinctions. This work demonstrated how ancient DNA can improve our understanding of human population movements across Eurasia.
38. Brotherton P, Haak W, Templeton J, Brandt G, Soubrier J, Jane Adler C, Richards SM, Sarkissian CD, Ganslmeier R, Friederich S, Dresely V, van Oven M, Kenyon R, Van der Hoek MB, Korlach J, Luong K, Ho SY, Quintana-Murci L, Behar DM, Meller H, Alt KW, Cooper A, The Genographic Consortium. 2013. Neolithic mitochondrial haplogroup H genomes and the genetic origins of Europeans. Nature Communications 4:1764
- Haplogroup H dominates present-day Western European mitochondrial DNA variability (>40%), yet was less common (~19%) among Early Neolithic farmers (~5450 BC) and virtually absent in Mesolithic hunter-gatherers. This project investigated maternal population history of modern Europeans by sequencing 39 complete haplogroup H mitochondrial genomes from ancient remains; and comparing this ‘real-time’ genetic data with cultural changes taking place between the Early Neolithic (~5450 BC) and Bronze Age (~2200 BC) in Central Europe. Results revealed that the current diversity and distribution of haplogroup H were largely established by the Mid Neolithic (~4000 BC), but with substantial genetic contributions from later pan-European cultures such as the Bell Beakers expanding out of Iberia in the Late Neolithic (~2800 BC). Newly dated haplogroup H genomes enabled the reconstruction of the evolutionary history of the haplogroup, and revealed a mutation rate 45% higher than previous estimates.
39. Elhaik E, Greenspan E, Staats S, Krahn T, Tyler-Smith C, Xue Y, Tofanelli S, Francalacci P, Cucca F, Pagani L, Jin L, Li H, Schurr TG, Greenspan B, Spencer Wells R, The Genographic Consortium. 2013. The GenoChip: a new tool for genetic anthropology. Genome Biology & Evolution 5(5): 1021-1031
- The Genographic Project is an international effort aimed at charting human migratory history. The first phase of the project was focused on haploid DNA markers (Y-chromosome and mtDNA), while the current phase focuses on markers from across the entire genome using the newly created GenoChip. GenoChip was designed to enable higher resolution research into outstanding questions in genetic anthropology. It includes ancestry informative markers obtained for over 450 human populations, an ancient human (Saqqaq), and two archaic hominins (Neanderthal and Denisovan) and it was designed to identify all known Y-chromosome and mtDNA haplogroups. The chip was also carefully vetted to avoid inclusion of medically relevant markers. To demonstrate its capabilities, we compared the FST distributions of GenoChip SNPs to those of two commercial arrays. Although all arrays yielded similarly shaped FST distributions, the GenoChip autosomal and X-chromosomal distributions had the highest mean FST, attesting to its ability to discern subpopulations. In summary, the GenoChip is a dedicated genotyping platform for genetic anthropology. With an unprecedented number of approximately 12,000 Y-chromosomal and approximately 3,300 mtDNA SNPs and over 130,000 autosomal and X-chromosomal SNPs with no health, medical, or phenotypic relevance, the GenoChip is a useful tool for genetic anthropology and human population genetics.
40. Boattini A, Martinez-Cruz B, Sarno S, Harmant C, Useli A, Sanz P, Yang-Yao D, Manry J, Ciani G, Luiselli D, Quintana-Murci L, Comas D, Pettener D; The Genographic Consortium. 2013. Uniparental markers in Italy reveal a sex-biased genetic structure and different historical strata. PLoS ONE 8(5): e65441
- Italy played an important role in the history of human settlements and movements of Southern Europe and the Mediterranean. Populated since Paleolithic times, the complexity of human movements during the Neolithic, the Metal Ages and the most recent history of the two last millennia, shaped the pattern of the modern Italian genetic structure. With the aim of disentangling this pattern, this project analyzed the haploid markers in ∼900 individuals from across the Italian peninsula, Sardinia and Sicily. Results show a sex-biased pattern, indicating different demographic histories for males and females. Besides the genetic outlier position of Sardinians, a North West-South East Y-chromosome structure appeared through continental Italy, likely a result of historical and demographic events. In contrast, mitochondrial (maternal) diversity is distributed homogeneously in accordance with older pre-historic events, as was the presence of an Italian Refugium during the last glacial period in Europe.
41. Sandoval JR, Lacerda DR, Jota MS, Salazar-Granara A, Vieira PP, Acosta O, Cuellar C, Revollo S, Fujita R, Santos FR, The Genographic Consortium. 2013. The genetic history of indigenous populations of the Peruvian and Bolivian Altiplano: the legacy of the Uros. PLoS ONE 8(9): e73006
- Since pre-Columbian times, different cultures established themselves around the Titicaca and Poopo Lakes. Yet by the time of Spanish colonization, the Inca Empire and the Aymara and Quechua languages were dominant in the region. This study focused on the pre-Columbian history of the Altiplano populations, particularly the Uros, which claim to be directly descend from the first settlers of the Andes. Results indicate that the Uros populations stand out among others in the Altiplano, while appearing more closely related to the Aymara and Quechua from Lake Titicaca and surrounding regions, than to the Amazon Arawaks. Moreover, the Uros populations from Peru and Bolivia are genetically differentiated from each other, indicating a high heterogeneity in this ethnic group. Lastly, the results support the distinctive ancestry for the Uros populations of Peru and Bolivia, likely derived from ancient Andean lineages, but further complicated by a partial replacement during more recent farming expansion, and the establishment of complex civilizations in the Andes, such as the Inca.
42. Brandt G, Haak W, Adler CJ, Roth C, Szécsényi-Nagy A, Karimnia S, Möller-Rieker S, Meller H, Ganslmeier R, Friederich S, Dresley V, Nicklish N, Pickrell JK, Siroko F, Reich D, Cooper A, Alt KW, The Genographic Consortium 2013. Ancient DNA Reveals Key Stages in the Formation of Central European Mitochondrial Genetic Diversity. Science 342, no.6155: 257-261.
- Genographic project scientists, in collaboration with archeologists from Germany, successfully sequenced and analyzed DNA from 364 individuals that lived in Central Europe between 5,500 and 1,500 BC. What they found was that the shift in the frequency of DNA lineages closely matched the changes and appearances of new Central European cultures across time. In other words, the people who lived in Central Europe 7,000 years ago had different DNA lineages than those that lived there 5,000 years ago, and again different to those that lived 3,500 years ago. Central Europe was dynamic place during the Bronze age, and the genetic composition of the people that lived there demonstrates that. Ultimately, Central Europe is a melting pot of genetic lineages from different prehistoric cultures that lived there at different periods of time, each new one partially replacing the one before it.
_____________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Services
Genealogy Research
Like this:
Like Loading...