Have you ever dealt with something so massive and overwhelming it took a few days just to get your head wrapped around it? Well, that’s how I’ve been feeling about the new National Geographic Geno 2.0 announcement. It’s not just what has been announced, but the utterly massive amount of scientific research behind the scenes, and what it means to the rest of us.
If you think of all of the discoveries and progress that has been made in the 12 years since the advent of genetic genealogy, what you’re about to hear today dwarfs it all. Hold on tight – this is a white knuckle ride of a lifetime. The day I heard about this, I wandered around somewhat starry-eyed in amazement and kept muttering something terribly intelligent like “Wow, oh Wow.”
I’d like to share with you some of today’s big news and hope that you too share my sense of awe to be alive in such an exciting time, and to have not only a front row seat, but participating in making history. This isn’t a movie, it’s the real McCoy!
Let’s start with a bit of history about Nat Geo 1.0, the Genographic Project. Fasten your seatbelt, your E ticket ride starts here and now!
Nat Geo 1.0
Eight years ago, in April 2005, the National Geographic Genographic project was announced. The goal was to sell a total of 100,000 kits over 5 years to help fund the indigenous part of the project, which was to collect samples from indigenous peoples around the world to better understand population migration.
According to Nat Geo, this has been the most successful program they have ever undertaken. That in and of itself it an amazing statement, especially considering that there was a lively debate within Nat Geo prior to the project launch.
Someone opined to Spencer Wells that they wouldn’t even sell 10,000 kits, let alone 100,000. Well, they were wrong, 10,000 kits were sold the first day alone. I’m guessing that Bennett and Max at Family Tree DNA, whose test kits Nat Geo uses, has a sense of controlled panic about that time. The 100,000 kits were sold in the first 8 months and they still sell between 40,000 and 50,000 kits per year today.
How is that project doing? Well, it was scheduled to run for 5 years, and it’s now into its 7th year. They have collected over 75,000 samples from indigenous people and on the public side, over 750,000 people in over 130 countries have bought kits to help fund the research. 32 publications either have been released or will be shortly. Of the 45 million dollars the project has grossed, National Geographic has contributed more than 1.7 million dollars to the Legacy Fund for investment back into the indigenous communities that participated in the Genographic project.
You might recall that the original Nat Geo project only tested 12 markers for men and the HVR1 region on the maternal side. At that time, 7 years ago, $99 for each of those was a great deal and the projects received a lot of new participants. About 20% of the Nat Geo participants transferred their result to Family Tree DNA, for free, so they could join projects and participate in genetic genealogy.
Today, 12 markers is quite light and so is HVR1 testing alone. Project administrators cringe when we see those, because we know it’s really not enough to do much with today. We’ve learned so much in the past 7 years. You don’t realize how much things have changed until you take a minute to look back.
At the same time we were learning, technology was also advancing. Seven years ago, running autosomal tests was simply cost prohibitive. If you consider that computer technology has decreased in price and doubled in speed every year or two (Moore’s Law), the advances in DNA sequencing technology and understanding are moving in the same directions (increased capability and decreased costs) by a factor of 5 as compared to computer technology. Literally, we are moving at the speed of light. See, I told you to hold on. I meant it!
Geno 2.0 – The Big Announcement
It’s amazing that something this big has been kept this quiet. Those of us involved have been bursting at the seams with excitement, and today is the big day. Last night about 9 o’clock we received word that the countdown had begun.
For a look at the new National Geographic webpage, go to www.genographic.com. This is the heart of the new Geno 2.0.
Geno 2.0 is still comprised of the 3 core components as before, the indigenous portion, the Legacy fund and the public participation portion. However the technology is changing, dramatically, and the public participation arena is expanding. Public participation will now include some “citizen science” projects, grants, an educational segment meaning kits in classrooms, and community based projects. All of this is made possible by advances in the core sciences and technology. This, plus the focus of the “Dream Team” of genetic genealogy and population genetics.
Thankfully, Spencer Wells at National Geographic and Bennett Greenspan and Max Blankfeld at Family Tree DNA prepared us in advance for what was coming, as much as you can prepare for a technological tsunami!
Let’s take a look at the technology and scientific advances that have occurred and what it means to us today.
New Chips and New Partnerships
The days of sequencing 12 markers in the lab are gone forever, replaced by high-speed sequencing that looks at half a million markers, or more, at a time, and for the same price as a 12 marker test and the mitochondrial DNA test, together, would have cost in Nat Geo 1.0.
However, when you’re looking at just the Y DNA and the mitochondrial, you’re missing 98% of the human genome, the part that isn’t Y or mitochondrial DNA. And that 98% holds many secrets, the secrets of our ancestors.
The National Geographic Society recruited one of the top geneticists in the world at Johns Hopkins, focused on autosomal genetic markers. He has spent the past two years identifying every known marker relevant to ancestry or population genetics that is NOT medically relevant. This includes the X and Y chromosomes, mitochondrial DNA and the balance of the autosomal markers.
Are you sitting down? Here’s the first of several bombs!
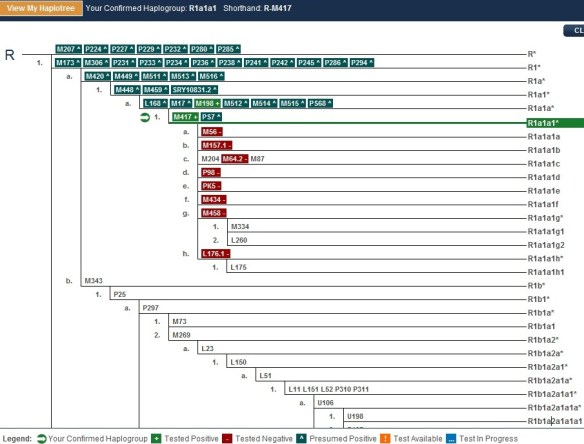
Relative to Y-line DNA, in 2010, just 2 years ago, the YCC SNP 2010 tree had a total of just over 800 SNPs that has been discovered. Today it still hasn’t reached 900. You can see the current tree at http://www.isogg.org/tree/index.html. Notice that all of the L SNPs were discovered by Thomas Krahn in the Family Tree DNA lab with the assistance of Family Tree DNA’s customers and project administrators. This is truly “crowd-science” in the flash mob sense.
Today, after a concerted effort of discovery involving many people, there are a total of 12,000 Y SNPS and of that, 10,000 of them are unique and new and have never been seen or published before. This means that your haplogroup will automatically be determined to the furthest branch of the tree with no additional SNPs to be tested. As this test becomes available to Family Tree DNA clients as an upgrade, it will signal the demise of the deep clade test.
If there is a project administrator sitting next to you, they have just fainted. The magnitude of this is simply mind-boggling.
Relative to mitochondrial DNA, 3352 unique (non-haplogroup defining) mutations have been discovered. To measure all of the relevant mitochondrial DNA mutations, including insertions and deletions, over 31,000 probes (locations) are needed on the new high density chips. Before this new approach, chip technology was unable to account for insertions and deletions, but that has been remedied by a new approach to an old problem. This means that haplogroups will be determined to their deepest level and they will be accurate, including insertions and deletions critical to haplogroup assignment.
Relative to autosomal DNA, over 75,000 Ancestrally Informative Markers (AIMs) have been discovered and included on the new chip, and that’s after removing any that might be considered medically informative. This astronomical number of SNPs will allow us to detect ethnicity and improve accuracy on a scale that we’ve never even dreamed about before. I specifically asked Spencer Wells if this will help resolve those “messy” situations where we have European, Native American and African admixture, and he indicated that it would. I can hardly wait. For those of us what have been waiting patiently, and some not so patiently, to be able to identify small amounts of admixture, this is the best news you could ever hope to hear! I told you that something wonderful was on the way!
Relative to admixture with Neanderthal, Denisovan and Melanesian man, meaning interbreeding, more than 30,000 SNPs have been identified that will signal interbreeding where it occurred between modern humans and ancient hominids. And yes, this means that it did occur! So indeed once again, you can begin wondering about your brother-in-law. He’s probably wondering about you too.
Relative to the X chromosome, it’s included. The X chromosome, because of its special inheritance pattern, gives us an additional, special tool when working with genetic genealogy. We’ll cover this in a future blog.

The New Chip
In total, the new SNP count to be included on the new Nat Geo 2.0 chip (photo above) includes both new and known existing SNPs in the following amounts:
- Autosomal including X – 147,000
- Neanderthal – 26,000
- Denisovan – 1,500
- Aboriginal – 13,000
- Eskimo – 12,000
- Chimpanzee – 1,100
- Y Chromosome – 12,000
- mtDNA – 31,000
This chip has been designed to distinguish between populations.
OMG – What Happened to the Haplotree?
We’re not done yet with bombshells.
After this new chip was created by Illumina specifically for National Geographic, about 1200 samples were run as proof of concept, including 400 WTY (Walk the Y), 350 mitochondrial full sequence and 500 Y samples. All of the samples run are checked and tested for all of the SNPs on the chip. Of course, females’ samples will fail on all of the Y haplogroup locations, etc.
Just based on this test run alone of 900 Y chromosome kits, the haplotree expanded from 862 SNPs to a total of 6153. If you’ve just said something akin to “Holy Cow,” you’re on the right track. Imagine what it will do with another 1000 or 10,000 or 100,000 tests. Right now, we’re making discoveries so fast we can hardly deal with them.
What Does This Mean?
In reality, what this means is that we will very soon use SNPs to determine heritage down to a genealogical meaningful timeframe, meaning 500 to maybe 1000 years. The standard STR (Short Tandem Repeat) markers we know and love will become the leaves on the branches of the tree and these will likely be used when there are no more SNPs to determine family groupings and line marker mutations within families.
New National Geographic Geno 2.0 Website
Needless to say, all of this discovery has prompted National Geographic to redo their website entirely. New maps are forthcoming. Yeah!! New maps include the migration maps as well as new haplogroup “heat maps” where the colors are graduated based on frequency.

There are entirely new capabilities too. The new website will show you as the center of a circle and you’ll be able to contact people who have tested at Nat Geo who are located near to you in the circle. Those closest to you, you’re most closely related to. Further away, more distantly related. Before, there was no matching between Nat Geo participants.
And yes, Geno 2.0 participants will still be able to transfer into Family Tree DNA for free. I hope they make that option much more visible or interactive.
A New Test Kit
Anyone wanting to participate in Geno 2.0 will have to order a new kit from National Geographic. The previous Nat Geo kits, if you recall, were anonymous unless you chose to transfer to Family Tree DNA, plus the permission you gave was specifically for mtdna or Y-line, not autosomal testing.
Furthermore, the DNA in many kits will be too old and will have degraded too much to use. Everyone ordering the new Geno 2.0 kit will receive a new swab kit, in an heirloom box. The comprehensive Y-line (haplogroup only), mtdna (haplogroup only) and autosomal testing will cost $199.

For Family Tree DNA clients who will be offered the upgrade in the late summer or fall, you will be able to upgrade if your DNA is less than 4 or 5 years old. Otherwise, you’ll receive a new swab kit too.
All processing will be done at the Family Tree DNA Houston facility.
New Results Pages
The new test of course requires all new results pages for participants.

Take a look at a few of the pages you can expect.

The results will be presented as a personal story.

Your story will also include information such as maps of where your ancestors lived and where they migrated.

I asked Spencer if participants will be able to download their results so that we can continue to compare them as we do today, using various phasing tools. Spencer replied, “Yes, raw results WILL be available for download. In the Genographic Project, you will always own your DNA results, and the genotype data will be yours to do with as you please. I feel very strongly that this is a cornerstone of ethical DTC genetic testing.” Way to go Spencer!!
As Geno 2.0 moves forward, additional analytical tools will be added.
Ordering
National Geographic is accepting pre-orders now. They will ship before the end of October, and they expect to be shipping significantly before that.
In Summary
Our world is changing, rapidly, and for the better. The door we’ve been peeking through for a decade now is swinging wide open. More brick walls will fall. We’ll find and meet new cousins. Ethnicities will be identified at a level never before possible. We’ll learn about our ancestors and the story of our past through their DNA that we carry today. It is the frontier within. DNA is truly the gift that keeps on giving!
“One small step for man, one giant leap for mankind.”
Neil Armstrong, July 20, 1969
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Services
Genealogy Research