“Charlemagne Agostino Cornacchini Vatican 2” by Photo: Myrabella / Wikimedia Commons. Licensed under CC BY-SA 3.0 via Commons
Above, a 1725 statue representing Charlemagne housed in St. Peter’s Basilica in Rome.
Wow, do I ever have a lot of cousins. According to Graham Coop, everyone in Europe today is descended from Charlemagne. Which either means I’m special and so is everyone else, or we’re all just normal. National Geographic wrote an article about the results of the Coop study in more easily readable verbiage, here. The Coop team also wrote a nontechnical FAQ, here.
In 2002, Steven Olson wrote this verbiage in the Atlantic magazine about the work of statistician Joseph Chang:
The most recent common ancestor of every European today (except for recent immigrants to the Continent) was someone who lived in Europe in the surprisingly recent past—only about 600 years ago. In other words, all Europeans alive today have among their ancestors the same man or woman who lived around 1400. Before that date, according to Chang’s model, the number of ancestors common to all Europeans today increased, until, about a thousand years ago, a peculiar situation prevailed: 20 percent of the adult Europeans alive in 1000 would turn out to be the ancestors of no one living today (that is, they had no children or all their descendants eventually died childless); each of the remaining 80 percent would turn out to be a direct ancestor of every European living today.
Now, granted, Charlemagne was indeed prolific, siring 18 known children. I guess I’m just lucky in that I descend from two known children, one who was the King of Italy and one who was the King of France. I must tell you, all this king stuff sounds very surreal to me.
Now, the bad news. In spite of all of these children, it appears that none of Charlemagne’s male lines survived more than a dozen or so generations. In generation 8, only two descendants produced a male child, so that severely limited the possibilities for his Y DNA to reproduce – and it didn’t. It apparently died out in 1122 with the death of the last male in an illegitimate line through Charlemagne’s son, Pepin of Italy. This means that today, we don’t know what Charlemagne’s Y DNA looked like. That’s very disappointing.
By the same token, Charlemagne could not pass on his own mtDNA. To find that information, we’d have to find a female sibling or someone from his mother’s line who descends from a matrilineal female through all females to the current generation. We don’t have that either.
And as for autosomal DNA…well, we have two problems. The first is that Charlemagne is so far back in my or anyone else’s lineage – between 40 and 49 generations roughly – that we would carry very little if any of his DNA today…and there is no way to assure that we don’t have other common lines too. In fact, at that distant point, it far more likely that we do share other common lines that we don’t, especially given what Graham Coop’s paper indicates.
So, Charlemagne’s autosomal DNA hasn’t been identified either. Short of digging him up, I’m doubting we’ll know much more. Actually, poor Charlemagne has been dug up, several times in fact, and parts of him given away as religious relics. In 1988 scientists tried to reassemble him, and their report was delivered 26 years later, without DNA unfortunately, but confirming as best they can that the remains they have, shown in the photo below, are indeed Charlemagne. I must say, it’s a very odd feeling to look at the bones of my ancestor, reassembled during the study. In fact, I don’t think I’ve ever seen the bones or even one bone of any of my ancestors before now. I’m not exactly sure what I think of this.
I’m striking out here genetically, although I’m hopeful for the future since his bones are already exhumed. To begin with, DNA could tell us if all of those bones are really from the same person. If they are, or most of them are from the same person, it’s more likely to be Charlemagne. We could potentially tell when and where this skeletal person lived from isotope testing as well, which could help us confirm or eliminate the possibility that these skeletal remains are Charlemagne. Y and mitochondrial DNA would tell us a lot about his ancestors, and therefore, ours too. I hope this avenue is being pursued.
Let’s see what we can discover about Charlemagne outside of genetic genealogy.
Charlemagne’s Birth and Ascent to Power
Charlemagne was born on April 2, in either 742, 747 or 748 and died on January 28, 814. No, those aren’t typos, they are genuinely three digit years. It’s hard for me to come to grips with the fact that I have ancestors that I can identify that were born 1270 years ago.
Charlemagne’s father was Pepin the Short and his mother, Bertrada of Laon. He was of the Carolingian dynasty and was, of course, Catholic. In fact, Charlemagne was DEVOUTLY Catholic, which plays a big part in the decisions he made in his lifetime. Either that, or he used his religious fervor as an excuse for his invasions of other non-Christian domains. It’s much easier to track the history of what he did rather than to discern his motivations.
The Carolingian dynasty was a Frankish noble family with origins in the Arnulfing and Pippinid clans of the 7th century AD.
This Carolingian family tree, above, dates from the Chronicon Universale of Ekkehard of Aura from the 12th century, but reflects earlier generations. The Carolingian empire came to a close not long after after Charlemagne’s rule, about 888.
The name “Carolingian” was in Medieval Latin, karolingi, an altered form of an unattested Old High German karling or kerling, meaning “descendant of Charles.”
Charlemagne’s birth year remains uncertain. The most likely year of Charlemagne’s birth is reconstructed from several sources. The date of 742, calculated from Einhard’s date of death of January 814 at age 72, predates the marriage of his parents in 744. Einhard was a Frankish scholar and servant of Charlemagne (and his son) who also served as Charlemagne’s biographer – thankfully.
The year given in the Annales Petaviani, a year by year history of the Carolingina empire, 747, would be more likely, except that it contradicts Einhard and a few other sources in making Charlemagne seventy years old at his death. The month and day of April 2 is established by a calendar from Lorsch Abbey.
In 747, that day fell on Easter, a coincidence that likely would have been remarked upon by chroniclers but was not. If Easter was being used as the beginning of the calendar year, then April 2, 747 could have been, by modern reckoning, April 2, 748 (not on Easter). The date favored by the preponderance of evidence is April 2, 742, based on Charlemagne’s being a septuagenarian at the time of his death. This date would appear to suggest that Charlemagne was born illegitimately, which is not mentioned by Einhard.
Einhard said the following:
It would be folly, I think, to write a word concerning Charles’ birth and infancy, or even his boyhood, for nothing has ever been written on the subject, and there is no one alive now who can give information on it. Accordingly, I determined to pass that by as unknown, and to proceed at once to treat of his character, his deeds, and such other facts of his life as are worth telling and setting forth, and shall first give an account of his deeds at home and abroad, then of his character and pursuits, and lastly of his administration and death, omitting nothing worth knowing or necessary to know.
We also don’t know where Charlemagne was born, but Aachen in today’s Germany has been suggested.
Charlemagne became king in 768 following the death of his father. He was initially co-ruler with his brother Carloman I. Carloman’s sudden death in 771 under unexplained circumstances, left Charlemagne as the undisputed ruler of the Frankish Kingdom. Charlemagne continued his father’s policy towards the papacy and became its protector, removing the Lombards from power in northern Italy, and leading an incursion into Muslim Spain. He also campaigned against the Saxons to his east, Christianizing them upon penalty of death, leading to events such as the Massacre of Verden in which 4500 captive Saxons were slaughtered. Charlemagne was not always a kind man – but history remembers him as an exceedingly effective ruler.
Above, Charlemagne on the left and his son, Pepin the Hunchback, who revolted against his father. Pepin the Hunchback was subsequently censured and exiled to a monastery instead of put to death after his father commuted his death sentence.
We don’t know how much this drawing actually looks like Charlemagne since it is a 10th century copy of a lost original from about 830.
In this next drawing, Charlemagne instructs his son, Louis the Pious.
Charlemagne was also known as both Charles the Great (Carolus Magnus) or Charles the First. He became the King of the Franks beginning in 768 with the death of his father and King of Italy beginning in 774. He was crowned Holy Roman Emperor (Imperator Augustus) in the now demolished Old St. Peter’s Basilica in Rome (shown sketched below between 1483-1506) on Christmas Day in the year 800 and he ruled in that position until his death.
This fresco below shows a cutaway view of the Old St. Peter’s from the 4th century, so it looked much like it would have to Charlemagne. This basilica is built over the location believed to be the burial site of St. Peter.
Today, a new St. Peter’s Basilica stands on this site, the dome visible from the Ponte Umberto I on the Tiber River, below.

“Vatican City at Large” by Sébastien Bertrand from Paris, France – Flickr. Licensed under CC BY 2.0 via Commons
Although he already ruled both Italy and France, becoming the Holy Roman Emperor bestowed upon him divine grace and a Godly legitimacy sanctioned by the Pope.
This painting from 1516-1517 by Raphael by depicts Charlemagne’s coronation.
Charlemagne’s Rule
Called the “Father of Europe” (pater Europae), Charlemagne united most of Western Europe for the first time since the Roman Empire and laid the foundations for both modern France and Germany. His rule spurred the Carolingian Renaissance, a period of energetic cultural and intellectual activity within the Church. Both the French and German monarchies considered their kingdoms to be descendants of Charlemagne’s empire.
Charlemagne didn’t seem to stay home much. In fact, this military and political history reads like a soap opera, with intrigue, betrayals, a brother who died in unexplained circumstances, but apparently of natural causes, invasions, rebellions and saving an injured Pope. His life was assuredly interesting and it’s nothing short of amazing that he managed to live past 70 and did not die on the battlefield.
Charlemagne was engaged in almost constant warfare throughout his reign, often at the head of his elite scara bodyguard squadrons, with his legendary sword Joyeuse in hand.
Joyeuse is stunningly beautiful and is on display in the Louvre today.
It just kills me that I have been in this building with this sword but didn’t know at that time that Charlemagne was my ancestor.
The 11th century “Song of Roland” describes the sword:
[Charlemagne] was wearing his fine white coat of mail and his helmet with gold-studded stones; by his side hung Joyeuse, and never was there a sword to match it; its colour changed thirty times a day.
Some seven hundred years later, Bulfinch’s Mythology described Charlemagne using Joyeuse to behead the Saracen commander Corsuble as well as to knight his comrade Ogier the Dane.
The town of Joyeuse, in Ardèche, is supposedly named after the sword. Joyeuse was allegedly lost in a battle and retrieved by one of the knights of Charlemagne; to thank him, Charlemagne granted him an appanage (estate) named Joyeuse.
Today, Joyeuse is used as the French coronation sword.
Charlemagne’s Additions to His Empire
Charlemagne spent his entire life increasing the size and power of his empire, some of which was done under the banner of expanding Christianity to the Muslim world and to the pagan Saxons as well.
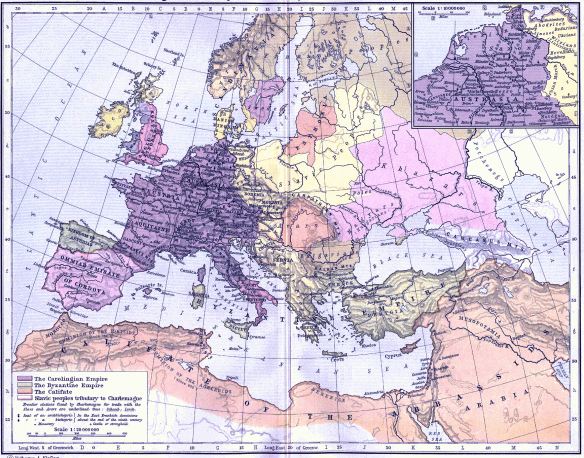
The map below shows the land that Charlemagne added to the Frankish Kingdom.

“Frankish Empire 481 to 814-en” by Sémhur – Own work, from Image:Frankish empire.jpg, itself from File:Growth of Frankish Power, 481-814.jpg, from the Historical Atlas by William R. Shepherd (Shepherd, William. Historical Atlas. New York: Henry Holt and Company, 1911.). Licensed under CC BY-SA 3.0 via Commons
Charlemagne’s reign of Aquitaine began in 768 with the death of his father, although it was initially a joint reign with his brother, Carloman, with whom he had, at best, lukewarm relations fostered by his mother.
It didn’t take long after Charlemagne’s father death for trouble to brew.
In 769, a small uprising in the Basque region was subdued, but that region was unstable for years, a constant thorn in Charlemagne’s side. Finally, in 781, Charlemagne proclaimed his son, Louis the Pious, then a young child, the first Frankish king of the area, assuring loyalty and displacing those whose loyalty he had reason to doubt.
In 770 Charlemagne married the daughter, Desiderada, of a Lombard King as a political move to form an alliance with her father and in doing so, surround brother Carloman with Charlemagne’s allies. However, by the end of 771, Carloman was dead and Charlemagne no longer needed the marriage with Desiderada, so he repudiated her and set her aside to marry 13 year old Hildegard.
In rejecting Desiderada, Charlemagne incurred the wrath of her father, the Italian King Desiderius. Carloman’s widow and children took refuge in King Desiderius’ court at Pavia for protection.
This drawing is of the Carolingian cavalry from that timeframe.

“Karolingische-reiterei-st-gallen-stiftsbibliothek 1-330×400”. Licensed under Public Domain via Commons
Sometimes Charlemagne’s military campaigns overlapped each other. In the Saxon Wars, spanning thirty years and eighteen battles, Charlemagne eventually conquered and subdued Saxonia. The conquering part wasn’t terribly difficult, most of the time, but the subdueing part proved nearly impossible. Charlemagne proceeded to convert the conquered to Christianity, beginning in 773 with his campaign against the Engrians where he cut down the Saxon pagan pillar of Irminsul. However, trouble in Italy caused that campaign to be cut short.
In 773, Charlemagne and his uncle crossed the Alps, chasing the Lombards back to Pavia which he then besieged.
Charlemagne met the Lombards at the pass in Susa Valley, shown below.
By 774, the siege of Pavia was over and Charlemagne had himself declared King of the Franks and Lombards and crowned with the traditional “Iron Crown of the Lombards.”

“Iron Crown” by James Steakley – photographed in the Theodelinda Chapel of the cathedral of Monza. Licensed under CC BY-SA 3.0 via Commons
This crown is called “The Iron Crown” because of the narrow band of iron within the crown said to have been beaten out of the nail used at the Crucifixion of Jesus Christ.
In 776, Charlemagne was back in Saxony, having subdued the Saxons and causing their leader, Widukind, to seek refuge in Denmark. Many Saxons were baptized as Christians.
Although the Lombards surrendered, all was not quiet on that front. In 776, Charlemagne rushed back from Saxony to squelch a rebellion in Lombardy.
In 778, Charlemagne turned back southwards and tried to overpower the Muslim Saracen rulers of Barcelona and nearby areas. He marched to face them, meeting them at Saragossa. He received homage from them, but their cities did not fall.
Charlemagne was facing the toughest battle of his career where the Muslims had the upper hand and forced him to retreat. He decided to go home, since he could not trust the Basques, whom he had subdued by conquering Pamplona. He turned to leave Iberia, but as he was passing through the Pass of Roncesvalles (shown below) one of the most famous events of his long reign occurred.
The Basques fell on his rearguard and baggage train, utterly destroying it. The Battle of Roncevaux Pass, though less a battle than a mere skirmish, left many famous dead, including the seneschal Eggihard, the count of the palace Anselm, and the warden of the Breton March, Roland, inspiring the subsequent creation of the Song of Roland (La Chanson de Roland).
This tapestry portraying the Battle of Roncevaux Pass was woven between 1475 and 1500.
In 779, while Charlemagne was focused elsewhere, Saxony again revolted and he again invaded and reconquered. I’m sure by now he wondered how many times he had to do this. He divided the land into missionary districts and personally assisted with the baptisms of masses at Lippe.
From 780-782, Saxony was quiet. Charlemagne was back in Italy during this time.
In 782, Charlemagne returned to Saxony and was not pleased that the majority of the population was still pagan. He implemented draconian laws prescribing death to Saxon pagans who refused to convert to Christianity. This spurred the return of their leader, Widukind, and was followed by three years of bloody battles precipitated by the Massacre of Verden wherein Charlemagne executed 4500 trapped Saxon soldiers. Three long years later, Widukind, defeated, accepted baptism. At this time, the Frisians were also brought to heel.
In 787, Charlemagne focused on bringing southern Italy into the fold. He besieged Salerno where Arechis who reigned independently submitted to vassalage. However, upon his death in 792, Arechis’ son proclaimed independence and Charlemagne never personally returned, and was never able to bring this region fully under his control.
In 788, Charlemagne was back in Gascony trying to once again reign in a rebellion. He had appointed his son “King” in that region and replaced Gascon individuals in power with his Frankish officers.
In 788, the pagan Asian Avars (Einhard called them Huns) had settled in Hungary and invaded Fruili and Bavaria. Charlemagne was busy elsewhere until 790, but at the time he marched down the Danube and ravaged Avar territory. A Lombard army did the same to Pannonia.
In 789, Charlemagne seeing an opportunity, marched into the Slavic Obotrite territory north of the Elbe, encountered little resistance, and subdued the Obotrites, sending in missionaries to convert them. They became loyal allies, fighting alongside Charlemagne in 795 when the Saxons broke the peace.
In 792, the Saxons rebelled again in Westphalia, breaking several years of peace and distracted Charlemagne from the Avars, occupying him instead with those relentless Saxons.
However, Charlemagne’s troops continued to assault the Avars’ ring-shaped strongholds. The great Ring of the Avars, their capital fortress, was taken twice. The booty was sent to Charlemagne at his capital, Aachen, and redistributed to all his followers and even to foreign rulers, including King Offa of Mercia. Soon the Avars had lost the will to fight and traveled to Aachen to subject themselves to Charlemagne as vassals and Christians. Charlemagne accepted their surrender and sent one native chief, baptized as Abraham, back to Avaria with the ancient title of khagan, meaning someone who rules an empire. Abraham kept his people in line, but in 800, the Bulgarians under Khan Krum also attacked the remains of Avar state.
From time to time, rebellions would flare up in the conquered Saxon territory. In 793 a rebellion erupted in Eastplania and Bordalbingia, but that uprising was over by 794. In 796, an Engrian rebellion followed.
In 794, Charlemagne set his eye upon Bavaria and was shortly thereafter dividing the land into Frankish counties, as he had done with Saxony.
From 791 to 806, Charlemagne was focused on taking the County of Toulouse for a power base and asserting his authority over the Pyrenees, making those counties vassals.
In 797, Barcelona, the greatest city of the region, previously held by the Moors, fell to Charlemagne, but was retaken in 799. However, Louis of Aquitaine marched the entire army of his kingdom over the Pyrenees and besieged it for two years, wintering there from 800 to 801, when it capitulated. Today, the Moorish influence can be seen in Barcelona in the architecture of the buildings.
The Medieval influence can be felt in any portion of the old city and in the plazas.
I absolutely loved Barcelona when I visited in 2011, having no idea that I had ancestral history here.
Charlemagne viewed his battle with the Muslim Moors as the battle of and for Christianity. He wanted to convert the Muslims to Christianity.
In 799, Charlemagne conquered the Balaeric Islands, often attacked by Saracen (Moorish and Muladi) pirates.
In either 797 or 801, Charlemagne send a delegation to Baghdad where the caliph of Baghdad presented Charlemagne with an Asian elephant and a clock. The elephant, Abul-Abbas, was transported to Charlemagne’s headquarters in Aachen where his life was chronicled. He died in 810, possibly a war elephant, although others report that the elephant was more than 40 years old, had rheumatism, developed pneumonia while on campaign with Charlemagne, and died suddenly. I don’t think any of my other ancestors had a pet elephant.
In 799, Pope Leo III had been mistreated by the Romans, who tried to put out his eyes and tear out his tongue. Pope Leo escaped and fled to Charlemagne at Paderborn, asking Charlemagne to intervene in Rome and restore him. Charlemagne, advised by scholar Alcuin of York, agreed to travel to Rome, doing so in November 800 and holding a council on December 1. On December 23, Leo swore an oath of innocence relative to the charges brought against him, and his accusers were exiled. Two days later, at Mass, on Christmas Day, December 25th, when Charlemagne knelt at the altar to pray, the Pope crowned him Imperator Romanorum, “Emperor of the Romans,” in Saint Peter’s Basilica. It is unclear whether Charlemagne knew this was going to happen or if the coronation was unexpected, a point debated by historians for hundreds of years.

Miniature of Charlemagne crowned emperor by Pope Leo III, from Chroniques de France ou de Saint Denis, vol. 1; France, second quarter of 14th century.
In 803, Charlemagne sent a Bavarian army into Pannonia ending the Avar confederation.
In 804, one last rebellion occurred in Saxony, but by this time, 30 years after being conquered, most of the original inhabitants were dead and the rest had never known anything but Charlemagne’s rule and warfare. They were tired of fighting in their homeland.
Einhard tells us:
The war that had lasted so many years was at length ended by their acceding to the terms offered by the King; which were renunciation of their national religious customs and the worship of devils, acceptance of the sacraments of the Christian faith and religion, and union with the Franks to form one people.
In 808, an attack arrived from an unexpected source, the pagan Danes, “a race almost unknown to his ancestors, but destined to be only too well known to his sons,” as Charles Oman described them. Wikukind, whose wife was Danish, and his allies had taken refuge among the Danes. The king of the Danes, Godfred, built the vast Danevirke (shown below) across the isthmus of Schleswig. This defense was at its beginning a 30 km (19 mi) long defensive earthenwork rampart. The Danevirke protected Danish land and gave Godfred the opportunity to harass Frisia and Flanders with pirate raids.
Godfred invaded Frisia, joked of visiting Aachen, but was murdered before he could do any more, either by a Frankish assassin or by one of his own men. Godfred was succeeded by his nephew Hemming, who concluded the Treaty of Heiligen with Charlemagne in late 811.
Europe at the end of Charlemagne’s rein looked a lot different than it did at the beginning. The balance of power had shifted dramatically.

“Europe in 814, Charlemagne, Krum, Nicephorus I” by Stolichanin – Europe_plain_rivers.pngThe map is made according to:”World Atlas”, part 3: Europe in Middle Ages, Larrouse, Paris, 2002, O. RenieAtlas “History of Bulgaria”, Sofia, 1988, Bulgarian Academy of Sciences, V. Kamburova”World Atlas”, N. Ostrovski, Rome, 1992, p.55Атлас “История на средните векове”, Sofia, 1982, G. Gavrilov”History in maps”, Johannes Herder, Berlin, 1999, p. 20″European Historical Globus”, R. Rusev, 2006, p.117. Licensed under CC BY-SA 3.0 via Commons
Education
Charlemagne was determined to have his children educated, including his daughters, as he himself was not. Below, we see Charlemagne’s monogram from the subscription of a royal diploma. Signum Karolvs Karoli gloriosissimi regis.
For an uneducated man, he made amazing changes, including the standardization of the monetary system and instituting principles of accounting practice.
Charlemagne’s children were taught all the arts, and his daughters were learned in the “ways of being a woman.” whatever that meant at the time. His sons participated in archery, horsemanship, and other outdoor activities. This renaissance of education was referred to as the Carolingian Renaissance because it ushered in a new cultural era in which scholarship, literature and art thrived.
Charlemagne was brought into contact with the culture and learning of other countries (especially Moorish Spain, Anglo-Saxon England, and Lombard Italy) as a result of his vast conquests. He greatly increased the provision of monastic schools and scriptoria (centres for book-copying) in Francia.
Most of the presently surviving works of classical Latin were copied and preserved by Carolingian scholars. Indeed, the earliest manuscripts available for many ancient texts are Carolingian. It is almost certain that a text which survived to the Carolingian age survives still, thanks to Charlemagne.
Charlemagne took a serious interest in scholarship, promoting the liberal arts at the court, ordering that his children and grandchildren be well-educated. In a time when even leaders who promoted education did not take time to learn, Charlemagne studied. Under the tutelage of Peter of Pisa, Charlemagne learned grammar; with Alcuin, he studied rhetoric, dialectic (logic), and astronomy (he was particularly interested in the movements of the stars); and Einhard assisted him in his studies of arithmetic.
Charlemagne’s great scholarly failure, as Einhard relates, was his inability to write: when in his old age he began attempts to learn – practicing the formation of letters in his bed during his free time on books and wax tablets he hid under his pillow – “his effort came too late in life and achieved little success.” Charlemagne’s ability to read – which Einhard is silent about, and which no contemporary source supports – has also been called into question.
It appears that the man who was personally responsible for the salvation of so much literature was, himself, illiterate. Charlemagne was however very foresighted and progressive. What an amazing legacy.
Children and Heirs
Charlemagne had eighteen children over the course of his life with eight of his ten known wives or concubines. Nonetheless, he only had four legitimate grandsons, the four sons of his fourth son, Louis. In addition, he had a grandson (Bernard of Italy, the only son of his third son, Pippin of Italy), who was born illegitimate, but included in the line of inheritance. So, despite eighteen children, the claimants to his inheritance were few.
Charlemagne’s personal life is somewhat colorful for a devout Catholic, although maybe the cultural aspect of the church at that time was different. Then again, being the protector of the Pope may have gained Charlemagne special favors or caused some behaviors to be overlooked.
As I look at the dates, I have to wonder if these women and children all lived in one place and knew each other. Did the wife and concubine glare at each other when they met in the hallway, or were they more like compatriot sisters? Did they live in different locations? Did they consider themselves sucky to be Charlemagne’s partner, or did they wish things were different and that he was monogamous, as they had surely be raised in the Catholic church to expect of a husband.
I found the many references to Charlemagne’s concubines confusing, given his Catholicism and marriages at the same time, so I turned to the Catholic encyclopedia for clarification:
The Council of Toledo, held in 400, in its seventeenth canon legislates as follows for laymen (for ecclesiastical regulations on this head with regard to clerics see Celibacy): after pronouncing sentence of excommunication against any who in addition to a wife keep a concubine, it says: “But if a man has no wife, but a concubine instead of a wife, let him not be refused communion; only let him be content to be united with one woman, whether wife or concubine” (Can. “Is qui”, dist. xxxiv; Mansi, III, col. 1001). The refractory are to be excommunicated until such time as they shall obey and do penance.
It would appear, based on that edict, that Charlemagne’s concubines and extra-curricular activities were overlooked by the church, although many of his children were very active within the church as abbots and abbesses. Charlemagne recognized and provided in some way for all of his children, legitimate or otherwise.
Charlemagne’s sons fought many wars on behalf of their father when they came of age.
Charles was mostly preoccupied with the Bretons, whose border he shared and who insurrected on at least two occasions and were easily put down, but he was also sent against the Saxons on multiple occasions. In 805 and 806, he was sent into the Böhmerwald (modern Bohemia) to deal with the Slavs living there (Bohemian tribes, ancestors of the modern Czechs). He subjected them to Frankish authority and devastated the valley of the Elbe, forcing a tribute on them.
Pippin had to hold the Avar and Beneventan borders but also fought the Slavs to his north. He was uniquely poised to fight the Byzantine Empire when finally that conflict arose after Charlemagne’s imperial coronation and a Venetian rebellion.
Finally, Louis was in charge of the Spanish March and also went to southern Italy to fight the duke of Benevento on at least one occasion. He took Barcelona in a great siege in 797.
Charlemagne’s attitude toward his daughters has been the subject of much discussion. He kept them at home with him and refused to allow them to contract sacramental marriages – possibly to prevent the creation of cadet branches of the family to challenge the main line, as had been the case with Tassilo of Bavaria – yet he tolerated their extramarital relationships, even rewarding their common-law husbands, and treasured the illegitimate grandchildren they produced for him. At least one of them, Bertha, had a recognized relationship, if not a marriage, with Angilbert, a member of Charlemagne’s court circle.
Charlemagne also refused to believe stories of their wild behavior. He’s certainly not the first father to do that!
After Charlemagne’s death the surviving daughters were banished from the court by their brother, Louis, to take up residence in the convents they had been bequeathed by their father. That’s what happens when you are a bit rowdy and your brother who is known as Louis the Pious becomes king.
Charlemagne’s Death
In 806, Charlemagne first made provision for the traditional division of the empire on his death. For Charles the Younger he designated Austrasia and Neustria, Saxony, Burgundy, and Thuringia. To Pippin he gave Italy, Bavaria, and Swabia. Louis received Aquitaine, the Spanish March, and Provence. There was no mention of the imperial title however, which has led to the suggestion that, at that particular time, Charlemagne regarded the title as an honorary achievement which held no hereditary significance.
This division might have worked, but it was never to be tested. Pippin died in 810 and Charles in 811. Charlemagne then reconsidered the matter, and in 813, called Louis the Pious, king of Aquitaine, his only surviving legitimate son, to his court. There Charlemagne crowned his son with his own hands as co-emperor, granting him a half-share of the empire with the rest to follow up on Charlemagne’s death. Louis the Pious then returned to Aquitaine. The only part of the Empire which Louis was not promised was Italy, which Charlemagne specifically bestowed upon Pippin’s illegitimate son Bernard.
Charlemagne then spent the autumn hunting before returning to Aachen on November 1st. In January, he fell ill with pleurisy. In a deep depression (mostly because many of his plans were not yet realized), he took to his bed on January 21st. With all Charlemagne did achieve, I can’t imagine what he yet wanted that was severe enough to induce a depression of that magnitude. Historians attribute his depression to his plans not being realized, but I have to wonder if the deaths of his three sons and two daughters in two years (810-811) didn’t contribute to his depression. Another daughter had died in 808. That’s a lot of death in a short time.
Einhard tells us:
He died January twenty-eighth, the seventh day from the time that he took to his bed, at nine o’clock in the morning, after partaking of the Holy Communion, in the seventy-second year of his age and the forty-seventh of his reign.
Charlemagne was buried the same day as his death, in Aachen Cathedral, although the cold weather and the nature of his illness made such a hurried burial unnecessary. This causes me to wonder why he was buried so quickly.
Above, the Aachen Cathedral today.
Charlemagne is buried in the Palatine Chapel. He commissioned its construction in 792 and it was consecrated in 805 by Pope Leo III in honor of the Virgin Mary.

“Aachener Dom Pfalzkapelle vom Münsterplatz 2014” by CaS2000 – Own work. Licensed under CC BY-SA 3.0 via Commons
Charlemagne’s floor plan, below, includes a sixteen-sided ambulatory with a gallery overhead encircling the central octagonal dome.
The plan and decoration owe much to the sixth-century Basilica of San Vitale in Ravenna. Indeed, Charlemagne visited Ravenna three times, the first in 787. In that year he wrote to Pope Hadrian I and requested “mosaic, marbles, and other materials from floors and walls” in Rome and Ravenna, for his palace.
Interior view of the chapel. Charlemagne was buried someplace here, although the exact location evades detection. Theories abound and I have to believe that Charlemagne enjoys every minute of this mystery that his bones visit upon his descendants – which remember, includes all or most of Europe today.

“Königsthron Aachener Dom” by German Wikipedia user Holger Weinandt. Licensed under CC BY-SA 3.0 via Commons
Charlemagne’s throne, above, resides in the chapel as well.
This piece of fabric is part of Charlemagne’s death shroud, manufactured in Constantinople (present day Istanbul), and represents a quadriga, a cart or chariot that was drawn by 4 horses abreast. Typically Gods were depicted in this manner.
A later story, told by Otho of Lomello, Count of the Palace at Aachen in the time of Otto III, about the year 1000, would claim that he and Emperor Otto had discovered Charlemagne’s tomb. They claimed that Charlemagne was seated upon a throne, wearing a crown and holding a sceptre, his flesh almost entirely incorrupt.
In 1165, Frederick I re-opened the tomb again and placed the emperor in a sarcophagus beneath the floor of the cathedral.
Sarcophagus, above and below.
In 1215 Frederick II re-interred him in a casket made of gold and silver.
Charlemagne’s death greatly affected many of his subjects, particularly those of the literary clique who had surrounded him at Aachen. An anonymous monk of Bobbio lamented:
From the lands where the sun rises to western shores, people are crying and wailing … the Franks, the Romans, all Christians, are stung with mourning and great worry … the young and old, glorious nobles, all lament the loss of their Caesar … the world laments the death of Charles … O Christ, you who govern the heavenly host, grant a peaceful place to Charles in your kingdom. Alas for miserable me.
Charlemagne was succeeded by his surviving son, Louis, who had been crowned the previous year. Charlemagne’s empire lasted only another generation in its entirety; its division, according to custom, between Louis’s own sons after their father’s death laid the foundation for the modern states of Germany and France.
Below, the mask reliquary of Charlemagne. If this is a death mask of the man, he was in wonderful shape for 72 years of age and the battles he had been through.
Reliquary’s were and remain very popular in the Catholic Church. Below, Charlemagne’s arm reliquary.

“Charlemagne arm at Cathedral Treasury Aachen Germany” by Prof-Declercq – Own work. Licensed under CC BY-SA 4.0 via Commons
Charlemagne’s throne remains today in the Achen Cathedral as well.
Below, the interior of Charlemagne’s chapel in the Aachen Cathedral.
What Was Charlemagne Like?
We’re exceedingly lucky that Einhard was Charlemagne’s biographer. Otherwise we would know very little about him.
Charlemagne spoke French and German in addition to Latin and understood some Greek but spoke it very poorly. He mandated that sermons be preached in either “Romance” (French) or “Theotiscan” (German) and not in Latin so that the common person could understand the lessons being imparted.
How Did He Look?
Charlemagne’s personal appearance is known from a description by Einhard in his biography of Charlemagne titled “Vita Karoli Magni.” Einhard describes in his twenty-second chapter:
He was heavily built, sturdy, and of considerable stature, although not exceptionally so, since his height was seven times the length of his own foot. He had a round head, large and lively eyes, a slightly larger nose than usual, white but still attractive hair, a bright and cheerful expression, a short and fat neck, and he enjoyed good health, except for the fevers that affected him in the last few years of his life. Toward the end, he dragged one leg. Even then, he stubbornly did what he wanted and refused to listen to doctors, indeed he detested them, because they wanted to persuade him to stop eating roast meat, as was his wont, and to be content with boiled meat.
The physical portrait provided by Einhard is confirmed by contemporary depictions of the emperor, such as coins and his 8-inch (20 cm) bronze statue kept in the Louvre.

“Charlemagne denier Mayence 812 814” by PHGCOM – Own work by uploader, photographed at Cabinet des Médailles, Paris.. Licensed under CC BY-SA 3.0 via Commons
It’s uncertain whether this bronze statue from the Louvre depicts Charlemagne or his grandson, Charles the Bald, who reportedly favored his grandfather, Charlemagne.

Photographs credited © RMN, Musée du Louvre / [etc.] are the property of the RMN. Non-commercial re-use is authorized, provided the source and author are acknowledged. Photo by Jean-Giles Berizzi
Charlemagne wore the traditional costume of the Frankish people, described by Einhard thus:
He used to wear the national, that is to say, the Frank, dress—next his skin a linen shirt and linen breeches, and above these a tunic fringed with silk; while hose fastened by bands covered his lower limbs, and shoes his feet, and he protected his shoulders and chest in winter by a close-fitting coat of otter or marten skins.
He wore a blue cloak and always carried a sword with him. The typical sword was of a golden or silver hilt. He wore fancy jeweled swords to banquets or ambassadorial receptions. To Charlemagne, a sword was his ever-present and indispensable weapon, just in case, and sometimes a fine piece of jewelry too.
Nevertheless:
He despised foreign costumes, however handsome, and never allowed himself to be robed in them, except twice in Rome, when he donned the Roman tunic, chlamys, and shoes; the first time at the request of Pope Hadrian, the second to gratify Leo, Hadrian’s successor.
He could rise to the occasion when necessary. On great feast days, he wore embroidery and jewels on his clothing and shoes. He had a golden buckle for his cloak on such occasions and would appear with his great diadem, but he despised such apparel, according to Einhard, and usually dressed like the common people.
The Beautification of the Blessed Charles Augustus
Charlemagne was accorded sainthood inside the Holy Roman Empire after the twelfth century. His canonization by Antipope Paschal III, to gain the favor of Frederick Barbarossa in 1165, was never recognized by the Holy See, which annulled all of Paschal’s ordinances at the Third Lateran Council in 1179. Charlemagne’s name does not appear among the 28 saints named Charles who are listed in the Roman Martyrology. However, his beatification has been acknowledged as cultus confirmed and is celebrated on 28 January. Even in death, Charlemagne was contentious and deeply involved in the politics of religion.
Ummmm…so how does one act appropriately if one is twice descended from a King who was also a Saint??? Do I need to learn how to curtsey or do that Queen wave maybe? No one prepared me for this when I was learning proper manners. I had no idea how proper “proper” was!
Afterlife
Charlemagne, being a model knight as one of the Nine Worthies, enjoyed an important afterlife in European culture.
The Nine Worthies are nine historical, scriptural and legendary personages who personify the ideals of chivalry as were established in the Middle Ages.
The Nine Worthies include three good pagans: Hector, Alexander the Great, and Julius Caesar, three good Jews: Joshua, David, and Judas Maccabeus, and three good Christians: King Arthur, Charlemagne, and Godfrey of Bouillon.
Gateway Ancestors
If you’d like to see if you too are descended from Charlemagne through known gateway ancestors, please click here for a list. But beware, you just might have to learn how to behave “properly.” If you figure out exactly what that means, let me know.
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research