Update: Please note that the Genographic Project discontinued public sales of kits in 2019. I am leaving this article for historical context.
Yesterday, my husband’s Geno 2.0 results arrived. I bought his kit for him for Christmas initially, but couldn’t wait that long so had him swab the very day it arrived. So much for Santa’s surprise. But wait, maybe his results will hold a surprise – and isn’t this the first day of Christmas? Oh, it’s not? Well, I’ve decided to establish a new tradition this year, that it is the first day of Christmas and this year we will simply have 15 days of Christmas. The extra three days will be dedicated to Y-line SNPs, a mitochondrial haplogroup and ethnicity results. Who wants a bird in a pear tree anyway?
 Meet Jim. Since you’re going to be looking as his “innards,” so to speak, I think you should formally meet him. Pretty soon you’re going to be wondering what a Neanderthal, Denisovan, German, Greek, Mediterranean Hun looks like. I wanted to use the photo of him kissing the dolphin in the Carribbean, but he wouldn’t let me. Harumph. I guess Huns have no sense of humor.
Meet Jim. Since you’re going to be looking as his “innards,” so to speak, I think you should formally meet him. Pretty soon you’re going to be wondering what a Neanderthal, Denisovan, German, Greek, Mediterranean Hun looks like. I wanted to use the photo of him kissing the dolphin in the Carribbean, but he wouldn’t let me. Harumph. I guess Huns have no sense of humor.
There’s a lot of information included on the new Genographic results webpage with some very cool features. We’re not going to look at everything in-depth today, but we’ll do a flyover so you can see everything. Right now, because everyone is checking the status of their results, the Genographic webpage is experiencing difficulties. So if you click and wait, well, go get a cup of coffee. It will eventually respond and multiple clicks only cause buffer problems when it does respond. I know this from experience. How many times have I said that patience is not a virtue attributed to me!
Oh yes, and I had much better luck with the webpage using Chrome as opposed to Internet Explorer 7.
The first screen you see is “Your Story.” It’s large so I’ll break it into three parts for review. Notice the 5 tabs at the top of the page. You’ll be using those. Actually, if you follow the story line, the story walks you through them all.
The middle portion of this page shows your mitochondrial DNA, autosomal in the center, followed by your Y-line, if you’re a male. Your haplogroup is displayed as well for mitochondrial, and your terminal SNP for males, which is the SNP that ultimately defines the deepest level of your haplogroup. Well, at least until a new terminal SNP is discovered.
There are a couple of items of note here.
The tab titled “Your Map” simply won’t load, using either browser. I’ve sent a note to National Geographic. I know this is loading for other folks though. I wonder if it has anything to do with Jim’s haplogroup.
Second, for Jim’s paternal line, CTS11962 is a new SNP not previously tested at Family Tree DNA. CTS stands for Chris Tyler-Smith, a researcher in the UK at the Sanger Institute who discovered this particular SNP, and named it.
We knew in advance that the new standard for both the Geno 2.0 and also Family Tree DNA will be to only provide the terminal SNP and no longer rename the haplogroups in the way they used to be, such as R1a or R1b1a2. However, the problem with this approach is that someone looking at this terminal SNP has no reference whatsoever at this point. There isn’t even a “main” haplogroup branch given, nor a link to any explanation, or at least not that I can find.
After transferring Jim’s results to Family Tree DNA, I noticed that he is now labelled R1a1a1g on his Family Tree DNA haplogroup page. It looked like his mitochondrial results didn’t transfer initially, but this morning after the Family Tree DNA webpage update, he does have results on his mtDNA Haplogroup Origins page, which is about all you can do with haplogroup only information. Nat Geo, of course, does not test beyond haplogroup designation for mitochondrial. However, it looks like his autosomal results didn’t transfer anyplace. Hmmm….maybe a bug. I’ll have to contact the helpdesk who is probably swamped this morning.
Looking at his Family Tree DNA Haplogroup and SNPs page, all of the SNPs Jim was tested for at Nat Geo appear to be listed, but CTS11962, his terminal SNP, seems to be missing, so others may be as well. Family Tree DNA updated the data base and web page today, but this didn’t seem to be resolved with the update this morning. It also looks like it might be a page real estate issue in that this section of his page is quite full.
The bottom portion of the introductory Geno 2.0 page provides an interesting tidbit.
Yesterday, when one of my blog followers sent me his initial results, the “you are 1 of” number was 524,384, and that number had been used for some time, so it’s not dynamic. This new number appeared a few hours later, so I’m thinking that the answer to the number of Geno 2.0 kits sold might be someplace in the ballpark of 35,131. Just speculation on my part.
Let’s go back now and look at each section.
The mitochondrial DNA area provides a basic description of the haplogroup’s journey, then you can click to see the map.
Since his map won’t load, taking a look at another account I have access to, we see the following map information.
See the “share” button on the top right hand side of the next page? This allows you to send e-mails or a number of other types of messages to friends with a nice summary page of your results. Be sure to type a message in the message box with your full name because otherwise the recipient will have no idea whose info they are viewing.
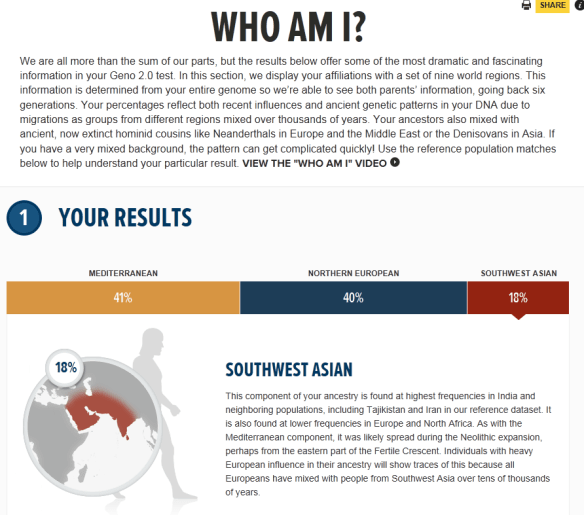
Moving to the autosomal section, we see that they have divided the world into 9 regions, plus Neanderthal and Denisovan.
Jim is Mediterranean, Northern European and Southwest Asian. His genealogy is entirely German and Hungarian. He’s a second generation immigrant, so none of his genealogy is lost here in the US. Of course, the term Hungarian should translate into “churning people” because the devastation of multiple wars spanning centuries has disrupted this population significantly. The same holds true for Germany. I would suggest that Germanic and Slavic would perhaps be a better description of his heritage.
In any case, the Mediterranean and Southwest Asian were somewhat of a surprise, but when you look at migration patterns, maybe not so much. After all, we all came through the Middle East on our way out of African and if your ancestors migrated to Germany, they probably went through the Mediterranean unless they went over the Caucus Mountains instead. And that would give you Southwest Asian heritage most likely.
The only problem is, now Jim wants to go on another Mediterranean cruise to commune with his ancestral homeland. DNA cruises anyone???
The next section explains what the results mean. This is actually quite interesting because it compares you to the actual reference population. In his case, in Germany.
His second reference population was somewhat of a surprise. It’s Greek. Perhaps it’s the best reference population they have for “Mediterranean” as the Greeks are truly a mixture of just about everyone in the Mediterranean basin.
They do provide links here to more information. By clicking on the overview of all reference populations, they provide information about how these populations as a whole are made up today.
I counted and there are 43 populations. Another option is to click to review the 9 world regions.
The Neanderthal and Denisovan ancestry is particularly interesting since it has been only recently, in the past couple of years, that humans were believed to have interbred with either group. Neanderthals have been found throughout most of Europe and western Asia, and Denisovans lived in the Altai Mountains of Siberia. If you click on the little links imbedded in many place on your page, you’ll see that there is more information about almost every topic. For example, they tell us that everyone outside of Africa carries some Neanderthal and Denisovan.
On Jim’s Genographic page, the terminal SNP shown is M458, shown below the map. Referring now to the ISOGG Y tree, we see that this equates to haplogroup R1a1a1b1a1. I didn’t expect CTS11962 to be on the ISOGG tree just yet, and it isn’t, but neither is it anyplace else, so we basically have our hands tied relative to where this falls on the R tree until someone places it someplace. I know that there is what is affectionately referred to as the “Big Paper” in the works that reworks the haplotree, and I surely hope it’s published sooner than later. Yes, I know, that patience thing again…..sigh.
You can also view this haplogroup information as a heatmap.
Under the “future” button, they ask you to complete your profile and contribute your story. You may have noticed that these options are stepping you across the tabs at the top of the introductory page.
Under the “Our Story” tab, which is where the “Contribute Your Story” link takes you, they explain about the community and social networking aspects of Geno 2.0.
Most interesting is the relationship circle, which looks like either a 45RPM record, a CD/DVD or the front of an i-pod, depending on your age. The stars and planets are the people who are “related” to you and the larger planets are those with stories attached. You just click on a planet to see their “story.” The closer they are to the center, the more closely related they are to you, but I’m unsure how “related” is gauged. For example if you click on the CTS11962 link, does this mean that everyone carries that mutation, or that it’s the terminal SNP for everyone listed? If it’s the terminal SNP for everyone listed, wouldn’t they all be equally as closely related? Jim’s V link doesn’t work, but the same questions apply.
And at the bottom of this page, you can add your own story.
I understand in the future that one will be able to contact these “related” people. If so, and if they haven’t tested their STR markers or their mitochondrial DNA at Family Tree DNA, we can encourage them, or invite them, to do so.
And speaking of which, the one thing I really DON’T like is what they’ve done with the download to Family Tree DNA option. In Geno 1.0, the transfer option said something like “to learn more,” but was tiny and obscure, at the bottom of a page where you had to scroll multiple times. I’m sure the reason that only 20% of the people ever transferred to Family Tree DNA was because only 20% of the people stumbled across the link and followed it. Actually, the fact that 20% actually found it is pretty amazing and speaks highly of how interested people actually are in the Genographic project and their results. This function and where it’s located is even worse in this version.
To initiate a transfer to Family Tree DNA, first, you have to know that you want to do this. There is no enticing “advertising” or education like there is for other information. This would be a wonderful opportunity for a nice video or at least a writeup and a link that at least says “to learn more” encourages people to investigate. Now, you need to go to your Profile link, and under the profile link, click on the Advanced Options.
Nope, I don’t think even 20% of the people will find this. I’ve very disappointed and hope they will reconsider and rework this option.
I am very hopeful that with some constructive and pleasant feedback that perhaps Nat Geo will reconsider and bring this transfer option into line with the otherwise wonderfully designed and world-class project pages. Their e-mail address is genographic@ngs.org.
Genographic has so much to offer and Family Tree DNA’s customers and project administrators have played a huge role in recruiting for both Geno 1.0 and 2.0. I’d really like to be sure that all Geno 2.0 participants receive everything they can out of the total genetic genealogy and anthropology experience. After all, this is the story of all of us, the human population, and we’re all connected. The only question is how closely we’re related, and when we get close enough in that tree, can we put names and faces on our ancestors, preferably with a few dates and locations as well? Hence, it’s a continuum from anthropology to genealogy. It needs to be a continuum from Nat Geo to FTDNA too.
One thing is for sure, we’re getting closer and closer as we learn more and more. The tools keep improving and one by one, those brick walls are falling.
Well, we’ve kicked the tires a bit and I think this one is a keeper. I think National Geographic has done a wonderful job of making a topic that has become increasingly complex over the past few years understandable to the average citizen who will take this test and look at their results with a sense of adventure and curiosity. The website is beautifully designed. Don’t shortchange yourself by hurrying through and not taking time to look at all of the photos, stories and videos, and be sure to click on the little “i” icon which means more information. There’s a lot here to savor, slowly, like a connoisseur of fine food or a wonderfully spellbinding book.
I think what I’ve decided though, relative to autosomal DNA testing and ethnicity is that there is no one answer, there is really only a matter of degrees. Today, with the different testing companies using different population data references, we receive differing results. I don’t worry too much unless they are significantly different, then I have to scratch my head a bit.
When my own results come back I’ll be doing some comparisons on how minority admixture stacks up using the different autosomal tests and analysis tools available. We kicked the tires today, but soon, we’ll be taking a test drive to see what we can actually do with all of this. So, stay tuned and in the mean time, savor those results! After all, on the first day of Christmas, my true love gave to me, a new SNP on my own family tree.
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research