Update: The WTY has been superceded by the Big Y test, but I’m leaving this article for historical continuity.
What the heck is WTY….and why do I care?
One of the reasons I started a blog is to continue what I do for my clients when I write their DNA reports. I make DNA understandable and fun for the normal air-breathing genealogist.
The past few days has been a whirlwind of information and announcements, some which tend to leave folks who don’t have a lot of experience in the dust.
For that, I do apologize. However, I’d like to tackle a much easier topic now, and that’s the WTY test. What is it and why is it so important?
WTY is short for Walk the Y, as in walk down the Y chromosome.
The tests we all order and love, at Family Tree DNA, that would be the 12, 25, 37, 67 and 111 marker tests, tell us about genealogy – who we are related to in the past several hundred years.
Deeper ancestry, anthropological in nature, a line I draw about the time when surnames were being adopted, is different and little information of that nature is exposed by the STR (short tandem repeat) genealogy markers.
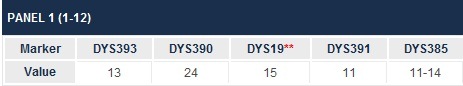
By the way, short tandem repeat means those locations in our DNA that are prone to develop repeated sequences. Think of them as genetic stutters. They are important to us as genealogists, because on the Y chromosome, we count the number of those stutters and that is the marker value reported.
For example, below, we see that for marker 393, we have a value of 13. That means there were 13 repeats of the same sequence. Obviously, combining all of these sequences, or marker values, together creates our own genealogical genetic profile or fingerprint. This, of course, is what we use to compare to others to see whom we match.
However, deep ancestry, identified by our haplogroup, is determined by a different kind of mutation, called a SNP, a single nucleotide polymorphism.
These are mutations that happen in only one location, and they are considered to be once in the lifetime of man mutations. In actuality, these mutations sometimes happen independently in different haplogroups, but the cumulative sequence of SNP mutations defines our haplogroup.
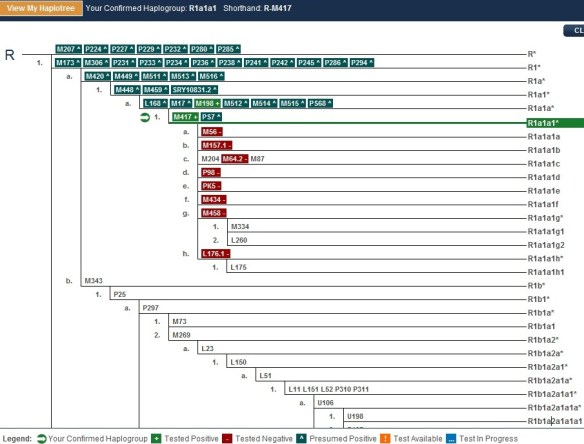
You can see, for example, below, a haplotree from a Family Tree DNA client’s results page.
This person tested positive for the light green SNP, M417. The plus means that they have this specific mutation. In his case, this is his terminal SNP, meaning the one furthest down the tree that defines his haplogroup, as we know it today. That would be R1a1a1.
The SNPs shown in red, below M417 are ones that he has also been tested for, but does not have, so he knows he is not a member of those haplogroups. These are shown with a minus sign, such as M56-.
Now for the problem that WTY has been helping to solve.
If your STR markers take you back about 500 years, in round numbers, and your haplogroup tells you where your ancestors were between 3000 and 4500 years ago, in this case, where were they in-between? What were they doing? Where did they live and how did they get from where they were 4500 years ago to where you find them 300 or 400 years ago, if you’re a lucky genealogist and can go back that far?
There is a significant gap in the timeframe between STR genealogy markers and haplogroup SNP markers. Finding additional SNPs will eventually close the gap between STR genealogy markers and haplogroups. We will have a complete timeline of our ancestors. In some cases, we’re even finding family-specific SNPs, known as “personal SNPs.” How cool is that? A new haplogroup is born in your family!
Did you notice on the tree above that some of the SNP markers begin with L? Every SNP discovered is prefaced with a letter that tells people which lab or university discovered the SNP. The L SNPs have all been discovered at Family Tree DNA’s Genomics Lab in Houston, Texas, run by Thomas Krahn. They are the product of the WTY discovery process.
When there is reason to believe that a SNP might be lurking undiscovered in the DNA of a person or a group, then the WTY becomes an option. Generally, the clue would be STR markers that are significantly different than any previously seen, or part of a small and quite unusual cluster.
Today, we test all of the known downstream SNPS, the ones in red above, and then if none are found, we would apply to Family Tree DNA to do a WTY test. This test is quite labor intensive. In essence, they manually look at between 450,000 and 500,000 positions to see if they spy any new mutations.
If they do, they begin the SNP naming process and the process of getting the SNP officially added onto the tree. You can see the most current haplotree (Y SNP tree) at the ISOGG site. Because of the long naming and authentication process, sometimes trees at different locations aren’t quite in sync. The ISOGG tree, maintained by volunteer genetic genealogists, has become what most people look to and use as the gold standard today.
In any case, this process is how new SNPs are discovered. The Geno 2.0 project includes 12,000 SNPs for the Y chromosome, an exponential growth from the current 862, or so. At least some of these SNPs were discovered at Family Tree DNA, as a result of savvy project administrators and others who are familiar enough with DNA results to suspect that a new SNP might exist, and who advocated with the tester and Family Tree DNA for WTY testing.
Hopefully, you now understand better about the WTY and why WTY tests are so critically important.
How might you know if you or a family member is a good candidate?
If you have tested to 67 or more markers and have no matches, you may be a candidate. You would need to do a deep clade test, which tests all relevant downstream SNPS at this point. In the past this has been the Deep Clade test, but today it would be the Geno 2.0 test. If you think you might be a candidate, you’ll want to work with your haplogroup administrator to see if there are any experimental SNPS to test for after the deep clade/Geno 2.0 is completed.
The WTY is the perfect example of collaborative citizen science. Participants fund part of the testing, haplogroup administrators identify good candidates, Family Tree DNA underwrites part of the testing fee and of course performs the test, and everyone benefits. Before you know it, you’ve got 12,000 new SNPs combined with new technology that promises to do more than we’ve ever dared dream before!!!
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research