Pre-release information from the paper, “Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans” which included results and analysis of DNA sequencing of 24,000 year old skeletal remains of a 4 year old Siberian boy caused quite a stir. Unfortunately, it was also misconstrued and incorrectly extrapolated in some articles. Some people misunderstood, either unintentionally or intentionally, and suggested that people with haplogroups U and R are Native American. That is not what either the prerelease or the paper itself says. Not only is that information and interpretation incorrect, the paper itself with the detailed information wasn’t published until November 20th, in Nature.
The paper is currently behind a paywall, so I’m going to discuss parts of it here, along with some additional information from other sources. To help with geography, the following google map shows the following locations: A=the Altai Republic, in Russia, B=Mal’ta, the location of the 24,000 year old skeletal remains and C=Lake Baikal, the region from where the Native American population originated in Asia.

Nature did publish an article preview. That information is in bold, italics and I will be commenting in nonbold, nonitalics.
The origins of the First Americans remain contentious. Although Native Americans seem to be genetically most closely related to east Asians1, 2, 3, there is no consensus with regard to which specific Old World populations they are closest to4, 5, 6, 7, 8. Here we sequence the draft genome of an approximately 24,000-year-old individual (MA-1), from Mal’ta in south-central Siberia9, to an average depth of 1×. To our knowledge this is the oldest anatomically modern human genome reported to date.
Within the paper, the authors also compare the MA-1 sequence to that of another 40,000 year old individual from Tianyuan Cave, China whose genome has been partially sequenced. This Chinese individual has been shown to be ancestral to both modern-day Asians and Native Americans. This comparison was particularly useful, because it showed that MA-1 is not closely related to the Tianyuan Cave individual, and is more closely related to Native Americans. This means that MA-1’s line and Tianyuan Cave’s line had not yet met and admixed into the population that would become the Native Americans. That occurred sometime later than 24,000 years ago and probably before crossing Beringia into North America sometime between about 18,000 and 20,000 years ago.
The MA-1 mitochondrial genome belongs to haplogroup U, which has also been found at high frequency among Upper Palaeolithic and Mesolithic European hunter-gatherers10, 11, 12, and the Y chromosome of MA-1 is basal to modern-day western Eurasians and near the root of most Native American lineages5.
The paper goes on to say that MA-1 is a member of mitochondrial (maternal) haplogroup U, very near the base of that haplogroup, but without affiliation to any known subclade, implying either that the subclade is rare or extinct in modern populations. In other words, this particular line of haplogroup U has NOT been found in any population, anyplace. According to the landmark paper, “A ‘‘Copernican’’ Reassessment of the Human Mitochondrial DNA Tree from its Root,” by Behar et al, 2012, haplogroup U itself was born about 46,500 years ago (plus or minus 3.200 years) and today has 9 major subclades (plus haplogroup K) and about 300 branching clades from those 9 subclades, excluding haplogroup K.
The map below, from the supplemental material included with the paper shows the distribution of haplogroup U, the black dots showing locations of haplogroup U comparison DNA.

In a recent paper, “Ancient DNA Reveals Key Stages in the Formation of Central European Mitochondrial Genetic Diversity” by Brandt et al (including the National Geographic Consortium) released in October 2013, the authors report that in the 198 ancient DNA samples collected from 25 German sites and compared to almost 68,000 current results, all of the ancient Hunter-Gatherer cultural results were haplogroup U, U4, U5 and U8. No other haplogroups were represented. In addition, those haplogroups disappeared from the region entirely with the advent of farming, shown on the chart below.

So, if someone who carries haplogroup U wants to say that they are distantly related to MA-1 who lived 24,000 years ago who was also related to their common ancestor who lived sometime prior to that, between 24,000 and 50,000 years ago, probably someplace between the Middle East where U was born, Mal’ta, Siberia and Western Europe, they would be correct. They are also distantly related to every other person in the world who carries haplogroup U, and many much more closely that MA-1 whose mitochondrial DNA line is either rare as chicken’s teeth (i.e. never found) or has gone extinct.
Let me be very clear about this, there is no evidence, none, that mitochondrial haplogroup U is found in the Native American population today that is NOT a result of post-contact admixture. In other words, in the burials that have been DNA tested, there is not one example in either North or South America of a burial carrying mitochondrial haplogroup U, or for that matter, male Y haplogroup R. Native American haplogroups found in the Americas remain subsets of mitochondrial haplogroups A, B, C, D and X and Y DNA haplogroups C and Q. Mitochondrial haplogroup M has potentially been found in one Canadian burial. No other haplogroups have been found. Until pre-contact remains are found with base haplogroups other than the ones listed above, no one can ethically claim that other haplogroups are of Native American origin. Finding any haplogroup in a contemporary Native population does not mean that it was originally Native, or that it should be counted as such. Admixture and adoption have been commonplace since Europeans first set foot on the soil of the Americas.
Now let’s talk about the Y DNA of MA-1.
The authors state that MA-1’s results are found very near the base of haplogroup R. They note that the sister lineage of haplogroup R, haplogroup Q, is the most common haplogroup in Native Americans and that the closest Eurasian Q results to Native Americans come from the Altai region.
The testing of the MA-1 Y chromosome was much more extensive than the typical STR genealogy tests taken by consumers today. MA-1’s Y chromosome was sequenced at 5.8 million base pairs at a coverage of 1.5X.
The resulting haplotree is shown below, again from the supplementary material.


The current haplogroup distribution range for haplogroup R is shown below, again with comparison points as black dots.

The current distribution range for Eurasian haplogroup Q is shown on the map below. Haplogroup Q is the most common haplogroup in Native Americans.

Similarly, we find autosomal evidence that MA-1 is basal to modern-day western Eurasians and genetically closely related to modern-day Native Americans, with no close affinity to east Asians. This suggests that populations related to contemporary western Eurasians had a more north-easterly distribution 24,000 years ago than commonly thought. Furthermore, we estimate that 14 to 38% of Native American ancestry may originate through gene flow from this ancient population. This is likely to have occurred after the divergence of Native American ancestors from east Asian ancestors, but before the diversification of Native American populations in the New World. Gene flow from the MA-1 lineage into Native American ancestors could explain why several crania from the First Americans have been reported as bearing morphological characteristics that do not resemble those of east Asians2, 13.
Kennewick Man is probably the most famous of the skeletal remains that don’t neatly fit into their preconceived box. Kennewick man was discovered on the bank of the Columbia River in Kennewick, Washington in 1996 and is believed to be from 7300 to 7600 years old. His anatomical features were quite different from today’s Native Americans and his relationship to ancient people is unknown. An initial evaluation and a 2010 reevaluation of Kennewick Man let to the conclusion by Doug Owsley, a forensic anthropologist, that Kennewick Man most closely resembles the Ainu people of Japan who themselves are a bit of an enigma, appearing much more Caucasoid than Asian. Unfortunately, DNA sequencing of Kennewick Man originally was ussuccessful and now, due to ongoing legal issues, more technologically advanced DNA testing has not been allowed. Nova sponsored a facial reconstruction of Kennewick Man which you can see here.
Sequencing of another south-central Siberian, Afontova Gora-2 dating to approximately 17,000 years ago14, revealed similar autosomal genetic signatures as MA-1, suggesting that the region was continuously occupied by humans throughout the Last Glacial Maximum. Our findings reveal that western Eurasian genetic signatures in modern-day Native Americans derive not only from post-Columbian admixture, as commonly thought, but also from a mixed ancestry of the First Americans.
In addition to the sequencing they set forth above, the authors compared the phenotype information obtainable from MA-1 to the Tyrolean Iceman, typically called Otzi. You can see Otzi’s facial reconstruction along with more information here. This is particularly interesting in light of the pigmentation change from darker skin in Africa to lighter skin in Eurasia, and the question of when this appearance change occurred. MA-1 shows a genetic affinity with the contemporary people of northern Europe, the population today with the highest frequency of light pigmentation phenotypes. The authors compared the DNA of MA-1 with a set of 124 SNPs identified in 2001 by Cerquira as informative on skin, hair and eye pigmentation color, although they also caution that this method has limited prediction accuracy. Given that, they say that MA-1 had dark hair, skin and eyes, but they were not able to sequence the full set of SNPs. MA-1 also had the SNP value associated with a high risk of male pattern baldness, a trait seldom found in Native American people and was not lactose tolerant, a trait found in western Eurasians. MA-1 also does not carry the mutation associated with hair thickness and shovel shaped incisors in Asians.
The chart below from the supplemental material shows the comparison with MA-1 and the Tyrolean Iceman.

The Tarim Mummies, found in the Tarim Basin in present-day Xinjiang, China are another example of remains that seem out of place. The earliest Tarim mummies, found at Qäwrighul and dated to 1800 BCE, are of a Europoid physical type whose closest affiliation is to the Bronze Age populations of southern Siberia, Kazakhstan, Central Asia, and the Lower Volga.
The cemetery at Yanbulaq contained 29 mummies which date from 1100–500 BCE, 21 of which are Mongoloid—the earliest Mongoloid mummies found in the Tarim Basin—and eight of which are of the same Europoid physical type found at Qäwrighul.
Notable mummies are the tall, red-haired “Chärchän man” or the “Ur-David” (1000 BCE); his son (1000 BCE), a small 1-year-old baby with brown hair protruding from under a red and blue felt cap, with two stones positioned over its eyes; the “Hami Mummy” (c. 1400–800 BCE), a “red-headed beauty” found in Qizilchoqa; and the “Witches of Subeshi” (4th or 3rd century BCE), who wore 2-foot-long (0.61 m) black felt conical hats with a flat brim. Also found at Subeshi was a man with traces of a surgical operation on his neck; the incision is sewn up with sutures made of horsehair.
Their costumes, and especially textiles, may indicate a common origin with Indo-European neolithic clothing techniques or a common low-level textile technology. Chärchän man wore a red twill tunic and tartan leggings. Textile expert Elizabeth Wayland Barber, who examined the tartan-style cloth, discusses similarities between it and fragments recovered from salt mines associated with the Hallstatt culture.
DNA testing revealed that the maternal lineages were predominantly East Eurasian haplogroup C with smaller numbers of H and K, while the paternal lines were all R1a1a. The geographic location of where this admixing took place is unknown, although south Siberia is likely. You can view some photographs of the mummies here.
In closing, the authors of the MA-1 paper state that the study has four important implications.
First, we find evidence that contemporary Native Americans and western Eurasians shareancestry through gene flow from a Siberian Upper Palaeolithic population into First Americans.
Second, our findings may provide an explanation for the presence of mtDNA haplogroup X in Native Americans, which is related to western Eurasians but not found in east Asian populations.
Third, such an easterly presence in Asia of a population related to contemporary western Eurasians provides a possibility that non-east Asian cranial characteristics of the First Americans derived from the Old World via migration through Beringia, rather than by a trans-Atlantic voyage from Iberia as proposed by the Solutrean hypothesis.
Fourth, the presence of an ancient western Eurasian genomic signature in the Baikal area before and after the LGM suggests that parts of south-central Siberia were occupied by humans throughout the coldest stages of the last ice age.
The times, they are a changin’.
Dr. Michael Hammer’s presentation at the 9th Annual International Conference on Genetic Genealogy may shed some light on all of this seeming confusing and somewhat conflicting information.
The graphic below shows the Y haplogroup base tree as documented by van Oven.
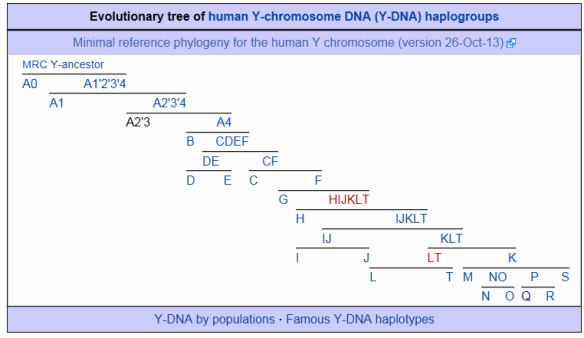

You can see, in the lower right corner, that Y haplogroup K (not to be confused with mtDNA haplogroup K discussed in conjunction with mtDNA haplogroup U) was the parent of haplogroup P which is the parent of both haplogroups Q and R.
It has always been believed that haplogroup R made its way into Europe before the arrival of Neolithic farmers about 10,000 years ago. However, that conclusion has been called into question, also by the use of Ancient DNA results. You can view additional information about Hammer’s presentation here, but in a nutshell, he said that there is no early evidence in burials, at all, for haplogroup R being in Europe at an early age. In about 40 burials from several location, haplogroup R has never been found. If it were present, especially in the numbers expected given that it represents more than half of the haplogroups of the men of Europe today, it should be represented in these burials, but it is not. Hammer concludes that evidence supports a recent spread of haplogroup R into Europe about 5000 years ago. Where was haplogroup R before spreading into Europe? In Asia.
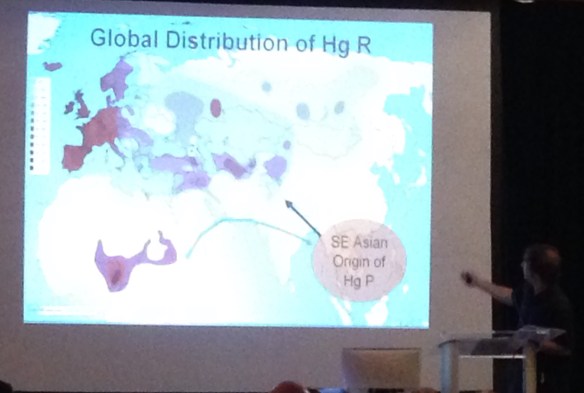
It appears that haplogroup K diversified in Southeast Asian, giving birth to haplogroups P, Q and R. Dr. Hammer said that this new information, combined with new cluster information and newly discovered SNP information over the past two years requires that haplogroup K be significantly revised. Between the revision of haplogroup K, the parent of both haplogroup R, previously believed to be European, and haplogroup Q, known to be Asian, European and Native, we may be in for a paradigm shift in terms of what we know about ancient migrations and who is whom. This path for haplogroup R into Europe really shouldn’t be surprising. It’s the exact same distribution as haplogroup Q, except haplogroup Q is much less frequently found in Europe than haplogroup R.
What Can We Say About MA-1?
In essence, we can’t label MA-1 as paternally European because of Y haplogroup R which now looks to have had an Asian genesis and was not known to have been in Europe 24,000 years ago, only arriving about 5,000 years ago. We can’t label haplogroup R as Native American, because it has never been found in a pre-Columbian New World burial.
We can say that mitochondrial haplogroup U is found in Europe in Hunter-Gatherer groups six thousand years ago (R was not) but we really don’t know if haplogroup U was in Europe 24,000 years ago. We cannot label haplogroup U as Native because it has never been found in a pre-Columbian New World burial.
We can determine that MA-1 did have ancestors who eventually became European due to autosomal analysis, but we don’t know that those people lived in what is now Europe 24,000 years ago. So the migration might have been into Europe, not out of Europe. MA-1, his ancestors and descendants, may have lived in Asia and subsequently settled in Europe or lived someplace inbetween. We can determine that MA-1’s line of people eventually admixed with people from East Asia, probably in Siberia, and became today’s First People of North and South America.
We can say that MA-1 appears to have been about 30% what is today Western Eurasian and that he is closely related to modern day Native Americans, but not eastern Asians. The authors estimate that between 14% and 38% of Native American ancestry comes from MA-1’s ancient population.
Whoever thought we could learn so much from a 4 year old?
For anyone seriously interested in Native American population genetics, “Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans” is a must read.
It’s been a great month for ancient DNA. Additional recent articles which pertain to this topic include:
http://www.nytimes.com/2013/11/21/science/two-surprises-in-dna-of-boy-found-buried-in-siberia.html?src=me&ref=general&_r=0
http://www.sciencedaily.com/releases/2013/11/131120143631.htm
http://dienekes.blogspot.com/2013/11/ancient-dna-from-upper-paleolithic-lake.html
http://blogs.discovermagazine.com/gnxp/2013/11/long-first-age-mankind/#.Uo0eOcSkrIU
http://cruwys.blogspot.com/2013/11/day-1-at-royal-societys-2013-ancient.html
http://cruwys.blogspot.co.uk/2013/11/day-2-at-royal-societys-2013-ancient.html
http://www.sciencedaily.com/releases/2013/11/131118081251.htm
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
Genealogy Services
Genealogy Research
Like this:
Like Loading...













































