At the November 2016 Family Tree DNA International Conference on Genetic Genealogy, I was invited to give a presentation about my Native American research findings utilizing the Genographic Project data base in addition to other resources. I was very pleased to be offered the opportunity, especially given that the 2016 conference marked the one year anniversary of the Genographic Project Affiliate Researcher program.
The results of this collaborative research effort have produced an amazing number of newly identified Native American mitochondrial haplogroups. Previously, 145 Native American mitochondrial haplogroups had been identified. This research project increased that number by 79% added another 114 haplogroups, raising the total to 259 Native American haplogroups.
Guilt by Genetic Association
Bennett Greenspan, President of Family Tree DNA, gave a presentation several years ago wherein he described genetic genealogy as “guilt by genetic association.” This description of genetic genealogy is one of the best I have ever heard, especially as it pertains to the identification of ancestral populations by Y and mitochondrial DNA.
As DNA testing has become more mainstream, many people want to see if they have Native ancestry. While autosomal DNA can only measure back in time relative to ethnicity reliably about 5 or 6 generations, Y and mitochondrial DNA due to their unique inheritance paths and the fact that they do not mix with the other parent’s DNA can peer directly back in time thousands of years.
Native American Mitochondrial DNA
Native American mitochondrial DNA consists of five base haplogroups, A, B, C, D and X. Within those five major haplogroups are found many Native as well as non-Native sub-haplogroups. Over the last 15 years, researchers have been documenting haplogroups found within the Native community although progress has been slow for various reasons, including but not limited to the lack of participants with proven Native heritage on the relevant matrilineal genealogical line.
In the paper, “Large scale mitochondrial sequencing in Mexican Americans suggests a reappraisal of Native American origins,” published in 2011, Kumar et al state the following:
For mtDNA variation, some studies have measured Native American, European and African contributions to Mexican and Mexican American populations, revealing 85 to 90% of mtDNA lineages are of Native American origin, with the remainder having European (5-7%) or African ancestry (3-5%). Thus the observed frequency of Native American mtDNA in Mexican/Mexican Americans is higher than was expected on the basis of autosomal estimates of Native American admixture for these populations i.e. ~ 30-46%. The difference is indicative of directional mating involving preferentially immigrant men and Native American women.
The actual Native mtDNA rate in their study of 384 completely sequenced Mexican genomes was 83.3% with 3.1% being African and 13.6% European.
This means that Mexican Americans and those south of the US in Mesoamerica provide a virtually untapped resource for Native American mitochondrial DNA.
The Genographic Project Affiliate Researcher Program
At the Family Tree DNA International Conference in November 2015, Dr. Miguel Vilar announced that the Genographic Project data base would be made available for qualified affiliate researchers outside of academia. There is, of course, an application process and aspiring affiliate researchers are required to submit a research project plan for consideration.
I don’t know if I was the first applicant, but if not, I was certainly one of the first because I wasted absolutely no time in submitting my application. In fact, my proposal likely arrived in Washington DC before Dr. Vilar did!
One of my original personal goals for genetic genealogy was to identify my Native American ancestors. It didn’t take long before I realized that one of the aspects of genetic genealogy where we desperately needed additional research was relative to Native people, specifically within Native language groups or tribes and from individuals who unquestionably know their ancestry and can document that their direct Y or mtDNA ancestors were Native.
Additionally, we needed DNA from pre-European-contact burials to ascertain whether haplogroups found in Europe and Africa were introduced into the Native population post-contact or existed within the Native population as a result of a previously unknown/undocumented contact. Some of both of these types of research has occurred, but not enough.
Slowly, over the years, additional sub-haplogroups have been added for both the Y and mitochondrial Native DNA. In 2007, Tamm et al published the first comprehensive paper providing an overview of the migration pathways and haplogroups in their landmark paper, “Beringian Standstill and the Spread of Native American Founders.” Other research papers have added to that baseline over the years.
In essence, whether you are an advocate of one migration or multiple migration waves, the dates of 10,000 to 25,000 years ago are a safe range for migration from Asia, across the then-present land-mass, Beringia, into the Americas. Recently another alternative suggesting that the migration may have occurred by water, in multiple waves, following coastlines, has been proposed as well – but following the same basic pathway. It makes little difference whether the transportation method was foot or kayak, or both, or one or more migration events. Our interest lies in identifying which haplogroups arrived with the Asians who became the indigenous people of the Americas.
Haplogroups
To date, proven base Native haplogroups are:
Y DNA:
- Q
- C
Mitochondrial DNA
- A
- B
- C
- D
- X
Given that the Native, First Nations or aboriginal people, by whatever name you call them, descended from Asia, across the Beringian land bridge sometime between roughly 10,000 and 25,000 years ago, depending on which academic model you choose to embrace, none of the base haplogroups shown above are entirely Native. Only portions, meaning specific subgroups, are known to be Native, while other subgroups are Asian and often European as well. The descendants of the base haplogroups, all born in Asia, expanded North, South, East and West across the globe. Therefore, today, it’s imperative to test mitochondrial DNA to the full sequence level and undergo SNP testing for Y DNA to determine subgroups in order to be able to determine with certainty if your Y or mtDNA ancestor was Native.
And herein lies the rub.
Certainty is relative, pardon the pun.
We know unquestionably that some haplogroups, as defined by Y SNPs and mtDNA full sequence testing, ARE Native, and we know that some haplogroups have never (to date) been found in a Native population, but there are other haplogroup subgroups that are ambiguous and are either found in both Asia/Europe and the Americas, or their origin is uncertain. One by one, as more people test and we obtain additional data, we solve these mysteries.
Let’s look at a recent example.
Haplogroup X2b4
Haplogroup X2b4 was found in the descendants of Radegonde Lambert, an Acadian woman born sometime in the 1620s and found in Acadia (present day Nova Scotia) married to Jean Blanchard as an adult. It was widely believed that she was the daughter of Jean Lambert and his Native wife. However, some years later, a conflicting record arose in which the husband of Radegonde’s great-granddaughter gave a deposition in which he stated that Radegonde came from France with her husband.
Which scenario was true? For years, no one else tested with haplogroup X2b4 that had any information as to the genesis of their ancestors, although several participants tested who descended from Radegonde.
Finally, in 2016, we were able to solve this mystery once and for all. I had formed the X2b4 project with Marie Rundquist and Tom Glad, hoping to attract people with haplogroup X2b4. Two pivotal events happened.
- Additional people tested at Family Tree DNA and joined the X2b4 project.
- Genographic Project records became available to me as an affiliate researcher.
At Family Tree DNA, we found other occurrences of X2b4 in:
- The Czech Republic
- Devon in the UK
- Birmingham in the UK
Was it possible that X2b4 could be both European and Native, meaning that some descendants had migrated east and crossed the Beringia land bridge, and some has migrated westward into Europe?
Dr. Doron Behar in the supplement to his publication, “A Copernican” Reassessment of the Human Mitochondrial DNA Tree from its Root” provides the creation dates for haplogroup X through X2b4 as follows:
These dates would read 31,718 years ago plus or minus 11,709 (eliminating the numbers after the decimal point) which would give us a range for the birth of haplogroup X from 43,427 years ago to 20,009 years ago, with 31,718 being the most likely date.
Given that X2b4 was “born” between 2,992 and 8,186 years ago, the answer has to be no, X2b4 cannot be found both in the Native population and European population since at the oldest date, 8,100 years ago, the Native people had already been in the Americas between 2,000 and 18,000 years.
Of course, all kinds of speculation could be (and has been) offered, about Native people being taken to Europe, although that speculation is a tad bit difficult to rationalize in the Czech Republic.
The next logical question is if there are documented instances of X2b4 in the Native population in the Americas?
I turned to the Genographic Project where I found no instances of X2b4 in the Native population and the following instances of X2b4 in Europe.
- Ireland
- Czech
- Serbia
- Germany (6)
- France (2)
- Denmark
- Switzerland
- Russia
- Warsaw, Poland
- Norway
- Romania
- England (2)
- Slovakia
- Scotland (2)
The conclusion relative to X2b4 is clearly that X2b4 is European, and not aboriginally Native.
The Genographic Project Data Base
As a researcher, I was absolutely thrilled to have access to another 700,000+ results, over 475,000 of which are mitochondrial.
The Genographic Project tests people whose identity remains anonymous. One of the benefits to researchers is that individuals in the public participation portion of the project can contribute their own information anonymously for research by answering a series of questions.
I was very pleased to see that one of the questions asked is the location of the birth of the participant’s most distant matrilineal ancestor.
Tabulation and analysis should be a piece of cake, right? Just look at that “most distant ancestor” response, or better yet, utilize the Genographic data base search features, sort, count, and there you go…
Well, guess again, because one trait that is universal, apparently, between people is that they don’t follow instructions well, if at all.
The Genographic Project, whether by design or happy accident, has safeguards built in, to some extent, because they ask respondents for the same or similar information in a number of ways. In any case, this technique provides researchers multiple opportunities to either obtain the answer directly or to put 2+2 together in order to obtain the answer indirectly.
Individuals are identified in the data base by an assigned numeric ID. Fields that provide information that could be relevant to ascertaining mitochondrial ethnicity and ancestral location are:
I utilized these fields in reverse order, giving preference to the earliest maternal ancestor (green) fields first, then maternal grandmother (teal), then mother (yellow), then the tester’s place of birth (grey) supplemented by their location, language and ethnicity if applicable.
Since I was looking for very specific information, such as information that would tell me directly or suggest that the participant was or could be Native, versus someone who very clearly wasn’t, this approach was quite useful.
It also allowed me to compare answers to make sure they made sense. In some cases, people obviously confused answers or didn’t understand the questions, because the three earliest ancestor answers cannot contain information that directly contradict each other. For example, the earliest ancestor place of birth cannot be Ireland and the language be German and the ethnicity be Cherokee. In situations like this, I omitted the entire record from the results because there was no reliable way to resolve the conflicting information.
In other cases, it was obvious that if the maternal grandmother and mother and tester were all born in China, that their earliest maternal ancestor was not very likely to be Native American, so I counted that answer as “China” even though the respondent did not directly answer the earliest maternal ancestor questions.
Unfortunately, that means that every response had to be individually evaluated and tabulated. There was no sort and go! The analysis took several weeks in the fall of 2016.
By Haplogroup – Master and Summary Tables
For each sub-haplogroup, I compiled, minimally, the following information shown as an example for haplogroup A with no subgroup:
The “Previously Proven Native” link is to my article titled Native American Mitochondrial Haplogroups where I maintain an updated list of haplogroups proven or suspected Native, along with the source(s), generally academic papers, for that information.
In some cases, to resolve ambiguity if any remained, I also referenced Phylotree, mtDNA Community and/or GenBank.
For each haplogroup or subgroup within haplogroup, I evaluated and listed the locations for the Genographic “earliest maternal ancestor place of birth” locations, but in the case of the haplogroup A example above, with 4198 responses, the results did not fit into the field so I added the information as supplemental.
By analyzing this information after completing a master tablet for each major haplogroup and subgroups, meaning A, B, C, D and X, I created summary tables provided in the haplogroup sections in this paper.
Family Tree DNA Projects
Another source of haplogroup information is the various mitochondrial DNA projects at Family Tree DNA.
Each project is managed differently, by volunteers, and displays or includes different information publicly. While different information displayed and lack of standardization does present challenges, there is still valuable information available from the public webpages for each mitochondrial haplogroup referenced.
Challenges
The first challenge is haplogroup naming. For those “old enough” to remember when Y DNA haplogroups used to be called by names such as R1b1c and then R1b1a2, as opposed to the current R-M269 – mitochondrial DNA is having the same issue. In other words, when a new branch needs to be added to the tree, or an entire branch needs to be moved someplace else, the haplogroup names can and do change.
In October and November 2016 when I extracted Genographic project data, Family Tree DNA was on Phylotree version 14 and the Genographic Project was on version 16. The information provided in various academic papers often references earlier versions of the phylotree, and the papers seldom indicate which phylotree version they are using. Phylotree is the official name for the mitochondrial DNA haplogroup tree.
Generally, between Phylotree versions, the haplogroup versions, meaning names, such as A1a, remain fairly consistent and the majority of the changes are refinements in haplogroup names where subgroups are added and all or part of A1a becomes A1a1 or A1a2, for example. However, that’s not always true. When new versions are released, some haplogroup names remain entirely unchanged (A1a), some people fall into updated haplogroups as in the example above, and some find themselves in entirely different haplogroups, generally within the same main haplogroup. For example, in Phylotree version 17, all of haplogroup A4 is obsoleted, renamed and shifted elsewhere in the haplogroup A tree.
The good news is that both Family Tree DNA and the Genographic project plan to update to Phylotree V17 in 2017. After that occurs, I plan to “equalize” the results, hopefully “upgrading” the information from academic papers to current haplogroup terminology as well if the authors provided us with the information as to the haplogroup defining mutations that they utilized at publication along with the entire list of sample mutations.
A second challenge is that not all haplogroup projects are created equal. In fact, some are entirely closed to the public, although I have no idea why a haplogroup project would be closed. Other projects show only the map. Some show surnames but not the oldest ancestor or location. There was no consistency between projects, so the project information is clearly incomplete, although I utilized both the public project pages and maps together to compile as much information as possible.
A third challenge is that not every participant enters their most distant ancestor (correctly) nor their ancestral location, which reduces the relevance of results, whether inside of projects, meaning matches to individual testers, or outside of projects.
A fourth challenge is that not every participant enables public project sharing nor do they allow the project administrators to view their coding region results, which makes participant classification within projects difficult and often impossible.
A fifth challenge is that in Family Tree DNA mitochondrial projects, not everyone has tested to the full sequence level, so some people who are noted as base haplogroup “A,” for example, would have a more fully defined haplogroup is they tested further. On the other hand, for some people, haplogroup A is their complete haplogroup designation, so not all designations of haplogroup A are created equal.
A sixth challenge is that in the Genographic Project, everyone has been tested via probes, meaning that haplogroup defining mutation locations are tested to determine full haplogroups, but not all mitochondrial locations are not tested. This removes the possibility of defining additional haplogroups by grouping participants by common mutations outside of haplogroup defining mutations.
A seventh challenge is that some resources for mitochondrial DNA list haplogroup mutations utilizing the CRS (Cambridge Reference Sequence) model and some utilize the RSRS (Reconstructed Sapiens Reference Sequence) model, meaning that the information needs to be converted to be useful.
Resources
Let’s look at the resources available for each resource type utilized to gather information.
The table above summarizes the differences between the various sources of information regarding mitochondrial haplogroups.
Before we look at each Native American haplogroup, let’s look at common myths, family stories and what constitutes proof of Native ancestry.
Family Stories
In the US, especially in families with roots in Appalachia, many families have the “Cherokee” or “Indian Princess” story. The oral history is often that “grandma” was an “Indian princess” and most often, Cherokee as well. That was universally the story in my family, and although it wasn’t grandma, it was great-grandma and every single line of the family carried this same story. The trouble was, it proved to be untrue.
Not only did the mitochondrial DNA disprove this story, the genealogy also disproved it, once I stopped looking frantically for any hint of this family line on the Cherokee rolls and started following where the genealogy research indicated. Now, of course this isn’t to say there is no Native IN that line, but it is to say that great-grandma’s direct matrilineal (mitochondrial) line is NOT Native as the family story suggests. Of course family stories can be misconstrued, mis-repeated and embellished, intentionally or otherwise with retelling.
Family stories and myths are often cherished, having been handed down for generations, and die hard.
In fact, today, some unscrupulous individuals attempt to utilize the family myths of those who “self-identify” their ancestor as “Cherokee” and present the myths and resulting non-Native DNA haplogrouip results as evidence that European and African haplogroups are Native American. Utilizing this methodology, they confirm, of course, that everyone with a myth and a European/African haplogroup is really Native after all!
As the project administrator of several projects including the American Indian and Cherokee projects, I can tell you that I have yet to find anyone who has a documented, as in proven lineage, to a Native tribe on a matrilineal line that does not have a Native American haplogroup. However, it’s going to happen one day, because adoptions of females into tribes did occur, and those adopted females were considered to be full tribal members. In this circumstance, your ancestor would be considered a tribal member, even if their DNA was not Native.
Given the Native tribal adoption culture, tribal membership of an individual who has a non-Native haplogroup would not be proof that the haplogroup itself was aboriginally Native – meaning came from Asia with the other Native people and not from Europe or Africa with post-Columbus contact. However, documenting tribal membership and generational connectivity via proven documentation for every generation between that tribally enrolled ancestor and the tester would be a first step in consideration of other haplogroups as potentially Native.
In Canada, the typical story is French-Canadian or metis, although that’s often not a myth and can often be proven true. We rely on the mtDNA in conjunction with other records to indicate whether or not the direct matrilineal ancestor was French/European or aboriginal Canadian.
In Mexico, the Caribbean and points south, “Spain” in the prevalent family story, probably because the surnames are predominantly Spanish, even when the mtDNA very clearly says “Native.” Many family legends also include the Canary Islands, a stopping point in the journey from Europe to the Caribbean.
Cultural Pressures
It’s worth noting that culturally there were benefits in the US to being Native (as opposed to mixed blood African) and sometimes as opposed to entirely white. Specifically, the Native people received head-right land payments in the 1890s and early 1900s if they could prove tribal descent by blood. Tribal lands, specifically those in Oklahoma owned by the 5 Civilized Tribes (Cherokee, Choctaw, Chickasaw, Creek and Seminole) which had been previously held by the tribe were to be divided and allotted to individual tribal members and could then be sold. Suddenly, many families “remembered” that they were of Native descent, whether they were or not.
Culturally and socially, there may have been benefits to being Spanish over Native in some areas as well.
It’s also easy to see how one could assume that Spain was the genesis of the family if Spanish was the spoken language – so care had to be exercised when interpreting some Genographic answers. Chinese can be interpreted to mean “China” or at least Asia, meaning, in this case, “not Native,” but Spanish in Mexico or south of the US cannot be interpreted to mean Spain without other correlating information.
Language does not (always) equal origins. Speaking English does not mean your ancestors came from England, speaking Spanish does not mean your ancestors came from Spain and speaking French does not mean your ancestors came from France.
However, if your ancestors lived in a country where the predominant language was English, Spanish or French, and your ancestor lived in a location with other Native people and spoke a Native language or dialect, that’s a very compelling piece of evidence – especially in conjunction with a Native DNA haplogroup.
What Constitutes Proof?
What academic papers use as “proof” of Native ancestry varies widely. In many cases, the researchers don’t make a case for what they use as proof, they simply state that they had one instance of A2x from Mexico, for example. In other cases, they include tribal information, if known. When stated in the papers, I’ve included that information on the Native American Mitochondrial Haplogroups page.
Methodology
I have adopted a similar methodology, tempered by the “guilt by genetic association” guideline, keeping in mind that both FTDNA projects and Genographic project public participants all provide their own genealogy and self-identify. In other words, no researcher traveled to Guatemala and took a cheek swab or blood sample. The academic samples and samples taken by the Genographic Project in the field are not included in the Genographic public data base available to researchers.
However, if the participant and their ancestors noted were all born in Guatemala, there is no reason to doubt that their ancestors were also found in the Guatemala region.
Unfortunately, not everything was that straightforward.
Examples:
- If there were multiple data base results as subsets of base haplogroups previously known to be Native from Mexico and none from anyplace else in the world, I’m comfortable calling the results “Native.”
- If there are 3 results from Mexico, and 10 from Europe, especially if the European results are NOT from Spain or Portugal, I’m NOT comfortable identifying that haplogroup as Native. I would identify it as European so long as the oldest date in the date ranges identifying when the haplogroup was born is AFTER the youngest migration date. For example, if the haplogroup was born 5,000 years ago and the last known Beringia migration date is 10,000 years ago, people with the same haplogroup cannot be found both in Europe and the Americas indigenously. If the haplogroup birth date is 20,000 years ago and the migration date is 10,000 years ago, clearly the haplogroup CAN potentially be found on both continents as indigenous.
- In some cases, we have the reverse situation where the majority of results are from south of the US border, but one or two claim Spanish or Portuguese ancestry, which I suspect is incorrect. In this case, I will call the results Native so long as there are a significant number of results that do NOT claim Spanish or Portuguese ancestry AND none of the actual testers were born in Spain or Portugal.
- In a few cases, the FTDNA project and/or Genographic data refute or at least challenge previous data from academic papers. Future information may do the same with this information today, especially where the data sample is small.
Because of ambiguity, in the master data table (not provided in this paper) for each base haplogroup, I have listed every one of the sub-haplogroups and all the locations for the oldest ancestors, plus any other information provided when relevant in the actual extracted data.
When in doubt, I have NOT counted a result as Native. When the data itself is questionable or unreliable, I removed the result from the data and count entirely.
I intentionally included all of the information, Native and non-Native, in my master extracted data tables so that others can judge for themselves, although I am only providing summary tables here. Detailed information will be provided in a series of articles or in an academic paper after both the Family Tree DNA data base and the Genographic data base are upgraded to Phylotree V17.
The Haplogroup Summary Table
The summary table format used for each haplogroup includes the following columns and labels:
- Hap = Haplogroup as listed at Family Tree DNA, in academic papers and in the Genographic project.
- Previous Academic Proven = Previously proven or cited as Native American, generally in Academic papers. A list of these haplogroups and papers is provided in the article, Native American Mitochondrial Haplogroups.
- Academic Confirmed = Academic paper haplogroup assignments confirmed by the Genographic Project and/or Family Tree DNA Projects.
- Previous Suspected = Not academically proven or cited at Native, but suspected through any number of sources. The reasons each haplogroup is suspected is also noted in the article, Native American Mitochondrial DNA Haplogroups.
- Suspected Confirmed = Suspected Native haplogroups confirmed as Native.
- FTDNA Project Proven = Mitochondrial haplogroup proven or confirmed through FTDNA project(s).
- Geno Confirmed = Mitochondrial haplogroup proven or confirmed through the Genographic Project data base.
Color Legend:
Additional Information:
- Possibly, probably or uncertain indicates that the data is not clear on whether the haplogroup is Native and additional results are needed before a definitive assignment is made.
- No data means that there was no data for this haplogroup through this source.
- Hap not listed means that the original haplogroup is not listed in the Genographic data base indicating the original haplogroup has been obsoleted and the haplogroup has been renamed.
The following table shows only the A haplogroups that have now been proven Native, omitting haplogroups proven not to be Native through this process, although the original master data table (not included here) includes all information extracted including for haplogroups that are not Native. Summary tables show only Native or potentially Native results.
Let’s look at the summary results grouped by major haplogroup.
Haplogroup A
Haplogroup A is the largest Native American haplogroup.
More than 43% of the individuals who carry Native American mitochondrial DNA fall into a subgroup of A.
Like the other Native American haplogroups, the base haplogroup was formed in Asia.
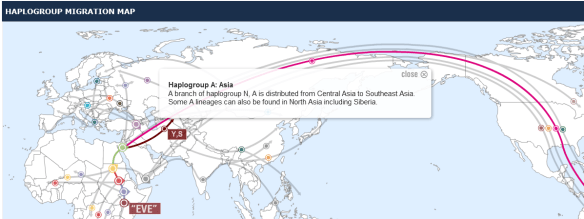
Family Tree DNA individual participant pages provide participants with both a Haplogroup Frequency Map, shown above, and a Haplogroup Migration Map, shown below.
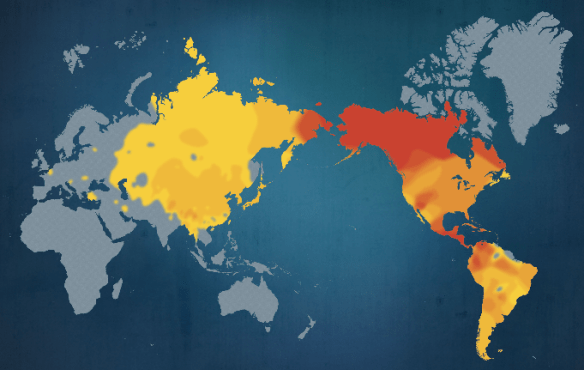
The Genographic project provides heat maps showing the distribution of major haplogroups on a continental level. You can see that, according to this heat map from when the Genographic Project was created, the majority of haplogroup A is found in the northern portion of the Americas.
Additionally, the Genographic Project data base also provides a nice tree structure for each haplogroup, beginning with Mitochondrial Eve, in Africa, noted as the root, and progressing to the current day haplogroups.
Haplogroup A Projects
I enjoy the added benefit of being one of the administrators, along with Marie Rundquist, of the haplogroup A project at Family Tree DNA, as well as the A10, A2 and A4 projects. However, in this paper, I only included information available on the projects’ public pages and not information participants sent to the administrators privately.
The Haplogroup A Project at Family Tree DNA is a public project, meaning available for anyone with haplogroup A to join, and fully publicly viewable with the exception of the participant’s surname, since that is meaningless when the surname traditionally changes with every generation. However, both the results, complete with the Maternal Ancestor Name, and the map, are visible. HVR1 and HVR2 results are displayed, but coding region results are never available to be shown in projects, by design.
The map below shows all participants for the entire project who have entered a geographic location. The three markers in the Middle East appear to be mis-located, a result of erroneous user geographic location input. The geographic locations are selected by participants indicating the location of their most distant mitochondrial ancestor. All 3 are Spanish surnames and one is supposed to be in Mexico. Please disregard those 3 Middle Eastern pins on the map below.
Haplogroup A Summary Table
The subgroups of haplogroup A and the resulting summary data are shown in the table below.
- Total haplogroups Native – 75
- Total haplogroups uncertain – 1
- Total haplogroups probable – 1
- Total new Native haplogroups – 38, 1 probable.
- Total new Native haplogroups proven by FTDNA Projects – 9, 1 possibly
- Total new Native haplogroups proven by Genographic Project – 35, 1 probable
Haplogroup B
Haplogroup B is the second largest Native American haplogroup, with 23.53% of Native participants falling into this haplogroup.
The Genographic project provides the following heat map for haplogroup B4, which includes B2, the primary Native subgroup.
The haplogroup B tree looks like this:
B4 and B5 are main branches.
You will note below that B2 falls underneath B4b.
Haplogroup B Projects
At Family Tree DNA, there is no haplogroup B project, but there is a haplogroup B2 project, which is where the majority of the Native results fall. Haplogroup B Project administrators have included a full project display, along with a map. All of the project participants are shown on the map below.
Please note that the pins colored other than violet (haplogroup B) should not be shown in this project. Only haplogroup B pins are violet.
Haplogroup B Summary Table
- Total haplogroups Native – 63
- Total haplogroups refuted – 1
- Total new Native haplogroups – 43
- Total new Native haplogroups proven by Family Tree DNA projects – 12
- Total new Native haplogroups proven by Genographic Project – 41
Haplogroup C
Haplogroup C is the third largest Native haplogroup with 22.99% of the Native population falling into this haplogroup.
Haplogroup C is primarily found in Asia per the Genographic heat map.
The haplogroup C tree is as follows:
Haplogroup C Project
Unfortunately, at Family Tree DNA, the haplogroup C project has not enabled their project pages, even for project members.
When I first began compiling this data, the Haplogroup C project map was viewable.
Haplogroup C Summary Table
- Total haplogroups Native – 61
- Total haplogroups refuted – 2
- Total haplogroups possible – 1
- Total haplogroups probable – 1
- Total new Native haplogroups – 8
- Total new Native haplogroups proven by Family Tree DNA projects – 6
- Total new Native haplogroups proven by Genographic Project – 5, 1 possible, 1 probable
Haplogroup D
Haplogroup D is the 4th largest, or 2nd smallest Native haplogroup, depending on your point of view, with 6.38% of Native participants falling into this haplogroup.
Haplogroup D is found throughout Asia, into Europe and throughout the Americas.
Haplogroups D1 and D2 are the two subgroups primarily found in the New World.
The haplogroup D1 heat map is shown above and D2 is shown below.
The Tree for haplogroup D is a subset of M.
Haplogroup D begins as a subhaplogroup of M80..
Haplogroup D Projects
D is publicly viewable, but shows testers last name, no ancestor information and no location, so I utilized maps once again.
Haplogroup D Summary Table
- Total haplogroups Native – 50
- Total haplogroups possibly both – 3
- Total haplogroups uncertain – 2
- Total haplogroups probable – 1
- Total haplogroups refuted – 3
- Total new Native Haplogroups – 25
- Total new Native haplogroups proven by Family Tree DNA projects – 2
- Total new Native haplogroups proven by Genographic Project – 22, 1 probably
Haplogroup X
Haplogroup X is the smallest of the known Native base haplogroups.
Just over 3% of the Native population falls into haplogroup X.
The heat map for haplogroup X looks very different than haplogroups A-D.
The tree for haplogroup X shows that it too is also a subgroup of M and N.
Haplogroup X Project
At Family Tree DNA, the Haplogroup X project is visible, but with no ancestral locations displayed. I utilized the map, which was visible.
This map of the entire haplogroup X project tells you immediately that the migration route for Native X was not primarily southward, but east. Haplogroup X is found primarily in the US and in the eastern half of Canada.
Haplogroup X Summary Table
- Total haplogroups Native – 10
- Total haplogroups uncertain, possible or possible both Native and other – 8
- Total New Native haplogroups – 0
Haplogroup M
Haplogroup M, a very large, old haplogroup with many subgroups, is not typically considered a Native haplogroup.
The Genographic project shows the following heat map for haplogroup M.
The heat map for haplogroup M includes both North and South America, but according to Dr. Miguel Vilar, Science Manager for the Genographic Project, this is because both haplogroups C and D are subsets of M.
The haplogroup M migration map from the Genographic Project shows haplogroup M expanding across southern Asia.
The tree for haplogroup M, above, is abbreviated, without the various subgroups being expanded.
The M1 and M1a1e haplogroups shown above are discussed in the following section, as is M18b, below.
The Haplogroup M Project
The haplogroup M project at Family Tree DNA shows the worldwide presence of haplogroup M and subgroups.
Native Presence
Haplogroup M was originally reported in two Native burials in the Americas. Dr. Ripan Malhi reported haplogroup M (excluding M7, M8 and M9) from two separate skeletons from the same burial in China Lake, British Columbia, Canada, about 150 miles north of the Washington State border, dating from about 5000 years ago. Both skeletons were sequenced separately in 2007, with identical results and are believed to be related.
While some researchers are suspicious of these findings as being incomplete, a subsequent paper in 2013, Ancient DNA-Analysis of Mid-Holocene Individuals from the Northwest Coast of North America Reveals Different Evolutionary Paths for Mitogenomes, which included Mahli as a co-author states the following:
Two individuals from China Lake, British Columbia, found in the same burial with a radiocarbon date of 4950+/−170 years BP were determined to belong to a form of macrohaplogroup M that has yet to be identified in any extant Native American population [24], [26]. The China Lake study suggests that individuals in the early to mid-Holocene may exhibit mitogenomes that have since gone extinct in a specific geographic region or in all of the Americas.
Haplogroup M Summary Table
One additional source for haplogroup M was found in GenBank noted as M1a1e “USA”, but there were also several Eurasian submissions for M1a1e as well. However, Doron Behar’s dates for M1a1e indicate that the haplogroup was born about 9,813 years ago, plus or minus 4,022 years, giving it a range of 5,971 to 13,835 years ago, meaning that M1a1e could reasonably be found in both Asia and the Americas. There were no Genographic results for M1a1e. At this point, M1a1e cannot be classified as Native, but remains on the radar.
Hapologroup M1 was founded 23,679 years ago +-4377 years. It is found in the Genographic Project in Cuba, Venezuela and is noted as Native in the Midwest US. M1 is also found in Colorado and Missouri in the haplogroup M project at Family Tree DNA, but the individuals did not have full sequence tests nor was additional family information available in the public project.
The following information is from the master data table for haplogroup M potentially Native haplogroups.
Haplogroup M Master Data Table for Potentially Native Haplogroups
The complete master data tables includes all subhaplogroups of M, the partial table below show only the Native haplogroups.
Haplogroup M18b is somewhat different in that two individuals with this haplogroup at Family Tree DNA have no other matches. They both have a proven connection to Native families from interrelated regions in North Carolina.
I initiated communications with both individuals who tested at Family Tree DNA who subsequently provided their genealogical information. Both family histories reach back into the late 1700s, one in the location where the Waccamaw were shown on maps in in the early 1700s, and one near the border of Virginia and NC. One participant is a member of the Waccamaw tribe today. A family migration pattern exists between the NC/VA border region and families to the Waccamaw region as well. An affidavit exists wherein the family of the individual from the NC/VA border region is sworn to be “mixed” but with no negro blood.
In summary:
- Haplogroups M and M1 could easily be both Native as well as Asian/European, given the birth age of the haplogroup.
- Haplogroup M1a1e needs additional results.
- Haplogroup M18b appears to be Native, but could also be found elsewhere given the range of the haplogroup birth age. Additional proven Native results could bolster this evidence.
- In addition to the two individuals with ancestors from North Carolina, M18b is also reported in a Sioux individuals with mixed race ethnicity
The Dark Horse Late Arrival – Haplogroup F
I debated whether I should include this information, because it’s tenuous at best.
The American Indian project at Family Tree DNA includes a sample of F1a1 full sequence result whose most distant matrilineal ancestor is found in Mexico.
Haplogroup F is an Asian haplogroup, not found in Europe or in the Americas.
Haplogroup F, according to the Genographic Project, expands across central and southern Asia.
According to Doron Behar, F1a1 was born about 10,863 years ago +- 2990 years, giving it a range of 7,873 – 13,853.
Is this Mexican F1a1 family Native? If not, how did F1a1 arrive in Mexico, and when? F1a1 is not found in either Europe or Africa.
In August, 2015, an article published in Science, Genomic evidence for the Pleistocene and recent population history of Native Americans by Raghaven et al suggested that a secondary migration occurred from further south in Asia, specifically the Australo-Melanesians, as shown in the diagram below from the paper. If accurate, this East Asian migration originating further south could explain both the haplogroup M and F results.
A second paper, published in Nature in September 2015 titled Genetic evidence for two founding populations of the Americas by Skoglund et al says that South Americans share ancestry with Australasian populations that is not seen in Mesoamericans or North Americans.
The Genographic project has no results for F1a1 outside of Asia.
I have not yet extracted the balance of haplogroup F in the Genographic project to look for other indications of haplogroups that could potentially be Native.
Haplogroup F Project
The haplogroup F project at Family Tree DNA shows no participants in the Americas, but several in Asia, as far south as Indonesia and also into southern Europe and Russia.
Haplogroup F Summary Table
Haplogroup F1a1 deserves additional attention as more people test and additional samples become available.
Native Mitochondrial Haplogroup Summary
Research in partnership with the Genographic Project as well as the publicly available portions of the projects at Family Tree DNA has been very productive. In total, we now have 259 proven Native haplogroups. This research project has identified 114 new Native haplogroups, or 44% of the total known haplogroups being newly discovered within the Genographic Project and the Family Tree DNA projects.
Acknowledgements
- National Geographic Society Genographic Project
 and Dr. Miguel Vilar, Science Manager
and Dr. Miguel Vilar, Science Manager - Family Tree DNA for providing the projects and support that enables us to further both scientific and genealogical research.
- The various haplogroup projects A, A2, A4, A10, B2, C, D, X, M, F and the project administrators of those projects.
- Dr. David Pike and Marie Rundquist, my co-administrators for the American Indian project
- Dr. Doron Behar for mtDNA Community and his publication, “A Copernican” Reassessment of the Human Mitochondrial DNA Tree from its Root”
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research
Discover more from DNAeXplained - Genetic Genealogy
Subscribe to get the latest posts sent to your email.

































































Roberta, you are one smart girl to put all this together. My head is still spinning. Great job!
Thanks for the detailed review! ^__^
Just a quick comment, looking at the genographic hotmap for haplogroup D. Weren’t Aleutians and Amazon basin Aboriginals the ones with a hint of Australasian? Could mt-haplogroup D be a hint for these Australasian migration?
I don’t know. I wouldn’t take that off the table.
It’s definitely too soon to conclude anything, and maybe it’s just due to founder effect, but maybe there is something to it after all. They’ll probably need to work more on haplogroup D in Asia though.
On a less quick comment, the part on haplogroup M and F are intriguing, I wish I could fast forwards in time to get more info about it.
Haplogroup X, still forming an odd distribution pattern, X2a still only found among Americas aboriginals. X2b isn’t totally ruled out yet although Radegonde Lambert X2a4 is.
It seems Catherine Pillard’s case isn’t settled with her A10.
Has any further information been gleaned regarding Catherine Pillard’s A10? Thank you!
Most of the matches to her descendants are to other known descendants. There are three or four that are more current, in the US, with unknown ancestors.
I’m in the process of reading DNA For Native American Genealogy for which I offer my congratulations and thanks. Six years ago we tested my second cousin who is in a direct maternal line from a woman who was the wife of a fur trader in early 19th Century Canada. She would have been either Anishinaabe or perhaps, Lakota. We have the paper traill. We don’t know her band. My cousin in C4c2. We’re curious as to why the information on this haplogroup seems so thin. Both of us have joined the American Indian Project of FTDNA. Is there more that we could contribute?
Hi Ellen. C4c2 is uncommon. If more people tested, especially Native people, we would know more. In terms of contributing more, please be sure to upload your tree and link the tester to their profile in the tree. Link other relatives too so that the system can do phasing for you. Do you have matches you can work with?
Not exactly sure where you would like the tree uploaded to. The couple from whom my cousin and I are decended had 9 children, 5 of whom left descendants. We have lots of matches. In addition, Thomas Taylor’s father (English) and mother (Cree) had 9 children, at least 5 of whom left descendants. The C4c2, however, comes from a woman identified only as “the second daughter of Jean Bte Cadot.”
You can upload your tree to Family Tree DNA. This article will help. https://dna-explained.com/2019/11/06/triangulation-in-action-at-family-tree-dna/
Roberta, Thank you for your comprehensive blog / report. Your blog will inspire more to test. As with the X2b4 study, I expect there will be new participants who join the mtDNA studies whose family lines and reports of earliest origins yield new answers to old, persistent questions. I am trying to live 100 years (watching what I eat and taking my vitamins!) so I can be around when some of these questions are finally resolved! Wonderful assessment. Thank you again.
I hope we both live long enough to see a lot of questions answered!!!
Roberta, and I hope we live long enough for some of those Family Bibles to come out of hiding…….
Pingback: Native American Mitochondrial Haplogroups | DNAeXplained – Genetic Genealogy
Still really curious about F1a1. At 23andme my paternal aunt (and a couple of cousins whose mother was a sister to my father) was predicted to be F1a. James Lick’s tool says F1a4a which is a haplogroup known to be in the Philippines where my paternal grandmother was from. And there was ties to New Spain prior to Mexico’s independence for 3 centuries as the Philippines was governed by New Spain during the Manila-Acapulco Galleon trade where some O haplogroups entered into Mexico. But know nothing about this F1a1. It would be interesting to see other descendants who has ties to the same place that this F1a1’s ancestress’ was from in order to see if it leads back to one woman or multiple.
now i can’t help but wonder if there’s other y-dna haplogroups in the americas like O
It’s certainly a possibility.
Hello, I recently discovered I am part of haplogroup B, and gedmatch has me up to 62% native. The problem is, I’m not sure where it comes from. I’m not too sure about my family history. How can I determine whether it comes from North or South America? I only have four matches, and the ones with more native have their locations hidden. Thanks.
Test your mtDNA with FTDNA and get a full sequence. You might be able to match up with someone else that know their direct ancestry. Also, I am surprised that you don’t know where your grandparents or earlier ancestors came from.
Very interesting post. The heat map for X is certainly interesting.
Pingback: Concepts – The Faces of Endogamy | DNAeXplained – Genetic Genealogy
what parts of the americas is mtdna b2r found in?
There is an academic paper being prepared with additional information.
ok, if so, it would be cool if it was associated with the tribes refered to as the chichimecas by the spanish and aztec/mexica people. I was just asking since it’s not listed on your native american haplogroups page, and my great grandpa(who died in 2000)’s only living brother left, appears to be b2r if jameslick’s mtdna tool is correct.
Excellent job! Great information!
Wondering if you have information about mtDNA D1e?
I don’t. Do you? If so, I would appreciate an e-mail at robertajestes@att.net.
I am D1e as well, have you found something?
I’m D1 e too, have you found any further information about this haplogroup?
You can keep current with what I have at this link: https://dna-explained.com/2013/09/18/native-american-mitochondrial-haplogroups/ If you have taken the mitochondrial DNA test at Family Tree DNA, please join the American Indian project. If you know your tribal affiliation, I’ll be glad to add that information to the D1e information.
Thankyou! Unfortunately I was adopted and know nothing about my biological relatives. Still surprised to find this particular haplogroup and huge number of DNA relatives in California, Texas and New Mexico, because I lived my role life in Brasil.
I am mtdna haplo D1 also. Where did your ggg grandmother live?
This says “New Native American Mitochondrial DNA Haplogroups, Estes, 2017,” however I don’t see any information about B2y.
It’s listed in the B chart. Look again. This is the first publication it was ever listed in, so there is little to no information about it elsewhere. Are you B2y?
Yes, and trying to find any info that I can about it.
Three distant cousins (discovered through genealogy) have common B2a2, B4’5. We know the region in Mexico where our family lived in 1820’s. Does this mean that it is meaningful to look at the Native population in that area as the home of the foremother’sl line?
I am B 4’5. After reviewing the information I ‘m not sure if my group is definitely Native or not. Can anyone please explain? Thanks.
Miss Roberta, it is true when a human mtDNA may can partially explain about our Ethnic group? Maybe because Men generally have a lot movement than their Female counterpart. About 30% of African American have European Y DNA and Autosomal DNA, but only 2 – 3% African American have European mtDNA and the Ethnic Group
But Polynesians, Micronesians and Eastern Indonesian have more “Native Pasific Islander” Maternal Haplogroup B4a and B4c then their Paternal Haplogroup C-M130, K-M9, M and S.
I suspect there are numerous cultural and geographic reasons for discrepancies.
1. Napoleon Bonaparte mtDNA belongs to Haplogroup H.
Napoleon Bonaparte Y DNA belongs to Haplogroup E1b1b (YAP+),
But I bet Napoleon have around 90 -100% European Autosomal DNA and their Autosomal World Region. So do with the Wright Brothers, Adolf Hitler, Albert Einstein, Nicholas Cage, etc.
2. A Northern Han Chinese and Southern Han Chinese share a common Paternal Line (3 Chinese Farmer Supergrandfathers): a Y DNA Haplogroup O3-M122+, O-P201+, O-P164+, O-M134+ and O-M117. But they have quite different for their Autosomal DNA and Maternal sides. An mtDNA Haplogroups M7 and E (M Type), R11, B, R9 an F (N – R Type) are far more common in Southern Han Chinese and Southeast Asian. Contrast with a Northern Han Chinese mtDNA Haplogroups M8, M8a, CZ, C, Z, D4, D5, M11, G (M Type) and Haplogroup A, N9, Y,….. (N Type).
3. My Uniparentals DNA :
– Y DNA Haplogroup O- CTS5492, descendant from Y DNA Hg K2-M526.
– mtDNA Haplogroup B4c2, descendant from an mtDNA Hg N – R.
– And myOrigines 2.0 Autosomal DNA: 73% Southeast Asian + 25% Northeast Asian and less then <1% Siberian DNA.
Can test results exaggerate your Native American DNA? The % of NA DNA that came back for some of my family members & myself was a lot more than we expected. We don’t have record of Native Americans in our recorded history, save for one man in the early 1800s. There is a highly detailed paper trail to Spain in the late 1700s, and presumably the mixing had to happen after that or someone mixed married in (ie the NA man in the 1800s). Being hispanic from New Mexico, we assumed we had some Native American DNA, but thought it would be negligible, especially considering we look Caucasian/European in appearance, not Native American. 23andme.com showed my maternal grandma to have 22% NA DNA and me to have 13%, and my maternal aunt to have 27%. I ran the data through another free site and my NA DNA dropped to 10%. We have the B2a2 marker, so confirmed NA there. I am just wondering if it’s possible to be LESS of something than a test indicates?
All ethnicity results are estimates although Native being less would be extremely rare.
Shell, what do you mean about B2a2 marker?
Sorry probably used the wrong term. My maternal haplogroup is B2a2.
Oh yeah…and according to 23andme this haplogroup is most closely associated with the Pueblo people of the southwest USA and maybe the very northwest region of Mexico bordering it. These people are distinct from other indigenous Mexicans (who vary quite a bit also). That makes sense given my family history.
I would be interested to know what town/towns your ancestors lived. In my study of New Mexico history, It seems I keep finding evidence of Genizaro communities everywhere. Even into the San Luis Valley of Colorado.
Hi! Belen, NM is where my grandmother’s side comes from. My grandfather’s side is from Albuquerque, but not sure of any more specific towns. I had to look up “genizaro”. It seems that means Native Americans who were servants in “Spanish” households. My grandmother said her family had at one point had “Indian servants” and they claimed they worked voluntarily for food and shelter. My grandma thinks they were closer to slaves and that’s why the NA DNA is in the female line without much written record to back it up. That’s all just hearsay of course.
Hearsay and myth, we sometimes find that there is a bit of truth to hearsay and myth, the remains of Richard III being found in the car park in England in recent years, a myth since he died in 1485. A myth in my family was that my great grandmother was NA. Since my mtdna is U3a1b, and she is my maternal ancestor, she could NOT have been NA. BUT, her husband, my great grandfather, after she died, did marry a NA woman and they each had the same surname and were very distant cousins. So that was how the myth started that my great grandmother was NA.
Genizaro comes from the term Jannisary. During the Ottoman Empire young boys would be collected as taxes and then trained to be soldiers for the Sultan’s army. In New Mexico young Indians would be taken as slaves and then used to populate communities that would provide protection from Indian raids. Spanish/Mexican colonialists tended to be concentrated in the Rio Grande Valley namely in Santa Fe, Albuquerque, and to some extent Taos. Genizaro communities were placed along routes entering the Valley. Abiquiu, Rio Hondo, Ojo Caliente, Belen, south side of river in Santa Fe, Galastio, and Pecos. I have found some references to Genizaros in Las Vegas, so I suspect that Las Vegas and even Grants may have been Genizaro communities also.
Is difficult to track down, because as the Crown, and then later Mexico City tried to eliminate Indian slavery they were always able to find loopholes around the new policies. Over time the term Genizaro was dropped and more cryptic language was used. For instance the Abiquiu land grant uses the term “half Indians”. Initially they would just outright take or buy slaves. Usually the slaves were acquired from Indians raiding other tribes. Later the practice was disguised as paying ransom for which then the hostage would be in servitude to pay off the debt. After the treaty of Guadalupe Hildago, the Navajos complained to the first territorial Indian agent that the all their children are being stolen. When the agent asked how many, they said about half the tribe. I have found accounts of Indians selling slaves in the trade fairs in Santa Fe and Taos. Acoounts that the going price for a young Navajo boy was three horses. Also accounts of Indians trading slaves for lead and gun powder.
The first pioneer family in the San Luis Valley came from Taos and brought a slave with them. In 1930 a woman slave died in Onate, she had been taken in 1860 and was passed down from one generation to the next.
So I suspect the practice was more widespread than most realize, and it’s just one of those dark secrets few talk about. And since they we taken at a young age and raised in Spanish speaking households, they lost all their tribal family and cultural ties. Of course is just my suspicion.
Hi Shell. As a person whose father’s family are Latinos from New Mexico and Colorado, I can assert that “New Mexicans” are mostly descended from Native Americans. The problem is that New Mexico history is filled with Eurocentric assumptions. I would go further that additional lineages (that DNA sampling is not available) would show even more Native American ancestry.
I disagree they’re mostly Native Americans. They put up a pretty good fight and preserved their cultured. They werent hispanicized as much as other parts of colonial new spain. Data shows the average new mexican who identifies as such or as hispanic is about 70% European dna.
Maybe for your family that’s not true, but mine definitely has mostly Iberian dna (aka Spanish). Certainly families will differ of course.
I have ancestors from Spain traced to the 1800s whereas any native ancestors were 1700s. Everyone else was mixed, Spanish or some other kind of European. My great aunt did a pretty thorough genealogy.
I had asked a question earlier but followed through without input. My female ancestor appears to come from the Opata tribe, in the area near Arevechi, Mexico. She was born around 1830. After reading David Yetman’s book, I feel confident that my lineage is correct. My male line that married her are definitely Spanish. Feels good to dig in a bit deeper.
A new match yesterday on GEDmatch, for both my father and I, listed her maternal haplogroup as D1. She is also a match to one of our 2nd cousins (a Herring). Where I share X-DNA of 187cM with my father, a male Herring shares 100cM on X-DNA with her. We have been trying to figure out the relationship today. Interesting, thanks Roberta!
Thank you..very interesting, i compaired my DNA with Montana Baby..and i was 4.5 or 18 where most of my friends didn’t match, can you tell me a little more about that, i think it eas like 13,000 yrs back . From Montana, first natives in the area. Thank you.Margaret Morinmarg
Here’s an article. https://dna-explained.com/2014/10/18/anzick-12707-12556-ancient-one-52-ancestors-42/
Hi Roberta,
I’ve found out that my mitochondrial DNA is B4a and even though my interest is just to find a matching ethnical or/ and geographical group, I can’t find it in any database. Do you have any matching results for that haplogroup somewhere in your research, please?
Thanks for all your work, I’ve been reading it and it’s fascinating to a lay person like me.
Kind regards,
Sandra Fischer
I have B4 and B4a1a. Where did you test. If it isn’t a full mitochondrial sequence test, it needs to be. Do you know if your mother’s mother’s mother’s line is Native?
Hi Roberta,
thanks for your comments. Like you, I also replied in email form as well. I suspect there might be a native bloodline from my mother, but unfortunately, it’s just that, a suspicion. No proof yet. I will investigate further.
In my email to you, I attach the result page from the test.
Hello Tim here, my father had his haplogroup checked at family tree and he came back as qm3 and b45. I have my haplogroup checked at 23andMe and I am qm3 and B2. Is b45 and B2 the same haplogroup. At family tree it says b45 is native and I read that B2 is a Beringa founder. Does b45 come from B2 or vice a versa?🤔 I know I’m asking a few questions here but can somebody straighten me out on this🙋
23andMe does not do a full sequence, and they are still using an outdated reference model. A full sequence test at Family Tree DNA is the only way to obtain your full haplogroup, and using the current reference model.
While 23andme gives you an idea of what your haplogroup is – B2, as suggested, a full mitochondrial DNA sequencing test at FTDNA will either confirm that or give you a more specific haplogroup.
The B4’5 designation that you got with FTDNA is because you tested only HVR1 and HVR2, which was purchased through their mtDNA Plus test.
And B2 comes from B4’5.
???? No Blackfoot Confederate
Thanks for your research and for sharing it with the public. I’m Colombian. I recently had a 23andme test that shows my haplogroup is C1. I’ve heard most women of the Americas have Native mitochondrial DNA because mainly European men came to the Americas. Do you know where could I find more information about this? Thanks!
I am new at this research. My husband has A2 on his maternal side and Q1a3a on his father’s side. I thought that Native American DNA could only be found in maternal, but since finding this on Gedmatch, I am now curious. Input?
The information on GedMatch is entered by people by hand. This article will help you understand. https://dna-explained.com/2012/10/01/4-kinds-of-dna-for-genetic-genealogy/
Both MtDNA and YDNA can show Native American. MtDNA only shows a person’s mother, her mother, her mother, etc. It does not show the father’s maternal line. And YDNA only shows a person’s father, his father, his father, etc. It does not show the mother’s paternal line.
Is Y-DNA more accurate than Mt-DNA (full sequence)?
They test completely different things. The Y tests the direct paternal line only. Usually the surname line. The mtdna tests only the direct matrilineal, the mother’s mother’s mother’s line.
I am hap U5 which is very surprising to me. All my life I have been mistaken for NA! I do see I have0.01% NA among my complicated results, why cant Family Tree just give me a pie chart. I just cant wrap my head around all this info!
Ancestry does a pie chart for autosomal results.
Why would you even list Haplogroup C4 if you don’t consider C4 Native American. Shouldn’t Haplogroup C4 be with the largest population of C4 which is the Australian Aborigines?
Your chart list different Haplogroups and beside D4 and D4e1 which you list as Not Native American but perhaps both-of what? there are no other Native American Haplogroups discarded except for D2 you just put a No with no explanation.
My interest with this is why so many people fake their Native Indian and being I live on Tribal land and certainly understand by trial and error the complex nature of Tribal governing it gives me an idea of whom I’m dealing with when Tribal mentality goes ape crazy with authoritarianism.
People rushing to be the next Native American Nation while there exist around 560 Nations in the United States others in the hundreds are applying for Nation status.
My question is directed to why you list C4 as Not Native American and yet include it with Native American?
On questionable haplogroups, many people were equating no entry with “don’t know,” so I list what we do know if it’s questionable and people might assume Native when it’s not or is undetermined.
Seeking guidance. I had my DNA done on 23 and me. I have African, European, and Native American. It just gives my ancestor group from Africa, and raw data has been received. How do I take this info and where do I go to pursue more info on my native heritage. What is a rsid what does it mean etc.
An rsid is a position number, like a street address. Native resources are listed here: https://dna-explained.com/2017/11/01/native-american-dna-resources/
I had the autosomal test done as well as mtdna. my haplogroup went from U to U5. when viewing my matches I see quite a few native american surnames but my results list me as 97% European with no native american listed. Can there still be NA in my family?
You could match those people on non-Native lines. Surnames aren’t native, specifically.
Please answer this question:
A man with R L21 or really any non-Native American Y dna marries a full-blood Indian woman. Their child..a male marries a white woman with H mtdna. Their child..a female would present with non-Indian H mtdna and would not be represented as Indian even though 25% Native American? Is this why we are finding the discrepancies?
My grandaughter is 5% Indian with an H mother according to her tests. That would make either her H mother 10% Indian or her father. I have an H mtdna too but I would be 20% Indian and my father 40% and grandfather 80%? If..on her mother’s side..the same. How does this coincide with your theories?…Under my theory, an H or any European mtdna could turn out as Indian. Am I missing something?
Yes, you’re missing something. The Y and mtDNA lines only show whether they are Native on THAT line – and only that line. They can still be Native, but not through that direct line. By finding mt and Y DNA of their other ancestral lines, you can narrow them down one at a time, and confirm when you’ve found one that is Native.
Hello, dear Roberta. After receiving messages from FT and 23andMe stating that I had an abundance of Jewish matches, I decided to look into it deeper. I also checked the locations and luckily there were enough photo to convince me. Match with other Native American haplogroups especially X Moundbuilders was thrilling. A mental picture is brewing in my head now.
My husband is waiting for results from FT Big Y expected in Sep. Will touch base with you when they come in.
Take care of yourself! Enjoy all of your successes! It is a pleasure knowing you!
Warmest regards!!!
FTDNA recently answered a question for me as to why my relative’s DNA had found no match. She belongs to Haplogroup D4j7 which they said is very rare and referred me to this article. While this is fascinating, and we have Czech roots, I don’t really know where to go with this information to learn more. My relative (who turns 102 tomorrow!) would be excited to know more about her genealogy.
In this article, haplogroup D4j7 is identified as southern Siberian. It is very rare. I have not found it in Native people. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0015214 Happy birthday to your relative!
Thank you for the reply! My relative Ida’s (referred to above in haplogroup D4j7) maternal grandmother is my father’s>great grandmother. Would she and I technically share the same haplogroup at the point of our common ancestor?
I’m not sure I would phrase it that way – but yes, your common ancestor had that haplogroup which means it’s relevant to your family history!
I have a relative who’s Bavarian in heritage who has the same Mtdna. I think that Czechia & Bavaria may have been settled by people from south Siberia. I have a lot of 100% European relatives with Native American-looking haplogroups like A4, C4 that aren’t actually Native American
A probably irrelevant question, but do you think the heat map and distribution map (blue flags) were/are affected by the transferal of various tribes by the Incas from areas east of Lake Titicaca in Bolivia. My adopted son (A2) comes from the Salasaca Tribe in Ecuador which has a traditional history of being moved to the coast and the mountains of current day Ecuador during early Inca domination. Would this indicate that A2 could have been originally much heavier populated in the Bolivia area than the heat map might suggest.
That certainly could be.
I am poking around Chaco Culture and history. The heat map for D1 was intriguing, as it neatly excludes, and sharply, the “OasisAmerican” area. Then D2 puts a blob right in that area and nowehere else close.
Can you explain why Haplogroup A2o is skipped in charts? Several years ago, my mom did the mtdna test so she could find out the ethnicity of her mother. My orphan grandmother was taken to Abilene, Texas by a “good samaritan” in 1918 at about age 5, her name was Consuelo, but her actual birthday was unknown. We learned that there isn’t much known about A2o and wondered if it’s because of extinction? My mom did the Ancestry test as well and discovered that my grandmother probably come from San Luis Potosi, Mexico. The Ancestry website shows 1st-5th maternal cousin matches but skips 3rd cousins altogether. My mom’s thinks that my grandmother came from one of the many extinct indigenous tribes that populated the area of Mexico/Texas during the Mexican Revolution. On my mom’s FamilyTree mtdna Haplogroup Origins page, there are matches for both A and A2o haplogroups with United States (Native American) and Mexico (Native American) listed in HVR1, HVR1 AND HVR2 sections, could you clarify what that means? Thank you for your time!
We need individuals with confirmed locations who have taken the full sequence and are haplogroup A2o. It would be great if your Mom would do so and join the haplogroup A and American Indian projects at Family Tree DNA. Has she taken the full sequence? I’ve love to be able to work with you to update the table.
Hi Roberta, is there a reason why haplogroup A2o is either skipped on charts or listed as unknown? My mom used the FamilyTree mtdna test to find out about her mother’s ethnicity. My orphan grandmother showed up in Abilene, Texas by about age 5 in 1918. My mom is 76% Native from Mexico, 9% French, and 6% Spanish. We found out that my grandmother may have come from a Mexican indigenous tribe in San Luis Potosi that didn’t survive the Mexican Revolution. If genocide or disease lasts long enough, can it eliminate a generation? Would sub-haplogroups get lost in the shuffle? Thanks for your time!
You have a reversion at that location and the branch has not been further named. Quite interesting.
Wish this Catherine Pillard mystery was settled! I am a direct descendant of Catherine twice in my tree. From her daughter Catherine and her son Pierre in two different lines 😊
I tested with FTDNA and my ancestry is 34% New World (North and Central America) 55% European (50% Iberia and 5% British Isles), 0% South America and other ancestries.
I tested with 23andMe and my ancestry is 30% Native American (Amerindian) 30% Spain and Portugal and other ethnicities. My Haplogroup is B2 and possibly Quechua – my mother’s family is from the north of Chile.
I was born in Chile and my ancestors on my father’s side have been in Chile since 1810, mostly in the south, and my mother’s ancestors in the north, even longer. I knew that my mother had strong Spanish blood lines and my father English. My great-great-grandparents on both sides were born in Chile and spoke Spanish.
I was very surprised about the high percentage of Native American from both tests. I had never heard of any Native Americans in my family. How come FTDNA says 34% North America and 0% South America. What does ca. 30% mean? Did 30% of my ancestors in total have NA blood on both sides N&F? Or would only one Native American ancestor be enough for me to have such a high percentage? if so, would it have to be a recent ancestor?
I am surprised and confused. How do I find out more? Are there Haplogroup B2 research groups? Can you point me in the right directions?
Thank you so much!
I can’t really answer this question out of context. If you would like a consult, I’m glad to do that. Regarding research groups, there is a haplogroup B project at Family Tree DNA that you can join.
Chileans are a very mixed people. I myself am Colombian and am nearly half Native and half Spanish. Just like a lot of Colombians, Chileans have mixed ancestry, especially Mestizo.
The sad truth about people in Chile is that they are very race conscious and deny their Native American heritage. I have met many Chileans that are obviously mestizos but always refer to themselves as white. It’s pretty sad. Most likely your family was trying to hide this as the whiter you are, the higher your social standing is in Chile.
As for the Native American, most of mine is also from North America. I would not look into this too much as it seems DNA tests sometimes find it difficult to pinpoint where the DNA comes from. According to other DNA sites I have tried I could even be Chinese/Japanese lol
The Native component is most likely from females in your family as your haplogroup is native to the Americas. Mine is A2.
I had a mtDNA test through Oxford Ancestors (after reading The Seven Daughters of Eve), which states that I belong to haplogroup C. i would like to know if this came via an ancestor from Asia, or the Americas. I would also like to know which First Nation peoples / Native Americans have a high percentage of this group.
Clifford Tomos M.Th, Cymru-Wales
You would need to order a full sequence test at Family Tree DNA as the first step. That would refine your haplogroup.
Hi, I’m lost at C1c with no matches at all on FTDNA. What are available options for a consult or is there no point in a consult when there is no matches? Thank you.
That’s interesting. I wonder if you have a significant mutation. Have you joined the haplogroup C mtDNA project and the American Indian project?
Thank you for the reply. I just joined the American Indian project and requested admission to the group C project. So, I look forward to hearing more from them or at least learning more. I’m curious how I do not have any matches for an existing haplogroup. If a significant mutation is the reason, can someone identify about when the mutation occurred (how far back)?
Interesting paper, Ancient and modern genomics of the Ohlone Indigenous population of California, https://doi.org/10.1073/pnas.2111533119, including mitochondrial haplotypes of ancient people here, the following supplementary materials are available to download https://10.1073/pnas.2111533119. Haplogroups: C1f, D1, Q1a2a1a1, M, B2b, C1c1b.
Roberta my mother’s mtDNA haplogroup is A12a. Am I correct in assuming this is Native American?
I believe so but check the public tree at FamilyTreeDNA.
A12a is originally from Siberia (Buryat).
See https://www.ncbi.nlm.nih.gov/nuccore/EF397560 and https://www.ncbi.nlm.nih.gov/nuccore/AY519488
Derenko et al., 2007. Phylogeographic analysis of mitochondrial DNA in northern Asian populations. Am. J. Hum. Genet. 81 (5), 1025-1041
Starikovskaya et al., 2005. Mitochondrial DNA diversity in indigenous populations of the
southern extent of Siberia, and the origins of Native American haplogroups. Ann. Hum. Genet. 69 (PT 1), 67-89.
Most cases tested by FTDNA self-report as from Great Britain.
https://www.familytreedna.com/public/mt-dna-haplotree/A;name=A12a
Two cases self-report as of having Amerindian ancestry. Since these are self-reports rather than the result of identifying the matriarch through genealogical research, these reports must be considered suspect. Many North Americans wish they had Amerindian ancestry and cultivate selection error, a form of ancestral romantism.
You should complete your line of mothers as far back as possible and find who was your matriarch in North America. A professional genealogist can do this for you.
Jacques P. Beaugrand, PhD
i have 1 cuestion haplogroup C1b4 is originated in mexico?
I’ve had time to check. This is a good question and the Million Mito Project will likely help to resolve this question. In Build 17, the previous haplogroup A4b became A12a, so it’s not the same as before. Haplogroup A12 has been found in the
Chumash people in the paper from Breschini and Haversat 2008. However, 2008 is very early for full sequence results in academic samples. We also have one person who reports an Iroquois ancestor from Canada. However, we have several people reporting A12a from England. Jacques suggested tracking your mother’s line as far back as possible and I would agree with that suggestion.
In 2002 I had an mtDNA test with Oxford Ancestors, which said that I belong to haplogroup C. In 2020 I had a DNA test with LivingDNA, which confirmed that I am halpgroup C, and that my subclade is C4a1a, and that this is common amongst the Ojibwe people. Can you confirm this for me?
In my book, I provided a breakdown for each haplogroup by known tribe.
I am one of those persons whose family lore says that my 2nd great-grandmother was Cherokee. I have her name, birthdate, husband and children’s names. She lived in south-central Virginia and possibly north central North Carolina, but I cannot find any other records about her. If she was NA, the tribes in that area were/are Coharie, the Eastern Band of Cherokee, Catawba, Saponi, Tutelo, Occanechee, and the Waccamaw-Siouan. I have found a second cousin once removed who is on the direct matrilineal line from the ancestor in question. I would like to offer her the FamilyTreeDNA Family Finder + mtFull Sequence test. I have a copy of your book, but I’d like clarification. If her results come back with a haplogroup other than A, B, C, D, or X, does that confirm that this ancestor was NOT Native American? If it IS in one of those groups, then more work needs to be done. Is that correct? Thank you for your amazing work.
Yes, any other haplogroup is NOT Native. But within those haplogroups, only some subgroups are Native. So you’ve got that right.
In regards to my question above, my second cousin, once removed received her results from FamilyTreeDNA and her confirmed haplogroup is U2e2a1c. Since this is not in haplogroup A, B, C, D, or X, I would assume that means our common ancestor, thought to be Native, is not Native. However, on the FamilyTreeDNA website, in the mtDNA Haplogroup origins section, under the HVR1, HVR2, and Coding Region Matches, there is a match for United States (Native American). Does that mean there remains a possibility that this ancestor is Native American? I’m not sure where to go from here.
Origins is what someone, another tester, entered.