The results for males just started coming in yesterday. One of our blog subscribers was kind enough to allow me to use his results. You’ll notice that there is no identifying information about you on this page, so if you forward this to someone, and that is something you can do, you’ll need to be sure to sign it with your name. It comes as a “no-reply” type of e-mail from National Geographic, not from you.
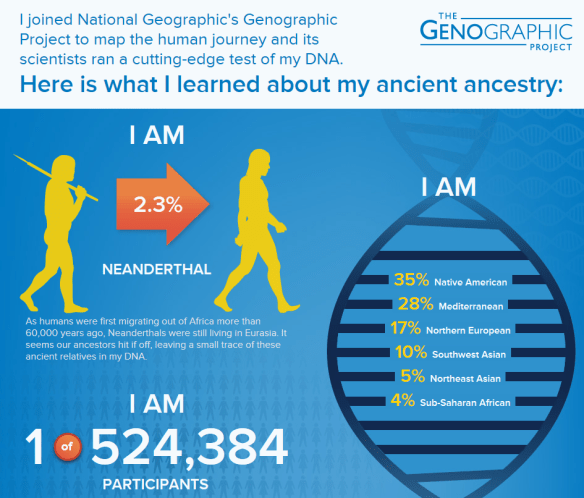
There’s lots of info provided here. First, you can see how much Neanderthal you carry. Ok, so no more Neanderthal jokes about your brother-in-law.
You can also see the division of your ethnicity. Compared to this person’s Family Finder results, this test seems to be more sensitive, picking up admixture not found in Family Finder. Their Family Finder results were 43% Europe, 5% East Asian (Siberian), 38% Native American and 13% Middle Eastern, rounded to the nearest percent. It looks like the Native American is about the same, the Middle Eastern may be absorbed by Mediterranean or Southwest Asia, and the Sub-Saharan African is new, but accurate according to this person’s genealogy.
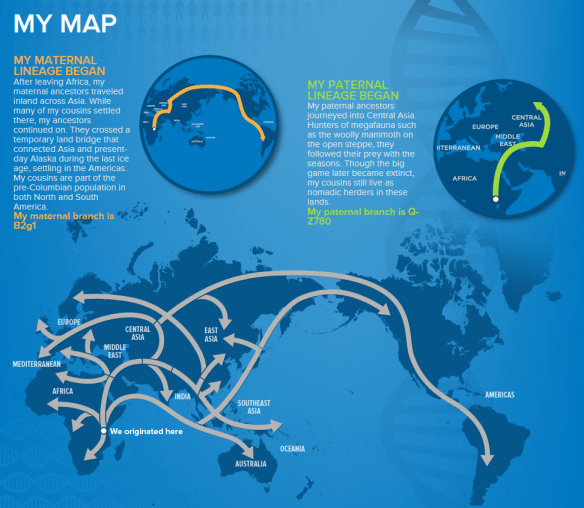
The second half of his display shows both the y-line and mitochondrial DNA map along with the migration path for the haplogroup. His mitochondrial DNA is B2g1. This is different from his B2 assigned at Family Tree DNA as a result of the full sequence test. His Y-line is haplogroup Q with a terminal SNP of Z780. He had tested for this SNP at Family Tree DNA also (as well as many others), and was classified as Q1a3.
It’s really exciting to see these results. Of course, now the questions begin, and there are already a lot of them. One of the first is about the ability to upload results to Family Tree DNA. Apparently you cannot do that if you have already SNP tested, have a mitochondrial DNA haplogroup assigned or have taken the Family Finder test. I sincerely hope this is simply a delay in development and that this will be addressed shortly. We need this information on our home pages.
Other questions are about the Y-line SNPs, which SNPs are included, and which aren’t, how to reference a new tree to see where you fit, and how has the tree, either the YCC tree at Family Tree DNA or the ISOGG tree, been shuffled. It’s obvious from seeing results for someone whose terminal SNP has not changed, but whose haplogroup has changed significantly that there has been major surgery to the tree. It’s difficult to figure out quite what you’re seeing at a deeper level.
And for autosomal of course, there are lots of questions about reference populations. Dave Dowell has already pointed out that there is a big discrepancy between his Geno 2.0 autosomal results and the ones returned by 23andMe last week. But 23andMe references 500 years and the Geno 2.0 test is billed as more of an anthropological test, so maybe they are measuring differently. Plus, there are all those new SNPs Nat Geo discovered and is using. I’m sure those are making a big difference too.
It’s been a great week or so for genetic genealogy, but yes, lots more questions than answers, so stay tuned.
______________________________________________________________
Disclosure
I receive a small contribution when you click on some of the links to vendors in my articles. This does NOT increase the price you pay but helps me to keep the lights on and this informational blog free for everyone. Please click on the links in the articles or to the vendors below if you are purchasing products or DNA testing.
Thank you so much.
DNA Purchases and Free Transfers
- Family Tree DNA
- MyHeritage DNA only
- MyHeritage DNA plus Health
- MyHeritage FREE DNA file upload
- AncestryDNA
- 23andMe Ancestry
- 23andMe Ancestry Plus Health
- LivingDNA
Genealogy Services
Genealogy Research
- Legacy Tree Genealogists for genealogy research
Discover more from DNAeXplained - Genetic Genealogy
Subscribe to get the latest posts sent to your email.



Hi,
It is not so much the autosomal SNPs but how you use them that determines how far back in time they trace. 23andMe is using a 100 SNP long run of SNPs in their new Ancestry Composition component. They are putting a 500 year time frame on the ancestral commonality of those 100 SNP long blocks being ‘in common’ with others.
Each individual SNP is much older and when it is an Ancestry Informative SNP you can go further back in time. You, of course, would need to ask Dr Wells just how his program works. 🙂
Best,
Rebekah
Thanks Rebekah. This is really helpful.
Thanks for this, I got my Geno 2.0 results yesterday but I hadn’t tried to email my results to myself or anyone. From the email I can see that I am 1% Neanderthal. I don’t know where to see this information on my Geno 2.0 results pages. At the bottom of my https://genographic.nationalgeographic.com/results/whoami page I see two circles that are supposed to be about Neanderthal and Denisovan ancestry, but I don’t see any percentages even when I click on the circles or “learn more”. Not a big deal.
Roberta, would you say this is the best test for me to take in order to secure a notion on how much of my lineage is Mediterranean? Or would another be better.
I took the National Geographic one several years ago and just was not excited about what I found.
When I get my own results to work with, I’ll be able to experiment some. It’s a learning curve for all of us right now, but what fun!
Granted, the Gen 2 vs 23andMe raises many questions but until they are recognized they can’t be answered can they. This is very exciting, keep the discussions and comparisons coming.
Do we know if the samples from the first Geno were used in the Geno2.0 dataset for the autosomal analysis? 524,384 participants seems to be all-inclusive. I also wonder if they used some of the public datasets as 23andMe did (eg. HGDP and 1000 Genomes Project)
By the way, I’m still at 60% and anxiously awaiting results.
They weren’t used for autosomal analysis, but the number of kits is included in the 524,384 number.
I sometimes think this is all getting overhyped. And I suspect the majority of British folk will be underwhelmed. This is not to belittle the work being done, but this is not why people take DNA tests to do family history in the UK. I think the migration chart at the top doesn’t tell us anything much that we did not know already. Now I have not spent much on autosomal DNA tests apart from dipping my toe in the water – but I want to have enough time to do real research using real written records. If funds are limited – and it is a toss-up between STR 111 markers and autosomal DNA testing – then it is a no-brainer in my book.
I think if the work of the People of the British Isles DNA Project leads to works of DNA analysis which can get far closer to the genetic homeland of people within a country – it will prove to be of far more interest to people than this broad-brush stuff.
A male friend just received his results today and was able to upload them to his FTDNA account, although he had not previously taken expanded SNP nor mtDNA tests, just the previous Geno.
Of course there was some surprise as it showed A2d1 as his maternal haplogroup when he had never been aware of any Native American heritage. (6% on his autosomal test) Both his and the reference examples shown (Greek and Iberian) had 10-15% Southwest Asian that was a little confusing to both of us, even after reviewing the site’s explanations. I’ll be very curious to see how that displays on my own results.
I stand corrected. My friend had taken the Deep Clade test with FTDNA in early 2010. So that did not prevent uploading Geno2.0 to FTDNA.
Pingback: Geno 2.0 Results – Kicking the Tires | DNAeXplained – Genetic Genealogy
Pingback: Proving Native American Ancestry Using DNA | DNAeXplained – Genetic Genealogy
Pingback: Proving Native American Ancestry Using DNA | Native Heritage Project
Pingback: 2012 Top 10 Genetic Genealogy Happenings | DNAeXplained – Genetic Genealogy
Pingback: Autosomal Testing Comparison | DNAeXplained – Genetic Genealogy
I received a notification last week from Genographic 2.0 about an update to the Y tree. Have you heard anything about that? I went from an R-DF19 to an R-v221 (if I recall correctly). FTDNA still shows me as and R-DF19.
Pingback: DNAeXplain Archives – General Information Articles | DNAeXplained – Genetic Genealogy